Abstract
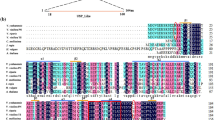
Dehydrins are proteins that accumulate in vegetative tissues subjected to various dehydrating stress conditions such as cold, drought, and salinity and in seeds at later stages of embryogenesis. Here, we report on two highly identical dehydrin genes, DHN1a and DHN1b, in wild and cultivated grapes, Vitis riparia and Vitis vinifera, and their expression in different tissues and under different environmental conditions. The two genes and their transcripts can easily be distinguished by RT-PCR because DHN1b has an 18 bp deletion compared to DHN1a. V. riparia expressed only DHN1a; V. vinifera expressed both DHN1a and DHN1b. Spliced transcripts, DHN1-S, encoding a putative YSK2-type dehydrin were present in low amounts in control leaves, but in high amounts in buds and seeds. Unspliced transcripts, DHN1-U, accumulated to high levels in buds and seeds. Cold, drought, and ABA treatment increased accumulation of both DHN1-S and DHN1-U in leaves, whereas short-day treatment increased only DHN1-S. The possible relation of these results with the difference in freezing stress tolerance between V. riparia and V. vinifera is discussed.





Similar content being viewed by others
Abbreviations
- DHN:
-
dehydrin
- DHN-S:
-
spliced dehydrin transcripts
- DHN-U:
-
unspliced dehydrin transcripts
- iPCR:
-
inverse polymerase chain reaction
- ORF:
-
open reading frame
- Vr:
-
Vitis riparia
- Vv:
-
Vitis vinifera
References
Allagulova CR, Gimalov FR, Shakirova FM, Vakhitov VA (2003) The plant dehydrins: structure and putative functions. Biochemistry (Moscow) 68:945–951
Alsheikh MK, Heyen BJ, Randall SK (2003) Ion binding properties of the dehydrin ERD14 are dependent upon phosphorylation. J Biol Chem 278:40882–40889
Asghar R, Fenton RD, DeMason DA, Close TJ (1994) Nuclear and cytoplasmic localization of maize embryo and aleurone dehydrin. Protoplasma 177:87–94
Borovskii GB, Stupnikova IV, Antipina AI, Vladimirova SV, Voinikov VK (2002) Accumulation of dehydrin-like proteins in the mitochondria of cereals in response to cold, freezing, drought and ABA treatment. BMC Plant Biol [electronic resource] 2:5
Bournay AS, Hedley PE, Maddison A, Waugh R, Machray GC (1996) Exon skipping induced by cold stress in a potato invertase gene transcript. Nucleic Acids Res 24:2347–2351
Bravo LA, Gallardo J, Navarrete A, Olave N, Martinez J, Alberdi M, Close TJ, Corcuea LJ (2003) Cryoprotective activity of a cold-induced dehydrin purified from barley. Physiol Plant 118:262–269
Brown JWS, Simpson CG (1998) Splice site selection in plant pre-mRNA splicing. Annu Rev Plant Physiol Plant Mol Biol 49:77–95
Choi DW, Close TJ (2000) A newly identified barley gene, Dhn12, encoding a YSK2 DHN, is located on chromosome 6H and has embryo-specific expression. Theor Appl Genet 100:1274–1278
Choi DW, Zhu B, Close TJ (1999) The barley (Hordeum vulgare L.) dehydrin family: sequences, allele types, chromosome assignments, and expression characteristics of 11 Dhn genes of cv Dicktoo. Theor Appl Genet 98:1234–1247
Close TJ (1996) Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiol Plant 97:795–803
Close TJ (1997) Dehydrins: a commonality in the response of plants to dehydration and low temperature. Physiol Plant 100:291–296
Close TJ, Fenton RD, Moonan F (1993) A view of plant dehydrins using antibodies specific to the carboxy terminal peptide. Plant Mol Biol 23:279–286
Danyluk J, Perron A, Houde M, Limin A, Fowler B, Benhamou N, Sarhan F (1998) Accumulation of an acidic dehydrin in the vicinity of the plasma membrane during cold acclimation of wheat. Plant Cell 10:623–638
Dhanaraj AL, Slovin JP, Rowland LJ (2005) Isolation of a cDNA clone and characterization of expression of the highly abundant, cold acclimation-associated 14 kDA dehydrin of blueberry. Plant Sci 168:949–957
Goday A, Jensen AB, Culianez-macia FA, Alba MM, Figueras M, Serratosa J, Torrent M, Pages M (1994) The maize abscisic acid-response protein Rab17 is located in the nucleus and interacts with nuclear localization signals. Plant Cell 6:351–360
Godoy J, Lunar R, Torres-Schumann S, Moreno J, Rodrigo RM, Pintor-Toro JA (1994) Expression, tissue distribution and subcellular localization of dehydrin TAS14 in salt-stressed tomato plants. Plant Mol Biol 26:1921–1934
