Abstract
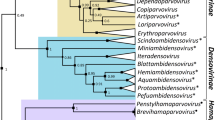
An exhaustive evolutionary analysis of the picornavirus family has been carried out using the amino acid sequences of several proteins of the viruses including: the capsid proteins (1D, 1B, and 1C) situated at the 5′ end of the genome and responsible for the serotype of the viruses, and the viral polymerase (3D), located at the 3′ end of the genome. The evolutionary relationships found among the viruses studied support the new classification, recently suggested, in contrast to the classical one, and the existence of a new genus for the picornavirus family. In the new taxonomic organization, five genera form the picornavirus family: (1) aphthoviruses, (2) cardioviruses, (3) hepatoviruses (previously classified as enteroviruses), (4) renteroviruses (which mainly constitute a combination of the previous genera rhinovirus and enterovirus), and (5) a new genus, with a new and unique representative: the echovirus 22. Our analysis also allowed us, for the first time, to propose the most probable sequence of speciation events to have given rise to the current picornavirus family.
The bootstrap procedure was used to check the reliability of the phylogenetic trees obtained. The application of the method of the statistical geometry in distance space to internal branches of the tree revealed a high degree of evolutionary “noise,” which makes the resolution of some internal branching points difficult.
Similar content being viewed by others
References
Adachi J, Cao Y, Hasegawa M (1993) Tempo and mode of mitochondrial DNA evolution in vertebrates at the amino acid level: rapid evolution in warm-blooded vertebrates. J Mol Evol 36:270–281
Bae Y-S, Eun H-M, Yoon J-W (1989) Genomic differences between diabetigenic and nondiabetigenic encephalomyocarditis virus. Virology 170:282–287
Buonaugurio DA, Nakada S, Fitch WM, Palese P (1986) Epidemiology of influenza C virus in man: multiple evolutionary lineages and low rate of change. Virology 153:12–21
Carroll AR, Rowlands DJ, Clarke BE (1984) The complete nucleotide sequence of the RNA coding for the primary translation product of foot and mouth disease virus. Nucleic Acids Res 12:2461–2472
Chang K, Auvinen P, Hyypia T, Stanway G (1989) The nucleotide sequence of coxsackievirus A9; implications for receptor binding and enterovirus classification. J Gen Virol 70:3269–3280
Cohen JI, Ticehurst JR, Purcell RH, Bukler-White A, Baroudy BM (1987) Complete nucleotide sequence of wild-type hepatitis A virus: comparison with different strains of hepatitis A virus and other picornaviruses. J Virol 61:50–59
Cohen SH, Naviaux RK, Vanden Brink KM, Jordan GW (1988) Comparison of nucleotide sequences of diabetogenic and nondiabetogenic encephalomyocarditis virus. Virology 166:603–607
Cooper PD, Agol VI, Bachrach HL, Brown F, Ghendon Y, Gibbs AJ, Gillespie JH, Lonberg-Holm K, Mandel B, Melnick JL, Mohanty SB, Povey RC, Rueckert RR, Schafter FL, Tyrrell DAJ (1978) Picomaviridae: second report. Intervirology 10:165–180
Dayhoff M (1979) Atlas of protein sequence and structure, vol. 5. Supplement 3, 1978. National Biomed Research Fundation, Washington DC
Dopazo J (1994) Estimating errors and confidence intervals for branch lengths in phylogenetic trees by a bootstrap approach. J Mol Evol 38:300–304
Dopazo J, Dress A, von Haeseler A (1993) Split decomposition: a new technique to analyze viral evolution. Proc Natl Acad Sci USA 90:10320–10324
Dopazo J, Rodrigo MJ, Rodríguez A, Sáiz JC, Sobrino F (1994) Aphthovirus evolution. In: Gibbs A, Calisher CH (eds). Molecular evolution of viruses. Cambridge Univ Press, Cambridge. In press
Dopazo J, Sobrino F, Palma EL, Domingo E, Moya A (1988) VP1 protein gene of foot-and-mouth disease virus: a quasispecies model of molecular evolution. Proc Natl Acad Sci U S A 85:6811–6815
Duechler M, Skern T, Sommergruber W, Neubauer C, Druendler P, Fogy I, Blaas D, Kuechler E (1987) Evolutionary relationships within the human rhinovirus genus: comparison of serotypes 89, 2 and 14. Proc Natl Acad Sci U S A 84:2605–2609
