Abstract
Purpose
To catalogue Gene Ontology terms in the sperm of infertile human males vs. donors of proven fertility by analyzing five samples from each of the two groups (five aliquots from fresh sperm and five post-swim-up).
Methods
Microarray technology was employed to study the mRNA profile of both fresh and post-swim-up pooled samples from infertile males and donors.
Results
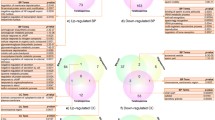
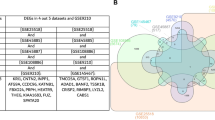
Genes that were differentially expressed in the two populations and expressed in only one of two were analyzed to determine the gene products in terms of their associated Gene Ontology terms. Each group presented a different number and pattern of Gene Ontology terms.
Conclusions
We found differences in Gene Ontology terms between the two groups. These differences could potentially be employed to establish markers of fertility success and to identify cellular processes and complex systems related with male infertility.




Similar content being viewed by others
References
Chatzimeletiou K, Morrison EE, Prapas N, Prapas Y, Handyside AH. The centrosome and early embryogenesis: clinical insights. Reprod Biomed Online. 2008;16:485–91.
Miller D, Ostermeier GC, Krawetz SA. The controversy, potential and roles of spermatozoal RNA. Trends Mol Med. 2005;11:156–63.
Lalancette C, Miller D, Li Y, Krawetz SA. Paternal contributions: new functional insights for spermatozoal RNA. J Cell Biochem. 2008;104:1570–9.
Miller D, Ostermeier GC. Towards a better understanding of RNA carriage by ejaculate spermatozoa. Hum Reprod Updat. 2006;12:757–67.
Miller D, Ostermeier GC. Spermatozoal RNA: why is it there and what does it do? Gynecol Obstet Fertil. 2006;34:840–6.
Boerke A, Dieleman SJ, Gadella BM. A possible role for sperm RNA in early embryo development. Theriogenology. 2007;68 Suppl 1:S147–55.
Garrido N, Martinez-Conejero JA, Jauregui J, Horcajadas JA, Simon C, Remohi J, et al. Microarray analysis in sperm from fertile and infertile men without basic sperm analysis abnormalities reveals a significantly different transcriptome. Fertil Steril. 2009;91:1307–10.
Garrido N, Remohí J, Martinez-Conejero JA, Garcia-Herrero S, Pellicer A, Meseguer M. Contribution of sperm molecular features to embryo quality and assisted reproduction success. Reprod Biomed Online. 2008;17:855–65.
Schena M, Shalon D, Davis RW, Brown PO. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995;270:467–70.
Hoheisel JD. Microarray technology: beyond transcript profiling and genotype analysis. Nat Rev Genet. 2006;7:200–10.
Werner T. Bioinformatics applications for pathway analysis of microarray data. Curr Opin Biotechnol. 2008;19:50–4.
Muriel L, Meseguer M, Fernandez JL, Alvarez J, Remohi J, Pellicer A, et al. Value of the sperm chromatin dispersion test in predicting pregnancy outcome in intrauterine insemination: a blind prospective study. Hum Reprod. 2006;21:738–44.
Martinez-Conejero JA, Garrido N, Remohi J, Pellicer A, Simon C, Meseguer M. MUC1 in human testis and ejaculated spermatozoa and its relationship to male fertility status. Fertil Steril. 2008;90:450–2.
Horcajadas JA, Sharkey AM, Catalano RD, Sherwin JR, Dominguez F, Burgos LA, et al. Effect of an intrauterine device on the gene expression profile of the endometrium. J Clin Endocrinol Metab. 2006;91:3199–207.
Peng X, Wood CL, Blalock EM, Chen KC, Landfield PW, Stromberg AJ. Statistical implications of pooling RNA samples for microarray experiments. BMC Bioinformatics. 2003;4:26.
Gene Ontology Consortium. The Gene Ontology (GO) project in 2006. Nucleic Acids Res. 2006;34:D322–6.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9.
Thomas PD, Mi H, Lewis S. Ontology annotation: mapping genomic regions to biological function. Curr Opin Chem Biol. 2007;11:4–11.
da Huang W, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, et al. DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007;35:W169–75.
Ruiz-Pesini E, Diez C, Lapena AC, Perez-Martos A, Montoya J, Alvarez E, et al. Correlation of sperm motility with mitochondrial enzymatic activities. Clin Chem. 1998;44:1616–20.
Erkkila K, Kyttanen S, Wikstrom M, Taari K, Hikim AP, Swerdloff RS, et al. Regulation of human male germ cell death by modulators of ATP production. Am J Physiol Endocrinol Metab. 2006;290:E1145–54.
Miki K. Energy metabolism and sperm function. Soc Reprod Fertil Suppl. 2007;65:309–25.
Sixt SU, Dahlmann B. Extracellular, circulating proteasomes and ubiquitin—incidence and relevance. Biochim Biophys Acta. 2008;1782:817–23.
Sutovsky P, Neuber E, Schatten G. Ubiquitin-dependent sperm quality control mechanism recognizes spermatozoa with DNA defects as revealed by dual ubiquitin-TUNEL assay. Mol Reprod Dev. 2002;61:406–13.
Muratori M, Marchiani S, Criscuoli L, Fuzzi B, Tamburino L, Dabizzi S, et al. Biological meaning of ubiquitination and DNA fragmentation in human spermatozoa. Soc Reprod Fertil Suppl. 2007;63:153–8.
Takubo K, Ohmura M, Azuma M, Nagamatsu G, Yamada W, Arai F, et al. Stem cell defects in ATM-deficient undifferentiated spermatogonia through DNA damage-induced cell-cycle arrest. Cell Stem Cell. 2008;2:170–82.
Acknowledgements
The authors would like to express their gratitude to technical personnel of the Andrology and IVF-Sperm Laboratory for their cooperation in this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Capsule Ontological analysis of genes that are differentially expressed in the sperm of infertile males undergoing assisted reproduction vs. fertile donors reveals that different biological processes, cellular components, molecular functions and metabolic pathways are attributable.
Rights and permissions
About this article
Cite this article
García-Herrero, S., Garrido, N., Martínez-Conejero, J.A. et al. Ontological evaluation of transcriptional differences between sperm of infertile males and fertile donors using microarray analysis. J Assist Reprod Genet 27, 111–120 (2010). https://doi.org/10.1007/s10815-010-9388-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-010-9388-5