Abstract
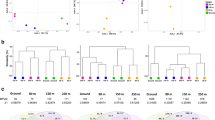
Bacteria are ubiquitous in the atmosphere, where they form a highly diverse community, albeit low in abundance. Several approaches are available for collecting airborne particles, though few comparative studies have been conducted to date. This study examined how different sampling strategies affect the apparent composition of the airborne community. Three devices were tested: an impactor, a liquid impinger, and a Teflon membrane filter. Comparative studies were conducted at one mountainous location in Norway and one seaside location in Sweden. At both locations, microbial samples were collected in parallel using the sampling devices. DNA extraction, construction of 16S rRNA gene clone libraries, and subsequent sequencing were used to identify the bacteria. The comparison between clone libraries retrieved using the different devices indicated good agreement regarding dominant species, overall diversity, and distribution of species among phylogenetic groups. Among the less common species, there were few shared sequences in different clone libraries, likely due to the high diversity of the assessed samples. Bacteria belonging to the Bacteroidetes and Proteobacteria phyla dominated at both locations, and the most common genera were Sphingomonas sp. and Pantoea sp. Chloroplast-like 16S rRNA gene sequences were detected in all samples.



Similar content being viewed by others
References
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215(3), 403–410.
Amato, P., Parazols, M., Sancelme, M., Laj, P., Mailhot, G., & Delort, A. M. (2007). Microorganisms isolated from the water phase of tropospheric clouds at the puy de dome: Major groups and growth abilities at low temperatures. FEMS Microbiology Ecology, 59(2), 242–254. doi:10.1111/J.1574-6941.2006.00199.X.
Baron, P. A., & Willeke, K. (2001). Aerosol measurement: Principle, techniques, and applications. New York: Wiley.
Bauer, H., Kasper-Giebl, A., Löflund, M., Giebl, H., Hitzenberger, R., Zibuschka, F., et al. (2002). The contribution of bacteria and fungal spores to the organic carbon content of cloud water, precipitation and aerosols. Atmospheric Research, 64, 109–119.
Bent, S. J., & Forney, L. J. (2008). The tragedy of the uncommon: Understanding limitations in the analysis of microbial diversity. The ISME Journal, 2, 689–695.
Bovallius, A., Bucht, B., Roffey, R., & Anäs, P. (1978). Long-range air transmission of bacteria. Applied and Environmental Microbiology, 35, 1231–1232.
Bowers, R. M., Lauber, C. L., Wiedinmyer, C., Hamady, M., Hallar, A. G., Fall, R., et al. (2009). Characterization of airborne microbial communities at a high-elevation site and their potential to act as atmospheric ice nuclei. Applied and Environmental Microbiology, 75(15), 5121–5130. doi:10.1128/Aem.00447-09.
Brodie, E. L., DeSantis, T. Z., Parker, J. P. M., Zubietta, I. X., Piceno, Y. M., & Andersen, G. L. (2007). Urban aerosols harbor diverse and dynamic bacterial populations. Proceedings of the National Academy of Sciences, USA, 104(1), 299–304.
Brown, J. K. M., & Hovmoller, M. S. (2002). Epidemiology—aerial dispersal of pathogens on the global and continental scales and its impact on plant disease. Science, 297(5581), 537–541.
Burton, N. C., Grinshpun, S. A., & Reponen, T. (2007). Physical collection efficiency of filter materials for bacteria and viruses. The Annals of Occupational Hygiene, 51(2), 143–151. doi:10.1093/Annhyg/Mel073.
Chao, A., Chazdon, R. L., Colwell, R. K., & Shen, T.-J. (2005). A new statistical approach for assessing similarity of species composition with incidence and abundance data. Ecology Letters, 8, 148–159.
Colwell, R. K. (2009). Estimates: Statistical estimation of species richness and shared species from samples. Version 8.2. User’s guide and application published at: http://purl.Oclc.Org/estimates.
Despres, V., Nowoisky, J., Klose, M., Conrad, R., Andreae, M. O., & Pöschl, U. (2007). Molecular genetics and diversity of primary biogenic aerosol particles in urban, rural, and high alpine air. Biogeosciences Discussion, 4, 349–384.
Fahlgren, C., Hagstrom, A., Nilsson, E. D., & Zweifel, U. L. (2010). Annual variation of the diversity, viability, and origin of airborne bacteria. Applied and Environmental Microbiology, 76, 3015–3025.
Fields, N. D., Oxborrow, G. S., Puleo, J. R., & Herring, C. M. (1974). Evaluation of membrane filter field monitors for microbiologial air sampling. Applied Microbiology, 27, 517–520.
