Abstract
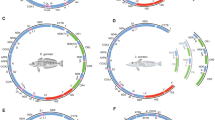
Mitochondrial gene content shows extensive variation among eukaryotes, but is remarkably compact and static in bilateral animals. Mitochondrial genomes of bilaterians typically contain two rRNA, 22 tRNA, and 13 protein-coding genes. In this study, we report that the mitochondrial genomes of Antarctic fishes of the suborder Notothenioidei (Perciformes) lack two adjacent genes encoding NADH 6 dehydrogenase (ND6) and tRNAGlu. Loss of the ND6 gene is reported for the first time in an animal mitochondrial genome, and is considered an extremely rare evolutionary event. Dot blot and ND6 transcript detection analyses found no evidence of mitochondrial ND6 gene copies in heteroplasmy or of a functional ND6 gene copy in the nuclear genome, respectively. Hence, we concluded that ND6 function was lost in Antarctic notothenioids, and could be compensated for by functional changes in other proteins of the mitochondrial respiratory system.




Similar content being viewed by others
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105
Adams KL, Palmer JD (2003) Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Mol Phylogenet Evol 29:380–395
Adams KL, Qiu YL, Stoutemyer M, Palmer JD (2002) Punctuated evolution of mitochondrial gene content: high and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc Natl Acad Sci USA 99:9905–9912
Bai YD, Attardi G (1998) The mtDNA-encoded ND6 subunit of mitochondrial NADH–dehydrogenase is essential for the assembly of the membrane arm and the respiratory function of the enzyme. EMBO J 17:4848–4858
Bargelloni L, Marcato S, Patarnello T (1998) Antarctic fish hemoglobins: evidence for adaptive evolution at subzero temperature. Proc Natl Acad Sci USA 9515:8670–8675
Bargelloni L, Marcato S, Zane L, Patarnello T (2000) Mitochondrial phylogeny of Notothenioids: a molecular approach to Antarctic fish evolution and biogeography. Syst Biol 49:114–129
Borner GV, Morl M, Janke A, Pääbo S (1996) RNA editing changes the identity of a mitochondrial tRNA in marsupials. EMBO J 15:5949–5957
Brand MD (2000) Uncoupling to survive? The role of mitochondrial inefficiency in ageing. Exp Gerontol 35:811–820
Cheng C-HC, Chen L (1999) Evolution of an antifreeze glycoprotein. Nature 401:443–444
Deng JH, Li Y, Park JS, Wu J, Hu P, Lechleiter J, Bai Y (2006) Nuclear suppression of mitochondrial defects in cells without the ND6 subunit. Mol Cell Biol 263:1077–1086
DeVries AL (1988) The role of antifreeze glycopeptides and peptides in the freezing avoidance of Antarctic fishes. Comp Biochem Physiol 90B:611–621
Di Prisco G, Giardina B (2003) Molecular aspects of temperature adaptation. In: Di Prisco G, Giardina B, Weber RE (eds). Hemoglobin function in vertebrates. Molecular adaptations in extreme and temperate environments. Springer, Milan, pp 1–21
Dörner M, Altmann M, Pääbo S, Morl M (2001) Evidence for import of a Lysyl-tRNA into marsupial mitochondria. Mol Biol Cell 12:2688–2698
Dreyer H, Steiner G (2006) The complete sequences and gene organisation of the mitochondrial genomes of the heterodont bivalves Acanthocardia tuberculata and Hiatella arctica – and the first record for a putative ATPase subunit 8 gene in marine bivalves. Front Zool 2006, 3:13
Eastman JT (2005) The nature of the diversity of Antarctic fishes. Polar Biol 28:93–107
Eastman JT, Eakin RR (2000) An updated species list for notothenioid fish (Perciformes; Notothenioidei), with comments on Antarctic species. Arch Fish Mar Res 48:11–20
Fischer W, Hureau JC (eds) (1985) FAO species identification sheets for fishery purposes. Southern Ocean: fishing areas 48, 58 and 88 (CCAMLR convention area). Prepared and published with the support of the Commission for the Conservation of Antarctic Marine Living Resources. FAO, Rome. p 471
Gray MW, Lang BF, Cedergren R, Golding B, Lemieux C, Sankoff D, Turmel M, Brossard N, Delage E, Littlejohn TG, Plante I, Rioux P, Saint-Louis D, Zhy Y, Burger G (1998) Genome structure and gene content in protist mitochondrial DNAs. Nucleic Acids Res 26:865–887
Gu X, Vander Velden K (2002) DIVERGE: Phylogeny-based analysis for functional-structural divergence of a protein. Bioinformatics 18:500–501
Guindon S, Gascuel O (2003) A simple fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hoffmann RJ, Boore JL, Brown WM (1992) A novel mitochondrial genome organization for the blue mussel, Mytilus edulis. Genetics 131:397–412
