Abstract
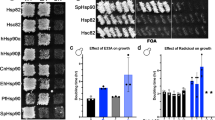
The Hsp60 and Hsp70 chaperones contain a number of conserved inserts that are restricted to particular phyla of bacteria. A one aa insert in the E. coli GroEL and a 21–23 insert in the DnaK proteins are specific for most Gram-negative bacteria. Two other inserts in DnaK are limited to certain groups of proteobacteria. The requirement of these inserts for cellular growth was examined by carrying out complementation studies with temperature-sensitive (T s) mutants of E. coli groEL or dnaK. Our results demonstrate that deletion or most changes in these inserts completely abolished the complementation ability of the mutant proteins. Studies with GroEL and DnaK from some other species that either lacked or contained these inserts also indicated that these inserts are essential for growth of E. coli. The DnaK from some bacteria contains a two aa insert that is not found in E. coli. Introduction of this insert into the E. coli DnaK also led to its inactivation, indicating that these inserts are specific for different groups. We postulate that these conserved inserts that are localized in loop regions on protein surfaces, are involved in some ancillary functions that are essential for the groups of bacteria where they are found.







Similar content being viewed by others
References
Ahmad S, Ahuja R, Venner TJ, Gupta RS (1990) Identification of a protein altered in mutants resistant to microtubule inhibitors as a member of the major heat shock protein (hsp70) family. Mol Cell Biol 10:5160–5165
Ang D, Georgopoulos C (1989) The heat-shock-regulated grpE gene of Escherichia coli is required for bacterial growth at all temperatures but is dispensable in certain mutant backgrounds. J Bacteriol 171:2748–2755
Arifuzzaman M, Oshima T, Mori H (2004) The ATPase domain of HscC (DnaK homolog) is essential for interfering sigma70 activity in E. coli. FEMS Microbiol Lett 230:99–104
Artsimovitch I, Svetlov V, Murakami KS, Landick R (2003) Co-overexpression of Escherichia coli RNA polymerase subunits allows isolation and analysis of mutant enzymes lacking lineage-specific sequence insertions. J Biol Chem 278:12344–12355
Baldauf SL, Palmer JD (1993) Animals and fungi are each other’s closest relatives: congruent evidence from multiple proteins. Proc Natl Acad Sci USA 90:11558–11562
Bork P, Koonin EV (1998) Predicting functions from protein sequences—where are the bottlenecks. Nat Genet 18:313–318
Braig K, Otwinowski Z, Hegde R, Boisvert DC, Joachimiak A, Horwich AL, Sigler PB (1994) The crystal structure of the bacterial chaperonin GroEL at 2.8 A. Nature 371:578–586
Bukau B, Horwich AL (1998) The Hsp70 and Hsp60 chaperone machines. Cell 92:351–366
Bukau B, Walker GC (1989) Cellular defects caused by deletion of the Escherichia coli dnaK gene indicate roles for heat shock protein in normal metabolism. J Bacteriol 171:2337–2346
Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P (2006) Toward automatic reconstruction of a highly resolved tree of life. Science 311:1283–1287
Craig EA, Gambill BD, Nelson RJ (1993) Heat shock proteins: molecular chaperones of protein biogenesis. Microbiol Rev 57:402–414
Danchin A (1999) From protein sequence to function. Curr Opin Struct Biol 9:363–367
Daubin V, Ochman H (2004) Bacterial genomes as new gene homes: the genealogy of ORFans in E. coli. Genome Res 14:1036–1042
Doolittle WF (1999) Phylogenetic classification and the universal tree. Science 284:2124–2128
Ellis RJ, Hartl FU (1999) Principles of protein folding in the cellular environment. Curr Opin Struct Biol 9:102–110
Embley TM, Stackebrandt E (1993) The molecular phylogeny and systematics of the actinomycetes. Annu Rev Microbiol 48:257–289
Fayet O, Ziegelhoffer T, Georgopoulos C (1989) The groES and groEL heat shock gene products of Escherichia coli are essential for bacterial growth at all temperatures. J Bacteriol 171:1379–1385
Fenton WA, Kashi Y, Furtak K, Horwich AL (1994) Residues in chaperonin GroEL required for polypeptide binding and release. Nature 371:614–619
