Abstract
Spatial organization of the genome within interphase nuclei is non-random. It has been shown that not only whole chromosomes but also individual genes occupy specific nuclear locations and these locations can be changed during different processes like differentiation or disease. Using a porcine in vitro adipogenesis stem cell differentiation system as a model to study nuclear organization, it was demonstrated that nuclear position of selected genes involved in porcine adipogenesis was altered with the up-regulation of gene expression, correlating with these genes becoming more internally located within nuclei, without whole territory relocation. Here, we investigated whether the gene relocation observed during porcine adipogenesis is related to spatial co-association with SC-35 domains. These domains are nuclear speckles enriched in numerous splicing and RNA metabolic factors. Using a DNA immuno-FISH approach we investigated the localisation of three adipogenic genes (PPARG, SREBF1, and FABP4) with SC-35 domains in porcine mesenchymal stem cells and after they were differentiated into adipocytes. We found that the location of these genes relative to SC-35 domains was non-random and correlated with the up-regulation of gene expression. In addition, we observed more frequent clustering of the studied genes located on different chromosomes around the same nuclear speckle in differentiated adipocytes than in mesenchymal stem cells. However, the choice of the domain was more random. This study adds to the evidence that SC-35 domains are hubs of gene activity and gene-domain association may be considered as a common mechanism to enhance gene expression.




Similar content being viewed by others
Abbreviations
- 3D:
-
Three-dimensional
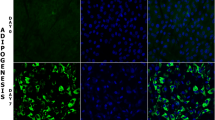
- AD14:
-
Differentiated adipocytes at day 14
- FISH:
-
Fluorescence in situ hybridisation
- MSC:
-
Mesenchymal stem cells
- N2:
-
Nitrogen
- PFA:
-
Paraformaldehyde
- PBS:
-
Phosphate buffered saline
- SSC:
-
Saline sodium citrate buffer
- SSC:
-
Sus scrofa chromosome
References
Brown JM, Green J, das Neves RP, Wallace HA, Smith AJ, Hughes J, Gray N, Taylor S, Wood WG, Higgs DR, Iborra FJ, Buckle VJ (2008) Association between active genes occurs at nuclear speckles and is modulated by chromatin environment. J Cell Biol 82:1083–1097
Dundr M, Ospina JK, Sung MH, John S, Upender M, Ried T, Hager GL, Matera AG (2007) Actin-dependent intranuclear repositioning of an active gene locus in vivo. J Cell Biol 179:1095–1103
Ferrai C, de Castro IJ, Lavitas L, Chotalia M, Pombo A (2010) Gene positioning. Cold Spring Harb Perspect Biol 2:a000588
Finlan LE, Sproul D, Thomson I, Boyle S, Kerr E, Perry P, Ylstra B, Chubb JR, Bickmore WA (2008) Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet 4:e1000039
Hall LL, Smith KP, Byron M, Lawrence JB (2006) Molecular anatomy of a speckle. Anat Rec A Discov Mol Cell Evol Biol 288:664–675
Hu Q, Kwon YS, Nunez E, Cardamone MD, Hutt KR, Ohgi KA, Garcia-Bassets I, Rose DW, Glass CK, Rosenfeld MG, Fu XD (2008) Enhancing nuclear receptor-induced transcription requires nuclear motor and LSD1-dependent gene networking in interchromatin granules. Proc Natl Acad Sci USA 105:19199–19204
Hu Y, Kireev I, Plutz M, Ashourian N, Belmont AS (2009) Large-scale chromatin structure of inducible genes: transcription on a condensed, linear template. J Cell Biol 185:87–100
Johnson CV, Primorac D, McKinstry M, McNeil JA, Rowe D, Lawrence JB (2000) Tracking COL1A1 RNA in osteogenesis imperfecta: splice-defective transcripts initiate transport from the gene but are retained within the SC-35 domain. J Cell Biol 150:417–432
Jolly C, Vourc’h C, Robert-Nicoud M, Morimoto RI (1999) Intron-independent association of splicing factors with active genes. J Cell Biol 145:1133–1143
Kim SH, McQueen PG, Lichtman MK, Shevach EM, Parada LA, Misteli T (2004) Spatial genome organization during T-cell differentiation. Cytogenet Genome Res 105:292–301
Kosak ST, Skok JA, Medina KL, Riblet R, Le Beau MM, Fisher AG, Singh H (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296:158–162