Janke A, Xu X, Arnason U (1997) The complete mitochondrial genome of the wallaroo (Macropus robustus) and the phylogenetic relationship among Monotremata, Marsupialia, and Eutheria. Proc Natl Acad Sci USA 94:1276–1281
Johnston IA, Calvo J, Guderley H, Fernandez D, Palmer L (1998) Latitudinal variation in the abundance and oxidative capacities of muscle mitochondria in perciform fishes. J Exp Biol 201:1–12
Kilpert F, Podsiadlowski L (2006) The complete mitochondrial genome of the common sea slater, Ligia oceanica (Crustacea, Isopoda) bears a novel gene order and unusual control region features. BMC Genomics 2006, 7:241
Kim KH, Eom KS, Park JK (2006) The complete mitochondrial genome of Anisakis simplex (Ascaridida: Nematoda) and phylogenetic implications. Int J Parasitol 36:319–328
Knudsen B, Miyamoto MM (2001) A likelihood ratio test for evolutionary rate shifts and functional divergence among proteins. Proc Natl Acad Sci USA 98:14512–14517
Lang BF, O’Kelly CJ, Nerad TA, Gray MW, Burger G (2002) The closest unicellular relatives of animals. Curr Biol 12:1773–1778
La Roche J, Snyder M, Cook DI, Fuller K, Zouros E (1990) Molecular characterization of a repeat element causing large-scale variation in the mitochondrial DNA of the sea scallop Placopecten magellanicus. Mol Biol Evol 7:45–64
Liò P, Goldman N, Thorne JL, Jones DT (1998) PASSML: combining evolutionary inference and protein secondary structure prediction. Bioinformatics 148:726–733
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16:404–405
Medina M, Collins AG, Takaoka TL, Kuehl JV, Boore JL (2006) Naked corals: skeleton loss in Scleractinia. Proc Natl Acad Sci USA 103:9096–9100
Milbury CA, Gaffney PM (2005) Complete mitochondrial DNA sequence of the eastern oyster, Crassostrea virginica. Marine Biotechnol 7:697–712
Near TJ, Pesavento JJ, Cheng C-HC (2004) Phylogenetic investigations of Antarctic notothenioid fishes (Perciformes: Notothenioidei) using complete gene sequences of the mitochondrial encoded 16S rRNA. Mol Phyl. Evol 32:881–891
Near TJ, Parker SK, Detrich HW III (2006) A genomic fossil reveals key steps in hemoglobin loss by the antarctic icefishes. Mol Biol Evol 23:2008–2016
Nicholas KB, Nicholas HB Jr, Deerfield DW II (1997) GeneDoc: analysis and visualization of genetic variation. EMBNEW News 4:14
Norman JR (1937). Coast fishes. Part II. The Patagonian region. Discovery Rep 16:1–150
Okimoto R, Macfarlane JL, Wolstenholme DR (1990) Evidence for the frequent use of TTG as the translation initiation codon of mitochondrial protein genes in the nematodes, Ascaris suum and Caenorhabditis elegans. Nucleic Acids Res 18:6113–6118
Palumbi S, Martin AP, Romano S, McMillan WO, Stice L, Grabowski G (1991) The Simple Fool’s Guide to PCR. Version 20. Honolulu, HI, University of Hawaii, pp 46
Pereira SL (2000) Mitochondrial genome organization and vertebrate phylogenetics. Genet Mol Biol 234:745–752
Rost B (1996) PHD: predicting one-dimensional protein structure by profile based neural networks. Meth Enzymol 266:525–539
San Mauro D, Gower DJ, Oommen OV, Wilkinson M, Zardoya R (2004) Phylogeny of caecilian amphibians (Gymnophiona) based on complete mitochondrial genomes and nuclear RAG1. Mol Phylogenet Evol 33:413–427
Signorovitch AY, Buss LW, Dellaporta SL (2007) Comparative genomics of large mitochondria in placozoans. PLoS Genet 3:44–50
Wolstenholme DR (1992) Animal mitochondrial DNA: structure and evolution. In: Mitochondrial genomes. Int Rev Cytol 141:173–216
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Yokobori S, Fukuda N, Nakamura M, Aoyama T, Oshima T (2004) Long-term conservation of six duplicated structural genes in cephalopod mitochondrial genomes. Mol Biol Evol 21:2034–2046
Zhang P, Zhou H, Liang D, Liu YF, Chen YQ, Qu LH (2005) The complete mitochondrial genome of a tree frog, Polypedates megacephalus (Amphibia: Anura: Rhacophoridae), and a novel gene organization in living amphibians. Gene 14:133–143
Acknowledgments
This work was supported by the Italian PNRA (grant to T.P.) and by the Ministerio de Educación y Ciencia of Spain (grant CGL2004-00401 to R.Z.). The authors would like to thank Rafaella Franch, Gabriella Mazzotta, Enrico Negrisolo, Lorenzo Zane and Vittorio Varotto for technical support, samples and advice, as well as Soraya Villalba for providing the adapted outline drawings in Fig. 1.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Papetti, C., Liò, P., Rüber, L. et al. Antarctic Fish Mitochondrial Genomes Lack ND6 Gene. J Mol Evol 65, 519–528 (2007). https://doi.org/10.1007/s00239-007-9030-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-007-9030-z