Abstract
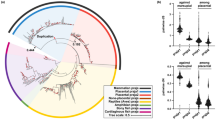
Guanine deaminase (GDA; guanase) is a ubiquitous enzyme that catalyzes the first step of purine metabolism by hydrolytic deamination of guanine, resulting in the production of xanthine. This hydrolase subfamily member plays an essential role in maintaining homeostasis of cellular triphosphate nucleotides for energy, signal transduction pathways, and nitrogen sources. In mammals, GDA protein levels can play a role in neuronal development by regulating dendritic arborization. We previously demonstrated that the most abundant alternative splice form of GDA in mammals, termed cypin (cytosolic PSD-95 interactor), interacts with postsynaptic density proteins, regulates microtubule polymerization, and increases dendrite number. Since purine metabolism and dendrite development were previously thought to be independent cellular processes, this multifunctional protein serves as a new target for the treatment of cognitive disorders characterized by aberrant neuronal morphology and purine metabolism. Although the enzymatic activity of GDA has been conserved during evolution from prokaryotes to higher eukaryotes, a detailed evolutionary assessment of the principal domains in GDA proteins has not yet been put forward. In this study, we perform a complete evolutionary analysis of the full-length sequences and the principal domains in guanine deaminases. Furthermore, we reconstruct the molecular phylogeny of guanine deaminases with neighbor-joining, maximum-likelihood, and UPGMA methods of phylogenetic inference. This study can act as a model whereby a universal housekeeping enzyme may be adapted to act also as a key regulator of a developmental process.







Similar content being viewed by others
References
Akum BF, Chen M, Gunderson SI, Riefler GM, Scerri-Hansen MM, Firestein BL (2004) Cypin regulates dendrite patterning in hippocampal neurons by promoting microtubule assembly. Nat Neurosci 7:145–152
Bergthorsson U, Andersson DI, Roth JR (2007) Ohno’s dilemma: evolution of new genes under continuous selection. Proc Natl Acad Sci USA 104:17004–17009
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Charych EI, Akum BF, Goldberg JS, Jörnsten RJ, Rongo C, Zheng JQ, Firestein BL (2006) Activity-independent regulation of dendrite patterning by postsynaptic density protein PSD-95. J Neurosci 26(40):10164-10176
Chen H, Firestein BL (2007) RhoA regulates dendrite branching in hippocampal neurons by decreasing cypin protein levels. J Neurosci 27:8378–8386
Chen M, Lucas KG, Akum BF, Balasingam G, Stawicki TM, Provost JM, Riefler GM, Jornsten RJ, Firestein BL (2005) A novel role for snapin in dendrite patterning: interaction with cypin. Mol Biol Cell 16:5103–5114
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fernandez JR, Welsh WJ, Firestein BL (2008) Structural characterization of the zinc binding domain in cytosolic PSD-95 interactor (cypin): role of zinc binding in guanine deamination and dendrite branching. Proteins 70:873–881
Firestein BL, Brenman JE, Aoki C, Sanchez-Perez AM, El-Husseini AE, Bredt DS (1999) Cypin: a cytosolic regulator of PSD-95 postsynaptic targeting. Neuron 24:659–672
Fogle PJ, Bieber AL (1975) Purification of rabbit liver guanine aminohydrolase. Prep Biochem 5:59–77
Galilea JM, Canela EI, Bozal J (1981) Characterization of bovine liver guanine aminohydrolase. Int J Biochem 13:773–776
Haldane JBS (1932) The causes of evolution. Cornell University Press, Ithaca, NY
Kimura M, Ota T (1974) On some principles governing molecular evolution. Proc Natl Acad Sci USA 71:2848–2852
Kumar S, Tewari KK, Krishnan PS (1965) Guanine-deaminase activity in rat brain and liver. Biochem J 95:797–802
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) ClustalW and ClustalX version 2. Bioinformatics 23:2947–2948
Lee SF, Kelly M, McAlister A, Luck SN, Garcia EL, Hall RA, Robins-Browne RM, Frankel G, Hartland EL (2008) A C-terminal class I PDZ binding motif of EspI/NleA modulates the virulence of attaching and effacing Escherichia coli and Citrobacter rodentium. Cell Microbiol 10(2):499–513
Liaw SH, Chang YJ, Lai CT, Chang HC, Chang GG (2004) Crystal structure of Bacillus subtilis guanine deaminase: the first domain-swapped structure in the cytidine deaminase superfamily. J Biol Chem 279:35479–35485
Maynes JT, Yuan RG, Snyder FF (2000) Identification, expression, and characterization of Escherichia coli guanine deaminase. J Bacteriol 182:4658–4660
Nygaard P, Bested SM, Andersen KA, Saxild HH (2000) Bacillus subtilis guanine deaminase is encoded by the yknA gene and is induced during growth with purines as the nitrogen source. Microbiology 146(Pt 12):3061–3069
Muller HJ (1936) Bar duplication. Science 83:528–530
Paletzki RF (2002) Cloning and characterization of guanine deaminase from mouse and rat brain. Neuroscience 109:15–26
Ponting CP, Phillips C, Davies KE, Blake DJ (1997) PDZ domains: targeting signalling molecules to sub-membranous sites. Bioessays 19:469–479
Prodanov K, Jerev S (1971) “Discharge” of guanine deaminase from the dog’s lung. Nauchni Tr Vissh Med Inst Sofiia 50:57–60
Rossi CA, Hakim G, Solaini G (1978) Purification and properties of pig brain guanine deaminase. Biochim Biophys Acta 526:235–246
Saint-Marc C, Daignan-Fornier B (2004) GUD1 (YDL238c) encodes Saccharomyces cerevisiae guanine deaminase, an enzyme expressed during post-diauxic growth. Yeast 21:1359–1363
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sheng M, Sala C (2001) PDZ domains and the organization of supramolecular complexes. Annu Rev Neurosci 24:1–29
Sohn J, Grant RA, Sauer RT (2007) Allosteric activation of DegS, a stress sensor PDZ protease. Cell 131(3):572–583
Sokal RR, Sneath PHA (1973) Principles of numerical taxonomy. Freeman, New York
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Yao L, Cukier RI, Yan H (2007) Catalytic mechanism of guanine deaminase: an ONIOM and molecular dynamics study. J Phys Chem B 111:4200–4210
Yuan G, Bin JC, McKay DJ, Snyder FF (1999) Cloning and characterization of human guanine deaminase. Purification and partial amino acid sequence of the mouse protein. J Biol Chem 274:8175–8180
Zuckerkandl E, Pauling L (1965) Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic Press, New York, pp 97–166
Acknowledgments
J.R.F. is a graduate trainee of the IGERT Program on Integratively Engineered Biointerfaces at Rutgers, under NSF Grant DGE-0333196. This work was supported in part by a Johnson and Johnson Discovery Grant, a New Jersey Governor’s Council on Autism Pilot Grant, National Science Foundation Grants IBN-0234206 and IBN-0548543, and March of Dimes Foundation Grant 1-FY04-107 (to B.L.F). We thank Dr. Chi-hua Chiu for comments on and critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fernández, J.R., Byrne, B. & Firestein, B.L. Phylogenetic Analysis and Molecular Evolution of Guanine Deaminases: From Guanine to Dendrites. J Mol Evol 68, 227–235 (2009). https://doi.org/10.1007/s00239-009-9205-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9205-x