Abstract
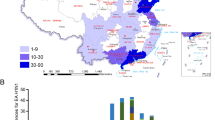
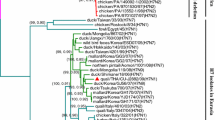
Isolates of the A(H1N1)pdm2009 virus were first identified in asymptomatic swine in Jiangsu province, China in January 2010, indicating that the virus has retro-infected swine after circulating through humans in mainland China. The purpose of this study was to determine whether the avian-origin European H1N1 swine influenza virus (SIV) and the A(H1N1)pdm2009 virus are cocirculating in swine in Jiangsu province of China. From May 2010 to May 2011, 1,030 nasal swab samples were collected from healthy swine in Jiangsu province of China and were tested for influenza A H1N1 using reverse transcription-PCR. Fragments of the complete genomes of viruses from the samples that were positive for influenza A H1N1 were sequenced and analysed. A total of 32 avian-origin European H1N1 SIVs were isolated, and no A(H1N1)pdm2009 viruses were identified; full-length genomes of 18 strains were sequenced. The eight gene segments of some of the isolated H1N1 viruses have 99.1–99.8% sequence identity with the human A/Jiangsu/ALS1/2011(H1N1) isolates in the same region. Our study indicates that the avian-origin European H1N1 SIVs remain endemic in swine and have retro-infected humans after circulating through swine, which may present a risk factor for public health.

Similar content being viewed by others
References
J. Liu et al., Emergence of European avian influenza virus-like H1N1 swine influenza A viruses in China. J. Clin. Microbiol. 47(8), 2643–2646 (2009)
Y. Guan et al., Emergence of avian H1N1 influenza viruses in pigs in China. J. Virol. 70(11), 8041–8046 (1996)
T. Ito et al., Molecular basis for the generation in pigs of influenza A viruses with pandemic potential. J. Virol. 72(9), 7367–7373 (1998)
F.S. Dawood et al., Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N. Engl. J. Med. 360(25), 2605–2615 (2009)
V. Shinde et al., Triple-reassortant swine influenza A (H1) in humans in the United States, 2005–2009. N. Engl. J. Med. 360(25), 2616–2625 (2009)
B. Cao et al., Clinical features of the initial cases of 2009 pandemic influenza A (H1N1) virus infection in China. N. Engl. J. Med. 361(26), 2507–2517 (2009). (2009)
G. Zhao et al., Isolation and phylogenetic analysis of pandemic H1N1/09 influenza virus from swine in Jiangsu province of China. Res. Vet. Sci. (2011) http://dx.doi.org/10.1016/j.rvsc.2011.06.009
World Health Organization. WHO laboratory guidelines for the collection of animal specimens for diagnosis of influenza infection, http://www.who.int/csr/disease/avian_influenza/guidelines/animalspecimens/zh/index.html
H.K. Chang et al., Development of multiplex RT-PCR assays for rapid detection and subtyping of influenza type A viruses from clinical specimens. J. Microbiol. Biotechnol. 18(6), 1164–1169 (2008)
B.F. Qiu et al., A reverse transcription-PCR for subtyping of the neuraminidase of avian influenza viruses. J. Virol. Methods 155(2), 193–198 (2009)
E. Hoffmann et al., Universal primer set for the full-length amplification of all influenza A viruses. Arch. Virol. 146(12), 2275–2289 (2001)
X. Qi et al., Isolation and genetic characterization of haemagglutinin of the first influenza A H1N1 (2009) viruses from Jiangsu Province. Microbiol. China 37(2), 305–310 (2010)
M. Matrosovich et al., Early alterations of the receptor-binding properties of H1, H2, and H3 avian influenza virus hemagglutinins after their introduction into mammals. J. Virol. 74(18), 8502–8512 (2000)
G.N. Rogers et al., Single amino acid substitutions in influenza haemagglutinin change receptor binding specificity. Nature 304(5921), 76–78 (1983)
G.K. Goh et al., Protein intrinsic disorder and influenza virulence: the 1918 H1N1 and H5N1 viruses. Virol. J. 6, 69 (2009)
A.J. Caton et al., The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 31(2Pt1), 417–427 (1982)
C. Scholtissek et al., How to overcome resistance of influenza A viruses against adamantane derivatives. Antivir. Res. 37(2), 83–95 (1998)
H. Suzuki et al., Emergence of amantadine-resistant influenza A viruses: epidemiological study. J. Infect. Chemother. 9(3), 195–200 (2003)
J.M. Katz et al., Molecular correlates of influenza A H5N1 virus pathogenesis in mice. J. Virol. 74(22), 10807–10810 (2000)
P. Massin et al., Residue 627 of PB2 is a determinant of cold sensitivity in RNA replication of avian influenza viruses. J. Virol. 75(11), 5398–5404 (2001)
Z. Li et al., Molecular basis of replication of duck H5N1 influenza viruses in a mammalian mouse model. J. Virol. 79(18), 12058–12064 (2005)
G.R. Bai et al., Amantadine- and oseltamivir-resistant variants of influenza A viruses in Thailand. Biochem. Biophys. Res. Commun. 390(3), 897–901 (2009)
X. Qi et al., Molecular characterization of highly pathogenic H5N1 avian influenza A viruses isolated from raccoon dogs in China. PLoS One 4(3), e4682 (2009)
C. Scholtissek et al., Genetic relatedness of hemagglutinins of the H1 subtype of influenza A viruses isolated from swine and birds. Virology 129(2), 521–523 (1983)
V. Gregory et al., Human infection by a swine influenza A (H1N1) virus in Switzerland. Arch. Virol. 148(4), 793–802 (2003)
K.P. Myers et al., Cases of swine influenza in humans: a review of the literature. Clin. Infect. Dis. 44(8), 1084–1088 (2007)
G.F. Rimmelzwaan et al., Antigenic and genetic characterization of swine influenza A (H1N1) viruses isolated from pneumonia patients in The Netherlands. Virology 282(2), 301–306 (2001)
H. Zhu et al., Novel reassortment of Eurasian avian-like and pandemic/2009 influenza viruses in swine: infectious potential for humans. J. Virol. 85(20), 10432–10439 (2011)
Acknowledgments
This study was supported by the Major State Basic Research Development Program (973 Program) (No. 2011CB505003), the Important National Science & Technology Specific Projects (Nos. 2008ZX10004-013 and 2009ZX10004-214), and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, G., Pan, J., Gu, X. et al. Isolation and phylogenetic analysis of avian-origin European H1N1 swine influenza viruses in Jiangsu, China. Virus Genes 44, 295–300 (2012). https://doi.org/10.1007/s11262-011-0704-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-011-0704-7