Flaherty KM, DeLuca-Flaherty C, McKay DB (1990) Three-dimensional structure of the ATPase fragment of a 70 K heat-shock cognate protein. Nature 346:623–628
Galperin MY, Koonin EV (2004) ‘Conserved hypothetical’ proteins: prioritization of targets for experimental study. Nucleic Acids Res 32:5452–5463
Gao B, Gupta RS (2005) Conserved indels in protein sequences that are characteristic of the phylum Actinobacteria. Int J Syst Evol Microbiol 55:2401–2412
Georgopoulos CP, Hendrix RW, Casjens SR, Kaiser AD (1973) Host participation in bacteriophage lambda head assembly. J Mol Biol 76:45–60
Gogarten JP, Doolittle WF, Lawrence JG (2002) Prokaryotic evolution in light of gene transfer. Mol Biol Evol 19:2226–2238
Gray MW (1999) Evolution of organellar genomes. Curr Opin Genet Dev 9:678–687
Griffiths E, Gupta RS (2004) Signature sequences in diverse proteins provide evidence for the late divergence of the order Aquificales. Int Microbiol 7:41–52
Griffiths E, Petrich A, Gupta RS (2005) Conserved indels in essential proteins that are distinctive characteristics of Chlamydiales and provide novel means for their identification. Microbiology 151:2647–2657
Gupta RS (1995) Evolution of the chaperonin families (Hsp60, Hsp10 and Tcp-1) of proteins and the origin of eukaryotic cells. Mol Microbiol 15:1–11
Gupta RS (1998) Protein phylogenies and signature sequences: a reappraisal of evolutionary relationships among archaebacteria, eubacteria, and eukaryotes. Microbiol Mol Biol Rev 62:1435–1491
Gupta RS (2000) The phylogeny of proteobacteria: relationships to other eubacterial phyla and eukaryotes. FEMS Microbiol Rev 24:367–402
Gupta RS (2004) The Phylogeny and signature sequences characteristics of Fibrobacters, Chlorobi and Bacteroidetes. Crit Rev Microbiol 30:123–143
Gupta RS, Golding GB (1993) Evolution of HSP70 gene and its implications regarding relationships between archaebacteria, eubacteria, and eukaryotes. J Mol Evol 37:573–582
Gupta RS, Griffiths E (2006) Chlamydiae-specific proteins and indels: novel tools for studies. Trends Microbiol 14:527–535
Gupta RS, Mok A (2007) Phylogenomics and signature proteins for the alpha proteobacteria and its main groups. BMC Microbiol 7:106
Gupta RS, Singh B (1994) Phylogenetic analysis of 70 kD heat shock protein sequences suggests a chimeric origin for the eukaryotic cell nucleus. Curr Biol 4:1104–1114
Gupta RS, Aitken K, Falah M, Singh B (1994) Cloning of Giardia lamblia heat shock protein HSP70 homologs: implications regarding origin of eukaryotic cells and of endoplasmic reticulum. Proc Natl Acad Sci USA 91:2895–2899
Gupta RS, Pereira M, Chandrasekera C, Johari V (2003) Molecular signatures in protein sequences that are characteristic of cyanobacteria and plastid homologues. Int J Syst Evol Microbiol 53:1833–1842
Harrison CJ, Hayer-Hartl M, Di Liberto M, Hartl F, Kuriyan J (1997) Crystal structure of the nucleotide exchange factor GrpE bound to the ATPase domain of the molecular chaperone DnaK. Science 276:431–435
Jindal S, Dudani AK, Singh B, Harley CB, Gupta RS (1989) Primary structure of a human mitochondrial protein homologous to the bacterial and plant chaperonins and to the 65-kilodalton mycobacterial antigen. Mol Cell Biol 9:2279–2283
Klein G, Georgopoulos C (2001) Identification of important amino acid residues that modulate binding of Escherichia coli GroEL to its various cochaperones. Genetics 158:507–517
Lake JA, Herbold CW, Rivera MC, Servin JA, Skophammer RG (2007) Rooting the tree of life using nonubiquitous genes. Mol Biol Evol 24:130–136
Macario AJL, Lange M, Ahring BK, De Macario EC (1999) Stress genes and proteins in the archaea. MMBR 63:923–967
Margulis L (1993) Symbiosis in cell evolution. W.H. Freeman, New York, p 452
Opalka N, Mooney RA, Richter C, Severinov K, Landick R, Darst SA (2000) Direct localization of a beta-subunit domain on the three-dimensional structure of Escherichia coli RNA polymerase. Proc Natl Acad Sci USA 97:617–622
Palmer JD, Delwiche CF (1998) The origin and evolution of plastids and their genomes. In: Sotis DE, Soltis PE, Doyle JJ (eds) Molecular systematics of plants II DNA sequencing. Kluwer, Norwell, pp 375–409
