Abstract
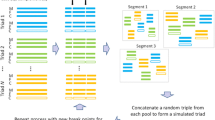
Existing simulation methods usually simulate linkage disequilibrium (LD) structures starting with an initial population that is randomly generated according to specified allele frequencies. These at random based methods might be unstable because the LD level of the initial population is generally extremely low. This study presents a new algorithm, SIMLD, to simulate genome populations with real LD structures. SIMLD begins from an initial population with possibly the highest LD level, and then the LD decays to fit the desired level through processes of mating and recombination over generations. SIMLD can produce case–control samples according to various disease models. Using empirical SNP marker information from three populations of HapMap data, we implement the proposed algorithm and demonstrate a set of experimental results.






Similar content being viewed by others
References
Baker BS, Carpenter ATC, Esposito MS, Esposito RE, Sandler L (1976) The genetic control of meiosis. Annu Rev Genet 10:53–134
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Bass MP, Martin ER, Hauser ER (2004) Pedigree generation for analysis of genetic linkage and association. Pac Symp Biocomput 9:93–103
Chen GK, Marjoram P, Wall JD (2009) Fast and flexible simulation of DNA sequence data. Genome Res 19:136–142
Dudek S, Mostinger AA, Velez D, Williams SM, Ritchie MD (2006) Data simulation software for whole-genome association and other studies in human genetics. Pac Symp Biocomput 11:499–510
Edwards TL, Bush WS, Turner SD, Dudek SM, Tortenson ES, Schmidt M, Martin E, Ritchie MD (2008) Generating linkage disequilibrium patterns in data simulations using GenomeSIMLA. EvoBIO, LNCS 4973:24–35
Hahn LW, Ritchie MD, Moore JH (2003) Multifactor dimensionality reduction software for detecting gene-gene and gene-environment interactions. Bioinformatics 19:376–382
Haldane JBS (1919) The combination of linkage values, and the calculation of distances between the loci of linked factors. J Genet 8:299–309
International HapMap Consortium (2003) The International HapMap Project. Nature 426:789–796
International HapMap Consortium (2005) A haplotype map of the human genome. Nature 437:1299–1320
International HapMap Consortium (2007) A second generation human haplotype map of over 3.1 million SNPs. Nature 449:851–861
Kosambi DD (1944) The estimation of the map distance from recombination values. Ann Eugen 12:172–175
Lewontin RC (1988) On measures of gametic disequilibrium. Genetics 120:849–852
Liang L, Zollner S, Abecasis GR (2007) Genome: a rapid coalescent-based whole genome simulator. Bioinformatics 23:1565–1567
Peng B, Amos CI (2010) Forward-time simulation of realistic samples for genome-wide association studies. BMC Bioinformatics 11:442
Ritchie MD, Hahn LW, Roodi N, Bailey LR, Dupont WD, Parl FF, Moore JH (2001) Multifactor-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am J Hum Genet 69:138–147
Schmidt M, Hauser ER, Martin ER, Schmidt S (2005) Extension of the SIMLA package for generating pedigrees with complex inheritance patterns: environmental covariates, gene-gene and gene-environment interaction. Stat Appl Genet Mol Biol 4, Article 15
Wright FA, Huang H, Guan X, Gamiel K, Jeffries C, Barry WT, de Villena FP, Sullivan PF, Wilhelmsen KC, Zou F (2007) Simulating association studies: a data-based resampling method for candidate regions or whole genome scans. Bioinformatics 23:2581–2588
Acknowledgments
This work was supported in part by the National Science Fund of China under Grant nos. 61070137, 60933009, and 60371044, and by the U.S. National Institutes of Health under Grants GM085665, HL090567, and NS029525.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10528_2011_9416_MOESM1_ESM.doc
Simulated LD for sim500SNPs_data. D′ values by distance are given in the comparisons between the simulated data and real populations, JPT/CHB (a), CEU (b), and YRI (c) (DOC 41 kb)
10528_2011_9416_MOESM2_ESM.doc
Simulated LD for sim1000SNPs_data. D′ values by distance are given in the comparisons of simulated data and real data for populations (a) JPT/CHB, (b) CEU, and (c) YRI (DOC 42 kb)
10528_2011_9416_MOESM3_ESM.doc
Simulated LD for sim2000SNPs_data. D′ values by distance are given in the comparisons of simulated data and real data for populations (a) JPT/CHB, (b) CEU, and (c) YRI (DOC 41 kb)
10528_2011_9416_MOESM4_ESM.doc
Simulated LD for sim5000SNPs_data. D′ values by distance are given in the comparisons of simulated data and real data for populations (a) JPT/CHB, (b) CEU, and (c) YRI (DOC 41 kb)
Rights and permissions
About this article
Cite this article
Yuan, X., Zhang, J. & Wang, Y. Simulating Linkage Disequilibrium Structures in a Human Population for SNP Association Studies. Biochem Genet 49, 395–409 (2011). https://doi.org/10.1007/s10528-011-9416-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-011-9416-x