Hara M, Terashima S, Kuboi T (2001) Characterization and cryoprotective activity of cold-responsive dehydrin from Citrus unshiu. J Plant Physiol 158:1333–1339
Hara M, Terashima S, Fukaya T, Kuboi T (2003) Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta 17:290–298
Hara M, Fujinaga M, Kuboi T (2004) Radical scavenging activity and oxidative modification of citrus dehydrin. Plant Phys Biochem 42:657–662
Hennig L (1999) WinGen/WinPep. User-friendly software for the analysis of amino acid sequences. Biotechniques 16:1170–1172
Heyen BJ, Alsheikh MK, Smith EA, Torvik CF, Seals DF, Randall SK (2002) The calcium-binding activity of a vacuole-associated, dehydrin-like protein is regulated by phosphorylation. Plant Physiol 130:675–687
Houde M, Daniel C, Lachapelle M, Allard F, Laliberte S, Sarhan F (1995) Immunolocalization of freezing-tolerance-associated proteins in the cytoplasm and nucleoplasm of wheat crown tissues. Plant J 8:583–593
Ismail AM, Hall AE, Close TJ (1999a) Purification and partial characterization of a dehydrin involved in chilling tolerance during seedling emergence of cowpea. Plant Physiol 120:237–244
Ismail AM, Hall AE, Close TJ (1999b) Allelic variation of a dehydrin gene cosegregates with chilling tolerance during seedling emergence. Proc Natl Aca Sci USA 96:13566–13570
Jia Y, del Rio HS, Robbins AL, Louzada ES (2004) Cloning and sequence analysis of a low temperature-induced gene from trifoliate orange with unusual pre-mRNA processing. Plant Cell Rep 23:159–166
Karlson DT, Zeng Y, Stirm VE, Joly RJ, Ashworth EN (2003) Photoperiodic regulation of a 24-kD dehydrin-like protein in red-osier dogwood (Cornus sericea L.) in relation to freeze-tolerance. Plant Cell Physiol 44:25–34
Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2004) A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low-temperature stress tolerance in tobacco by gene transfer. Plant Cell Physiol 45:346–350
Kazan T (2003) Alternative splicing and proteome diversity in plants: the tip of the iceberg has just emerged. Trends Plant Sci 8:468–471
Kazuoka T, Oeda K (1994) Purification and characterization of COR85-oligomeric complex from cold-acclimated spinach. Plant Cell Physiol 35:601–611
Kikuchi S, Satoh K, Nagata T, Kawagashira N, Doi K, Kishimoto N, Yazaki J, Ishikawa M, Yamada H, Ooka H et al Rice Full-Length cDNA Consortium; National Institute of Agrobiological Sciences Rice Full-Length cDNA Project Team; Foundation of Advancement of International Science Genome Sequencing & Analysis Group; RIKEN (2003) Collection, mapping, and annotation of over 28 000 cDNA clones from japonica rice. Science 301:376–379
Koag MC, Fenton RD, Stephan Wilkens S, Close TJ (2003) The binding of maize DHN1 to lipid vesicles. Gain of structure and lipid specificity. Plant Physiol 131:309–316
Kong J, Gong JM, Zhang ZG, Zhang JS, Chen SY (2003) A new AOX homologous gene OsIM1 from rice (Oryza sativa L.) with an alternative splicing mechanism under salt stress. Theor Appl Genel 107:326–331
Kruger C, Berkowith O, Stephan UW, Hell R (2002) A metal-binding member of the late embryogenesis abundant protein family transports iron in the phloem of Ricinus communis L. J Biol Chem 277:25062–25062
Levi A, Panta GR, Parmentier CM, Muthalif MM, Arora R, Shanker S, Rowland LJ (1999) Complementary DNA cloning, sequencing and expression of an unusual dehydrin from blueberry floral buds. Physiol Plant 107:98–109
Li L, Howe GA (2001) Alternative splicing of prosystemin pre-mRNA produces two isoforms that are active as signals in the wound response pathway. Plant Mol Biol 46:409–419
Lida K, Seki M, Sakurai T, Satou M, Akiyama K, Toyoda T, Konagaya A, Shinozaki K (2004) Genome-wide analysis of alternative pre-mRNA splicing in Arabidopsis thaliana based on full-length cDNA sequences. Nucleic Acids Res 32:5096–5103
Marrs KA, Walbot V (1997) Expression and RNA splicing of the maize glutathione S-transferase Bronze2 gene is regulated by cadmium and other stresses. Plant Physiol 113:93–102
Modrek B, Resch A, Grasso C, Lee C (2001) Genome-wide detection of alternative splicing in expressed sequences of human genes. Nucleic Acids Res 29:2850–2859
Mueller JK, Heckathorn SA, Fernando D (2003) Identification of a chloroplast dehydrin in leaves of mature plants. Int J Plant Sci 164:535–542