Duke GM, Hoffman MA, Palmenberg AC (1992) Sequence and structural elements that contribute to efficient encephalomyocarditis virus RNA translation. J Virol 66:1602–1609
Earle JAP, Skuce RA, Fleming CS, Hoey EM, Martin SJ (1988) The complete nucleotide sequence of a bovine enterovirus. J Gen Virol 69:253–263
Efron B (1982) The Jackknife, the bootstrap and other resampling plans. Society for Industrial and Applied Mathematics, Philadelphia
Efron B, Gong G (1983) A leisurely look at the bootstrap, the jackknife and cross-validation. Am Stat 37:36–48
Efron B, Tibshirani R (1991) Statistical data analysis in the computer age. Science 253:390–395
Eigen M, Lindemann BF, Tietze M, Winkler-Ostwatitsch R, Dress A, von Haeseler A (1989) How old is the genetic code? Statistical geometry of tRNA provides an answer. Science 244:673–679
Eigen M, Nieselt-Struwe K (1990) How old is the immunodeficiency virus? AIDS 4:S85-S93
Eigen M, Winkler-Ostwatitsch R (1981) Transfer-RNA: the early adaptor. Naturwissenchaften 68:217–228
Eigen M, Winkler-Ostwatitsch R (1990) Statistical geometry in sequence space. Methods Enzymol 183:505–530
Eigen M, Winkler-Ostwatitsch R, Dress A (1988) Statistical geometry in sequence space: a method of quantitative comparative sequence analysis. Proc Natl Acad Sci USA 85:5913–5917
Felsenstein J (1973) Maximum likelihood and minimum-steps method for estimating evolutionary trees from data on discrete characters. Syst Zool 22:240–249
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Felsenstein J (1988) Phylogenies from molecular sequences: inference and reliability. Annu Rev Genet 22:521–565
Felsenstein J (1993) PHYLIP (phylogeny inference package) version 3.5. Department of Genetics, University of Washington, Seattle
Fitch WM, Margoliash E (1967) Construction of phylogenetic trees. Science 155:279–284
Forss S, Strebel K, Beck E, Schaller H (1984) Nucleotide sequence and genome organization of foot-and-mouth disease virus. Nucleic Acids Res 12:6587–6601
Gorbalenya AE, Donchenko AP, Blinov VM, Koonin EV (1989) Cystein proteases of positive strand RNA viruses and chymotrypsinlike serine proteases. FEBS Lett 243:103–114
Graff J, Normann A, Feinstone SM, Flehmig B (1994) Nucleotide sequence of wild-type hepatitis A virus GBM in comparison to two cell culture adapted variants. J Virol 68:548–554
Higgins DG, Bleasby AJ, Fuchs R (1992) CLUSTAL V: improved software for multiple sequence alignment. Comput Appl Biosci 8:189–191
Hughes PJ, North C, Jellis CH, Minor PD, Stanway G (1988) The nucleotide sequence of human rhinovirus 1B: molecular relationships within the rhinovirus genus. J Gen Virol 69:49–58
Hughes PJ, North C, Minor PD, Stanway G (1989) The nucleotide sequence of coxsackievirus A21. J Gen Virol 70:2943–2952
Hyypia T, Horsnell M, Maronen M, Khan M, Kalkkinen N, Auvinen P, Kinnunen L, Stanway G (1992) A distinct picornavirus group identified by sequence analysis. Proc Natl Acad Sci USA 89:8847–8851
Iizuka N, Kuge S, Nomoto A (1987) Complete nucleotide sequence of the genome of coxsackievirus B1. Virology 156:64–73
Inoue T, Suzuki T, Sekiguchi K (1989) The complete nucleotide sequence of swine vesicular disease virus. J Gen Virol 70:919–934
Inoue T, Yamaguchi S, Kanno T, Sugita S, Saeki T (1993) The complete nucleotide sequence of a pathogenic swine vesicular disease virus isolated in Japan (J1′73) and phylogenetic analysis. Nucleic Acids Res 21:3896–3896
Jansen RW, Siegl G, Lemon SM (1990) Molecular epidemiology of human hepatitis A virus defined by an antigen-capture polymerase chain reaction method. Proc Natl Acad Sci USA 87:2867–2871
Jenkins O, Booth JD, Minor PD, Almond JW (1987) The complete nucleotide sequence of coxsackievirus B4 and its comparison to other members of the picornaviridae. J Gen Virol 68:1835–1848
Kinnunen L, Huovilainen A, Pöyry T, Hovi T (1990) Rapid molecular evolution of wild type poliovirus during infection in individual hosts. J Gen Virol 71:317–324
