Abstract
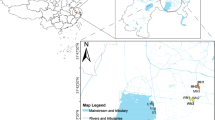
Samples of sediments and surrounding soda soils (SS) from the extremely saline and alkaline lakes of the Wadi el Natrun in the Libyan Desert, Egypt, were obtained in October 2000. Anaerobic enrichment cultures were grown from these samples, DNA isolated, and the bacterial diversity assessed by 16S rRNA gene clone analysis. Clones derived from lake sediments (LS) most closely matched Clostridium spp., Natronoincola histidinovorans, Halocella cellulolytica, Bacillus spp., and the Cytophaga–Flexibacter–Bacteroides group. Similar clones were identified in the SS, but Bacillus spp. predominated. Many of the clones were most closely related to organisms already identified in alkaline or saline environments. Two genomic DNA libraries were made from the pooled LS enrichments and a single SS-enrichment sample. A novel cellulase activity was identified and characterized in each. The lake cellulase ORF encoded a protein of 1,118 amino acids; BLASTP analysis showed it was most closely related to an endoglucanase from Xanthomonas campestris. The soil-cellulase ORF encoded a protein of 634 amino acids that was most closely related to an endoglucanase from Fibrobacter succinogenes.




Similar content being viewed by others
References
Altschul SF, Hadden TL, Schaffer AA, Zhang J, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Amann R, Ludwig W, Schleifer K (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Duckworth AW, Grant WD, Jones BE, van Steenberg R (1996) Phylogenetic diversity of soda lake alkaliphiles. FEMS Microbiol Ecol 19:181–191
Grant S, Grant WD, Jones BE, Kato C, Li L (1999) Novel archaeal phylotypes from an East African alkaline saltern. Extremophiles 3:139–145
Henne A, Schmitz RA, Bomeke M, Gottschalk G, Daniel R (2000) Screening of environmental DNA libraries for the presence of genes conferring lipolytic activity on Escherichia coli. Appl Environ Microbiol 66:3113–3116
Horikoshi K (1999) Alkaliphiles: some applications of their products for biotechnology. Microbiol Mol Biol Rev 63:735–750
Imhoff JF, Sahl HG, Soliman GSH, Truper HG (1979) The Wadi Natrun: chemical composition and microbial mass developments in alkaline brines of eutrophic desert lakes. Geomicrobiol J 1:219–234
Jones BE, Grant WD, Duckworth AW, Owenson GG (1998) Microbial diversity of soda lakes. Extremophiles 2:191–200
Jukes TH, Cantor CR (1969) Evolution of protein molecules In: Munro HN (ed). Mammalian protein metabolism. Academic Press, New York, pp 21–132
Lu J, NogiY, Takami H (2001) Oceanobacillus iheyensis gen. nov., sp. nov., a deep-sea extremely halotolerant and alkaliphilic species isolated from a depth of 1,050 m on the Iheya Ridge. FEMS Microbiol Lett 205:291–297
Malburg SR, Malburg LM Jr, Liu T, Iyo AH, Forsberg CW (1997) Catalytic properties of the cellulose-binding endoglucanase F from Fibrobacter succinogenes S85. Appl Environ Microbiol 63:2449–2453
Margesin R, Schinner F (2001) Potential of halotolerant and halophilic microorganisms for biotechnology. Extremophiles 5:73–83
Marrs B, Delagrave S, Murphy D (1999) Novel approaches for discovering industrial enzymes. Curr Opin Microbiol 2:241–245
Pfennig N, Lippert KD (1966) Uber das Vitamin B12-Bedurfnis phototropher Schwefelbakterien. Arch Mikrobiol 55:245–256
Rees HC, Grant S, Jones BE, Grant WD, Heaphy S (2003a) Detecting cellulase and esterase activities encoded by novel genes present in environmental DNA libraries. Extremophiles 7:415–421
Rees HC, Jones BE, Grant WD, Heaphy S (2003b) Diversity of Kenyan soda lake alkaliphiles assessed by molecular methods. Extremophiles 8:63–71
Stougaard P, Jorgensen F, Johnsen MG, Hansen OC (2002) Microbial diversity in ikaite tufa columns: an alkaline, cold ecological niche in Greenland. Environ Microbiol 4:487–493
Taher AG (1999) Inland saline lakes of Wadi El Natrun depression, Egypt. Int J Salt Lake Res 8:149–170
Teather RM, Wood PJ (1982) Use of Congo Red-polysaccharide interactions in enumeration and characterisation of cellulolytic bacteria from the bovine rumen. Appl Environ Microbiol 43:777–780
Van de Peer Y, De Wachter R (1994) TREECON: a software package for the construction and drawing of evolutionary trees. Comput Appl Biosci 9:177–182
Acknowledgements
This research was supported by the Russian Academy of Sciences Programme of Molecular and Cell Biology for DS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Horikoshi
Rights and permissions
About this article
Cite this article
Grant, S., Sorokin, D.Y., Grant, W.D. et al. A phylogenetic analysis of Wadi el Natrun soda lake cellulase enrichment cultures and identification of cellulase genes from these cultures. Extremophiles 8, 421–429 (2004). https://doi.org/10.1007/s00792-004-0402-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-004-0402-7