Nassuth A, Pollari E, Helmeczy K, Stewart S, Kofalvi S (2000) Improved RNA extraction and one-tube RT-PCR assay for the simultaneous detection of control plant RNA plus several viruses in plant extracts. J Virol Methods 90:37–49
Ner-Gaon H, Halachmi R, Savaldi-Goldstein S, Rubin E, Ophir R, Fuhr R (2004) Intron retention is a major phenomenon in alternative splicing in Arabidopsis. Plant J 39:877–886
Nylander M, Svensson J, Palva ET, Welin BV (2001) Stress-induced accumulation and tissue-specific localization of dehydrins in Arabidopsis thaliana. Plant Mol Biol 45:263–279
Okazaki Y, Furuno M, Kasukawa T, Adachi J, Bono H, Kondo S, Nikaido I, Osato N, Saito R, Suzuki H et al (2002) Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420:563–573
Parra MM, del Pozo O, Luna R, Godoy JA, Pintor-Toro JA (1996) Structure of the dehydrin tas14 gene of tomato and its developmental and environmental regulation in transgenic tobacco. Plant Mol Biol 32:453–460
Puhakainen T, Hess MW, Makela P, Svensson J, Heino P, Palva ET (2004) Overexpression of multiple dehydrin genes enhances tolerance to freezing in Arabidopsis. Plant Mol Biol 54:743–753
Rinne PLH, Kaikuranta PLM, van der Plas LHW, van der Schoot C (1999) Dehydrins in cold-acclimated apices of birch (Betula pubescens Ehrh.): production, localization and potential role in rescuing enzyme function during dehydration. Planta 209:377–388
Robertson M, Chandler PM (1994) A dehydrin cognate protein from pea (Pisum sativum L.) with an atypical pattern of expression. Plant Mol Biol 26:805–816
Rorat T, Grygorowicz WJ, Irzykowski W, Rey P (2004) Expression of KS-type dehydrins is primarily regulated by factors related to organ type and leaf developmental stage during vegetative growth. Planta 218:878–885
Sambrook J, Russell DN (2001) Molecular cloning. A laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York.
Sarry JE, Sommerer N, Sauvage F-X, Bergoin A, Rossignol M, Albagnac G, Romieu C (2004) Grape berry biochemistry revisited upon proteomic analysis of the mesocarp. Proteomics 4:201–215
Shi H, Xiong L, Stevenson B, Lu T, Zhu JK (2002) The Arabidopsis salt overly sensitive 4 mutants uncover a critical role for Vitamin B6 in plant salt tolerance. Plant Cell 14:575–588
Svensson J, Ismail AM, Palva ET, Close TJ (2002) Dehydrins. In: Storey KB, Storey JM (eds) Sensing, signaling and cell adaptation. Elsevier Science B.V., Amsterdam, pp 15–171
Wake CMF, Fennell A (2000) Morphological, physiological and dormancy responses of three Vitis genotypes to short photoperiod. Physiol Plant 109:203–210
Welin BV, Olson A, Nylander M, Tapio Palva E (1994) Characterization and differential expression of dhn/lea/rab-like genes during cold acclimation and drought stress in Arabidopsis thaliana. Plant Mol Biol 26:131–144
Welling A, Rinne P, Vihera-Aarnio A, Kontunen-Soppela S, Heino P, Palva ET (2004) Photoperiod and temperature differentially regulate the expression of two dehydrin genes during overwintering birch (Betula pubescens Ehrh.). J Exp Bot 55:507–516
Wisniewski M, Webb R, Balsamo R, Close TJ, Yu XM, Griffith M (1999) Purification, immunolocalization, cryoprotective, and antifreeze activity of PCA60: A dehydrin from peach (Prunus persica). Physiol Plant 105:600–608
Xu DP, Duan XL, Wang BY, Hong BM, Ho THD, Wu R (1996) Expression of a late embryogenesis abundant protein gene, HAV1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110:249–257
Zhou Y, Zhou C, Ye L, Dong J, Xu H, Cai L, Zhang L, Wei L (2003) Database and analyses of known alternatively spliced genes in plants. Genomics 82:584–595
Acknowledgements
The research was financially supported by Chateau des Charmes Wineries, St. Catherine, Ontario, and the Food Science Biotechnology Centre at the University of Guelph.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. C. Register
Rights and permissions
About this article
Cite this article
Xiao, H., Nassuth, A. Stress- and development-induced expression of spliced and unspliced transcripts from two highly similar dehydrin 1 genes in V. riparia and V. vinifera . Plant Cell Rep 25, 968–977 (2006). https://doi.org/10.1007/s00299-006-0151-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-006-0151-4