Pellecchia M, Montgomery DL, Stevens SY, Vander Kooi CW, Feng HP, Gierasch LM, Zuiderweg ER (2000) Structural insights into substrate binding by the molecular chaperone DnaK. Nat Struct Biol 7:298–303
Picketts DJ, Mayanil CS, Gupta RS (1989) Molecular cloning of a Chinese hamster mitochondrial protein related to the “chaperonin” family of bacterial and plant proteins. J Biol Chem 264:12001–12008
Popp S, Packschies L, Radzwill N, Vogel KP, Steinhoff HJ, Reinstein J (2005) Structural dynamics of the DnaK–peptide complex. J Mol Biol 347:1039–1052
Richardson A, Schwager F, Landry SJ, Georgopoulos C (2001) The importance of a mobile loop in regulating chaperonin/co-chaperonin interaction: humans versus Escherichia coli. J Biol Chem 276:4981–4987
Rist W, Graf C, Bukau B, Mayer MP (2006) Amide hydrogen exchange reveals conformational changes in hsp70 chaperones important for allosteric regulation. J Biol Chem 281:16493–16501
Rivera MC, Lake JA (1992) Evidence that eukaryotes and eocyte prokaryotes are immediate relatives. Science 257:74–76
Rodriguez-Quinones F, Maguire M, Wallington EJ, Gould PS, Yerko V, Downie JA, Lund PA (2005) Two of the three groEL homologues in Rhizobium leguminosarum are dispensable for normal growth. Arch Microbiol 183:253–265
Rokas A, Holland PW (2000) Rare genomic changes as a tool for phylogenetics. Trends Ecol Evol 15:454–459
Rusanganwa E, Gupta RS (1993) Cloning and characterization of multiple groEL chaperonin-encoding genes in Rhizobium meliloti. Gene 126:67–75
Sugimoto S, Higashi C, Saruwatari K, Nakayama J, Sonomoto K (2007) A gram-negative characteristic segment in Escherichia coli DnaK is essential for the ATP-dependent cooperative function with the co-chaperones DnaJ and GrpE. FEBS Lett 581:2993–2999
Suppini JP, Amor M, Alix JH, Ladjimi MM (2004) Complementation of an Escherichia coli DnaK defect by Hsc70-DnaK chimeric proteins. J Bacteriol 186:6248–6253
Sussman MD, Setlow P (1987) Nucleotide sequence of a Bacillus megaterium gene homologous to the dnaK gene of Escherichia coli. Nucleic Acids Res 15:3923
Tilly K, Hauser R, Campbell J, Ostheimer GJ (1993) Isolation of dnaJ, dnaK, and grpE homologues from Borrelia burgdorferi and complementation of Escherichia coli mutants. Mol Microbiol 7:359–369
Ventura M, Canchaya C, Tauch A, Chandra G, Fitzgerald GF, Chater KF, van Sinderen D (2007) Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol Mol Biol Rev 71:495–548
Viale AM, Arakaki AK, Soncini FC, Ferreyra RG (1994) Evolutionary relationships among eubacterial groups as inferred from GroEL (chaperonin) sequence comparisons. Int J Syst Bacteriol 44:527–533
Vickery LE, Cupp-Vickery JR (2007) Molecular chaperones HscA/Ssq1 and HscB/Jac1 and their roles in iron–sulfur protein maturation. Crit Rev Biochem Mol Biol 42:95–111
Wild J, Kamath-Loeb A, Ziegelhoffer E, Lonetto M, Kawasaki Y, Gross CA (1992) Partial loss of function mutations in DnaK, the Escherichia coli homologue of the 70-kDa heat shock proteins, affect highly conserved amino acids implicated in ATP binding and hydrolysis. Proc Natl Acad Sci USA 89:7139–7143
Zeilstra-Ryalls J, Fayet O, Georgopoulos C (1991) The universally conserved GroE (Hsp60) chaperonins. Annu Rev Microbiol 45:301–325
Acknowledgments
We are grateful to Drs. Costa Georgopoulos and Debbie Ang for sending us the T s mutant strains for groEL and dnaK that were used in these studies. We also thank Drs. Lori Burrows and Dr. T. Finan for the DNA from P. aeruginosa and S. meliloti, respectively. This work was supported by a research grant from the Canadian Institute of Health Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by D. Andersson.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Singh, B., Gupta, R.S. Conserved inserts in the Hsp60 (GroEL) and Hsp70 (DnaK) proteins are essential for cellular growth. Mol Genet Genomics 281, 361–373 (2009). https://doi.org/10.1007/s00438-008-0417-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-008-0417-3