Kinnunen L, Pöyry T, Hovi T (1991) Generation of virus genetic lineages during an outbreak of poliomyelitis. J Gen Virol 72:2483–2489
Kishino H, Miyata T, Hasegawa M (1990) Maximum likelihood inference of protein phylogeny and the origin of choroplasts. J Mol Evol 31:151–160
Klump WM, Bergmann I, Muller BC, Ameis D, Kandolf R (1990) Complete nucleotide sequence of infectious coxsackievirus B3 cDNA: two initial 5′ uridine residues are regained during plusstrand RNA synthesis J Virol 64:1573–1583
Koonin EV, Gorbalenya AE (1992) An insect picornavirus may have genome organization similar to that of caliciviruses. FEBS Lett 297:81–86
Law KM, Brown TDK (1990) The complete nucleotide sequence of the GDVII strain of Theiler's murine encephalomyelitis virus (TMEV). Nucleic Acids Res 18:6707–6708
Li W-H (1989) A statistical test of phylogenies estimated from sequence data. Mol Biol Evol 6:424–435
Li W-H, Gouy M (1990) Statistical tests of molecular phylogenies. Methods Enzymol 183:645–659
Li W-H, Tanimura M, Sharp PM (1987) An evaluation of the molecular clock hypothesis using mammalian DNA sequences. J Mol Evol 25:330–342
Margush T, McMorris FR (1981) Consensus n-trees. Bull Math Biol 43:239–244
Martínez MA, Dopazo J, Hernández J, Mateu MG, Sobrino F, Domingo E, Knowles NJ (1992) Evolution of the capsid protein genes of foot-and-mouth disease virus: antigenic variation without accumulation of amino acid substitutions over six decades. J Virol 66: 3557–3565
Martinez-Salas E, Ortin J, Domingo E (1985) Sequence of the viral replicase gene from foot-and-mouth disease virus C-1-Santa Pau (C-S8). Gene 35:55–61
Miyamura K, Tanimura M, Takeda N, Kono R, Yamazaki S (1986) Evolution of enterovirus 70 in nature: all isolates were recently derived from a common ancester. Arch Virol 89:1–14
Miyamura K, Takeda N, Tanimura M, Yamazaki S (1989) Evolution of enterovirus 70 and coxsackievirus A24 variant. In: Ishii K, Uchida Y, Niyamura K, Yamazaki S (eds) Acute hemorragic conjunctivitis. University of Tokyo Press, Tokyo, pp 399–418
Mueller L, Ayala FJ (1982) Estimation and interpretation of genetic distance in empirical studies. Genet Res 40:127–137
Najarian R, Caput D, Gee W, Potter S, Renard A, Merryweather J, Van Nest G, Dina D (1985) Primary structure and gene organization of human hepatitis A virus. Proc Natl Acad Sci USA 82:2627–2631
Nei M, Jin L (1989) Variance of the average numbers of substitutions within and between populations. Mol Biol Evo1 6:290–300
Nei M, Sthepens JC, Saitou N (1989) Methods for computing the standard errors of branching points in an evolutionary tree and their application to molecular data from human and apes. Mol Biol Evol 2:66–85
Nichol ST, Rowe JE, Fitch WM (1989) Glycoprotein evolution of vesicular stomatitis virus New Jersey. Virology 168:281–291
Nomoto A, Omata T, Toyoda H, Kuge S, Horie H, Kataoka Y, Genba Y, Nakano Y, Imura N (1982) Complete nucleotide sequence of the attenuated poliovirus Sabin 1 strain genome. Proc Natl Acad Sci USA 79:5793–5797
O'Hara Y, Stein S, Fu J, Stillman L, Klaman L, Roos RP (1988) Molecular cloning and sequence determination of DA strain of Theiler's murine encephalomyelitis viruses. Virology 164:245–255
Palese P, Young JF (1982) Variation of influenza A, B and C viruses. Science 215:1468–1474
Palmenberg A (1989) Sequence alignments of picornaviral capsid proteins. In: Semler BL, Ehrenfeld E (eds) Molecular aspects of picornavirus infection and detection. American Society for Microbiology, Washington, p 211
Pamilo P (1990) Statistical tests of phenograms based on genetic distances. Evolution 44:689–697
Perez-Bercoff R (1987) Picornavirus at the molecular level. In: Perez-Bercoff R (ed) The molecular basis of viral replication. Plenum Press, New York, p 197
Pevear DC, Borkowsky J, Calenoff M, Oh CK, Ostrawski B, Lipton HL (1988) Insights into Theiler's virus neurovirulence bases on genomic comparison of the neurovirulent GDVII and less virulent BaAn strains. Virology 165:1–12
Porter AG (1993) Picornavirus nonstructural proteins: emerging roles in virus replication and inhibition of host cell functions. J Virol 67:6917–6921