Lamond AI, Spector DL (2003) Nuclear speckles: a model for nuclear organelles. Nat Rev Mol Cell Biol 4:605–612
Meaburn KJ, Misteli T (2008) Locus-specific and activity-independent gene repositioning during early tumorigenesis. J Cell Biol 180:39–50
Mehta IS, Amira M, Harvey AJ, Bridger JM (2010) Rapid chromosome territory relocation by nuclear motor activity in response to serum removal in primary human fibroblasts. Genome Biol 11:R5
Moen PT Jr, Johnson CV, Byron M, Shopland LS, de la Serna IL, Imbalzano AN, Lawrence JB (2004) Repositioning of muscle-specific genes relative to the periphery of SC-35 domains during skeletal myogenesis. Mol Biol Cell 15:197–206
Nielsen JA, Hudson LD, Armstrong RC (2002) Nuclear organization in differentiating oligodendrocytes. J Cell Sci 115:4071–4079
Osborne CS, Chakalova L, Brown KE, Carter D, Horton A, Debrand E, Goyenechea B, Mitchell JA, Lopes S, Reik W, Fraser P (2004) Active genes dynamically colocalize to shared sites of ongoing transcription. Nat Genet 36:1065–1071
Osborne CS, Chakalova L, Mitchell JA, Horton A, Wood AL, Bolland DJ, Corcoran AE, Fraser P (2007) Myc dynamically and preferentially relocates to a transcription factory occupied by Igh. PLoS Biol 5:e192
Ragoczy T, Bender MA, Telling A, Byron R, Groudine M (2006) The locus control region is required for association of the murine beta-globin locus with engaged transcription factories during erythroid maturation. Genes Dev 20:1447–1457
Reddy KL, Zullo JM, Bertolino E, Singh H (2008) Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452:243–247
Shopland LS, Johnson CV, Byron M, McNeil J, Lawrence JB (2003) Clustering of multiple specific genes and gene-rich R-bands around SC-35 domains: evidence for local euchromatic neighborhoods. J Cell Biol 162:981–990
Smith KP, Moen PT, Wydner KL, Coleman JR, Lawrence JB (1999) Processing of endogenous pre-mRNAs in association with SC-35 domains is gene specific. J Cell Biol 144:617–629
Smith KP, Byron M, Johnson C, Xing Y, Lawrence JB (2007) Defining early steps in mRNA transport: mutant mRNA in myotonic dystrophy type I is blocked at entry into SC-35 domains. J Cell Biol 178:951–964
Szczerbal I, Foster HA, Bridger JM (2009) The spatial repositioning of adipogenesis genes is correlated with their expression status in a porcine mesenchymal stem cell adipogenesis model system. Chromosoma 118:647–663
Takizawa T, Gudla PR, Guo L, Lockett S, Misteli T (2008a) Allele-specific nuclear positioning of the monoallelically expressed astrocyte marker GFAP. Genes Dev 22:489–498
Takizawa T, Meaburn KJ, Misteli T (2008b) The meaning of gene positioning. Cell 135:9–13
Williams RR, Azuara V, Perry P, Sauer S, Dvorkina M, Jørgensen H, Roix J, McQueen P, Misteli T, Merkenschlager M, Fisher AG (2006) Neural induction promotes large-scale chromatin reorganisation of the Mash1 locus. J Cell Sci 119:132–140
Xing Y, Johnson CV, Moen PT, McNeil JA, Lawrence J (1995) Nonrandom gene organization: structural arrangements of specific pre-mRNA transcription and splicing with SC-35 domains. J Cell Biol 131:1635–1647
Xu M, Cook PR (2008) Similar active genes cluster in specialized transcription factories. J Cell Biol 181:615–623
Zeng C, Kim E, Warren SL, Berget SM (1997) Dynamic relocation of transcription and splicing factors dependent upon transcriptional activity. EMBO J 16:1401–1412
Zink D, Amaral MD, Englmann A, Lang S, Clarke LA, Rudolph C, Alt F, Luther K, Braz C, Sadoni N, Rosenecker J, Schindelhauer D (2004) Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J Cell Biol 166:815–825
Acknowledgment
This study was financed by the Polish Ministry of Science and Higher Education, grant N301 3381 33.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Pat Heslop-Harrison.
Rights and permissions
About this article
Cite this article
Szczerbal, I., Bridger, J.M. Association of adipogenic genes with SC-35 domains during porcine adipogenesis. Chromosome Res 18, 887–895 (2010). https://doi.org/10.1007/s10577-010-9176-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-010-9176-1