Abstract
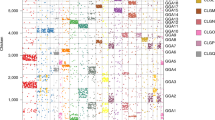
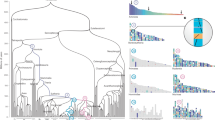
Evidence has shown that bacterial genomes have undergone random shuffling of genomic elements consisting of one to two genes. In order to delineate such genome-shuffling events in mammals, we constructed a high-resolution map of Sus scrofa chromosome 3 (SSC3) with a total of 116 genes/markers. Alignment of this pig map to orthologous regions in human, dog, mouse and rat led to the identification of 31 provisional conserved ancestral blocks (CABs) in these five species. Among them, only 3 CABs (<10%) had one gene, indicating that one-gene shuffling is not frequent in mammals. The sizes of CABs vary significantly within a species, but each may be relatively consistent in different species with a scale to species-genome evolution. The type and frequency of rearrangement events that takes place, either intra- or interchromosomal, depends on the evolutionary regions and species under comparison. Characterization of 36 tentative breakpoint regions flanking these 31 CABs indicated that they occupied ∼43 Mb in length and featured genome deserts, gene duplications, and birth/death of species-specific genes in humans. Identification of CABs provides an alternative for further determination of the evolutionary make-up of mammalian genomes.
Similar content being viewed by others
References
Bellgard MI, Itoh T, Watanabe H, Imanishi T, Gojobori T (1999) Dynamic evolution of genomes and the concept of genome space. Ann NY Acad Sci 870: 293–300.
Boehnke M, Lange K, Cox DR (1991) Statistical methods for multipoint radiation hybrid mapping. Am J Hum Genet 49: 1174–1188.
Crooijmans RP, Dijkhof RJ, Veenendaal T et al. (2001) The gene orders on human chromosome 15 and chicken chromosome 10 reveal multiple inter- and intrachromosomal rearrangements. Mol Biol Evol 18: 2102–2109.
Fan Y, Linardopoulou E, Friedman C, Williams E, Trask BJ (2002a) Genomic structure and evolution of the ancestral chromosome fusion site in 2q13–2q14.1 and paralogous regions on other human chromosomes. Genome Res 12: 1651–1662.
Fan Y, Newman T, Linardopoulou E, Trask BJ (2002b) Gene content and function of the ancestral chromosome fusion site in human chromosome 2q13–2q14.1 and paralogous regions. Genome Res 12: 1663–1672.
Gibbs RA, Weinstock GM, Metzker ML et al. (2004) Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428: 493–521.
Jiang Z, Michal JJ (2003) Linking porcine microsatellite markers to known genome regions by identifying their human orthologs. Genome 46: 798–808.
Jiang Z, Melville JS, Cao H, Kumar S, Filipski A, Gibbins AM (2002a) Measuring conservation of contiguous sets of autosomal markers on bovine and porcine genomes in relation to the map of the human genome. Genome 45: 769–776.
Jiang Z, He H, Hamasima N, Suzuki H, Verrinder G (2002b) Comparative mapping of Homo sapiens chromosome 4 (HSA4) and Sus scrofa chromosome 8 (SSC8) using orthologous genes representing different cytogenetic bands as landmarks. Genome 45: 147–156.
Kirkness EF, Bafna V, Halpern AL et al. (2003) The dog genome: survey sequencing and comparative analysis. Science 301: 1898–1903.
Kumar S, Hedges SB (1998) A molecular timescale for vertebrate evolution. Nature 392: 917–920.
Lander ES, Linton LM, Birren B et al. (2001) Initial sequencing and analysis of the human genome. Nature 409: 860–921.
Lange K, Boehnke M, Cox DR, Lunetta KL (1995) Statistical methods for polyploid radiation hybrid mapping. Genome Res 5: 136–150.
Lunetta KL, Boehnke M, Lange K, Cox DR (1996) Selected locus and multiple panel models for radiation hybrid mapping. Am J Hum Genet 59: 717–725.
Nadeau JH, Sankoff D (1998) Counting on comparative maps. Trends Genet 14: 495–501.
Nadeau JH, Taylor BA (1984) Lengths of chromosomal segments conserved since divergence of man and mouse. Proc Natl Acad Sci USA 81: 814–818.
Nei M, Gu X, Sitnikova T (1997) Evolution by the birth-and-death process in multigene families of the vertebrate immune system. Proc Natl Acad Sci USA 94: 7799–7806.
Pevzner P, Tesler G (2003a) Genome rearrangements in mammalian evolution: lessons from human and mouse genomes. Genome Res 13: 37–45.
Pevzner P, Tesler G (2003b) Human and mouse genomic sequences reveal extensive breakpoint reuse in mammalian evolution. Proc Natl Acad Sci USA 100: 7672–7677.
Rohrer GA, Alexander LJ, Hu Z, Smith TP, Keele JW, Beattie CW (1996) A comprehensive map of the porcine genome. Genome Res 6: 371–391.
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132: 365–386.
Scherer SW, Cheung J, MacDonald JR (2003) Human chromosome 7: DNA sequence and biology. Science 300: 767–772.
Thomas JW, Green ED, NISC Comparative Sequencing Program (2003) Comparative sequence analysis of a single-gene conserved segment in mouse and human. Mamm Genome 14: 673–678.
Venter JC, Adams MD, Myers EW et al. (2001) The sequence of the human genome. Science 291: 1304–1351.
Waterston RH, Lindblad-Toh K, Birney E et al. (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420: 520–562.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, Z., Michal, J.J., Melville, J.S. et al. Multi-alignment of orthologous genome regions in five species provides new insights into the evolutionary make-up of mammalian genomes. Chromosome Res 13, 707–715 (2005). https://doi.org/10.1007/s10577-005-1001-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-005-1001-x