Pöyry T, Kinnunen L, Kapsenberg J, Kew J, Hovi T (1990) Type 3 poliovirus/Finland/1984 is genetically related to common mediterranean strains. J Gen Virol 71:2535–2541
Rico-Hesse R, Pallansch M, Nottay BK, Kew OM (1987) Geographic distribution of wild poliovirus type 1 genotypes. Virology 160:311–322
Robertson BH, Grubman MJ, Weddell GN, Moore DM, Welsh JD, Fischer T, Dowbenko DJ, Yansura DG, Small B, Kleid DG (1985) Nucleotide and amino acid sequence coding for polypeptides of foot-and-mouth disease virus type A12. J Virol 54:651–660
Rueckert RR (1985) Picornaviruses and their replication. In: Fields BN (ed) Virology. Raven Press, New York, p 705
Ryan MD, Jenkins O, Hughes PJ, Brown A, Knowles NJ, Booth D, Minor PD, Almond JW (1990) The complete nucleotide sequence of enterovirus type 70: relationships with other members of the picornaviridae. J Gen Virol 71:2291–2299
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sáiz JC, Sobrino F, Dopazo J (1993) Molecular epidemiology of foot-and-mouth disease virus type O. J Gen Virol 74:2281–2285
Seechurn P, Knowles NJ, McCauley JW (1990) The nucleotide sequence of a pathogenic swine vesicular disease virus. Virus Res 16:255–274
Skern T, Sommergruber W, Blaas D, Gruendler P, Fraundorfer F, Pieler C, Fogy I, Kuechler E (1985) Human rhinovirus 2: complete sequence and proteolytic processing signals in the capsid protein region. Nucleic Acids Res 13:2111–2126
Sneath PHA, Sokal RR (1973) Numerical taxonomy. Freeman, San Francisco
Sobrino F, Palma EL, Beck E, Dávila M, de la Torre JC, Negro P, Villanueva N, Ortín J, Domingo E (1986) Fixation of mutations in the viral genome during and outbreak of foot-and-mouth disease: heterogeneity and rate variations. Gene 50:149–159
Stanway G (1990) Structure, function and evolution of picornaviruses. J Gen Virol 71:2483–2501
Stanway G, Hughes PJ, Mountford RC, Minor PD, Almond JW (1989) The complete nucleotide sequence of a common cold virus: human rhinovirus 14. Nucleic Acids Res 12:7859–7875
Supanaranond K, Takeda N, Yamazaki S (1992) The complete nucleotide sequence of a variant of Coxsackievirus A24, an agent causing acute hemorrhagic conjunctivitis. Virus Genes 6:149–158
Tajima F (1992) Statistical method for estimating the standard errors of branch lengths in a phylogenetic tree reconstructed without assuming equal rates of nucleotide substitutions among different lineages. Mol Biol Evol 9:168–181
Tanimura M, Miyamura K, Takeda N (1985) Construction of a phylogenetic tree of enterovirus 70. Jpn J Genet 60:137–150
Toyoda H, Kohara M, Kataoka Y, Suganuma T, Omata T, Imura N, Nomoto A (1984) Complete nucleotide sequences of all three poliovirus serotype genomes. J Mol Biol 174:561–585
Tsarev SA, Emerson SU, Balayan MS, Ticehurst JR, Purcell RH (1991) Simian hepatitis A virus (HAV) strain AGM-27: comparison of genome structure and growth in cell culture with other HAV strains. J Gen Virol 72:1677–1683
Winkler-Ostwatitsch R, Dress A, Eigen M (1986) Comparative sequence analysis. Chem Scripta 26B:59–66
Yamashita M, Krystal M, Fitch WM, Palese P (1988) Influenza B virus evolution: co-circulating lineages and comparison of evolutionary pattern with those of influenza A and C viruses. Virology 163:112–122
Zhang G, Wilsden G, Knowles NJ, McCauley JW (1993) Complete nucleotide sequence of a coxsackie B5 virus and its relationship to swine vesicular disease virus. J Gen Virol 74:845–853
Zharkikh A, Li W-H (1992) Statistical properties of bootstrap estimation of phylogenetic variability from nucleotide sequences: I. four taxa with a molecular clock. Mol Biol Evol 9:1119–1147
Author information
Authors and Affiliations
Additional information
Correspondence to: J. Dopazo
Rights and permissions
About this article
Cite this article
Rodrigo, M.J., Dopazo, J. Evolutionary analysis of the picornavirus family. J Mol Evol 40, 362–371 (1995). https://doi.org/10.1007/BF00164022
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00164022