Fierer, N., Liu, Z. Z., Rodriguez-Hernandez, M., Knight, R., Henn, M., & Hernandez, M. T. (2008). Short-term temporal variability in airborne bacterial and fungal populations. Applied and Environmental Microbiology, 74(1), 200–207. doi:10.1128/Aem.01467-07.
Frohlich-Nowoisky, J., Pickersgill, D. A., Despres, V. R., & Poschl, U. (2009). High diversity of fungi in air particulate matter. Proceedings of the National Academy of Sciences, USA, 106(31), 12814–12819. doi:10.1073/Pnas.0811003106.
Good, I. J. (1953). The population frequencies of species and the estimation of population parameters. Biometrika, 40(3–4), 237–264.
Griffin, D. W., Kellogg, C. A., Garrison, V. H., Lisle, J. T., Borden, T. C., & Shinn, E. A. (2003). Atmospheric microbiology in the Northern Caribbean during African dust events. Aerobiologia, 19, 143–157.
Hagström, Å., Pinhassi, J., & Zweifel, U. L. (2000). Biogeographical diversity among marine bacterioplankton. Aquatic Microbial Ecology, 21, 231–244.
Harrison, R. M., Jones, A. M., Biggins, P. D. E., Pomeroy, N., Cox, C. S., Kidd, S. P., et al. (2005). Climate factors influencing bacterial count in background air samples. International Journal of Biometeorology, 49(3), 167–178. doi:10.1007/S00484-004-0225-3.
Hervas, A., Camarero, L., Reche, I., & Casamayor, E. O. (2009). Viability and potential for immigration of airborne bacteria from Africa that reach high mountain lakes in Europe. Environmental Microbiology, 11(6), 1612–1623. doi:10.1111/J.1462-2920.2009.01926.X.
Jensen, P. A., Todd, W. F., Davis, G. N., & Scarpino, P. V. (1992). Evaluation of eight bioaersol samplers challenged with aerosols of free bacteria. American Industrial Hygiene Association Journal, 53, 660–667.
Kellogg, C. A., & Griffin, D. W. (2006). Aerobiology and the global transport of desert dust. Trends in Ecology & Evolution, 21(11), 638–644. doi:10.1016/J.Tree.2006.07.004.
Kesavan, J., Bottiger, J. R., & McFarland, A. R. (2008). Bioaerosol concentrator performance: Comparative tests with viable and with solid and liquid nonviable particle. Journal of Applied Microbiology, 104, 285–295.
Kesavan, J., Schepers, D., & McFarland, A. R. (2010). Sampling and retention efficiencies of batch-type liquid-based bioaerosol samplers. Aerosol Science and Technology, 44, 817–829.
King, M. D., Thien, B. F., Tiirikainen, S., & McFarland, A. R. (2009). Collection characteristics of a batch-type wetted wall bioaerosol sampling cyclone. Aerobiologia 25, 239–247.
Lighthart, B. (1997). The ecology of bacteria in the alfresco atmosphere. FEMS Microbiology Ecology, 23, 263–274.
Lundholm, I. M. (1982). Comparison of methods for quantitative-determinations of airborne bacteria and evaluation of total viable counts. Applied and Environmental Microbiology, 44(1), 179–183.
Maron, P. A., Lejon, D. P. H., Carvalho, E., Bizet, K., Lemanceau, P., Ranjard, L., et al. (2005). Assessing genetic structure and diversity of airborne bacterial communities by DNA fingerprinting and 16s rdna clone library. Atmospheric Environment, 39(20), 3687–3695. doi:10.1016/J.Atosenv.2005.03.002.
Möhler, O., DeMott, P. J., Vali, G., & Levin, Z. (2007). Microbiology and atmospheric processes: The role of biological particles in cloud physics. Biogeosciences, 4, 1059–1071.
Pommier, T., Canback, B., Riemann, L., Bostrom, K. H., Simu, K., Lundberg, P., et al. (2007). Global patterns of diversity and community structure in marine bacterioplankton. Molecular Ecology, 16(4), 867–880. doi:10.1111/J.1365-294x.2006.03189.X.
Porter, K. G., & Feig, Y. S. (1980). The use of dapi for identifying and counting aquatic microflora. Limnology and Oceanography, 25(5), 943–948.
Shannon, C. E. (1948). A mathematical theory of communication. Bell System Technical Journal, 27(4), 623–656.
Soltis, P. S., Soltis, D. E., & Doyle, J. J. (1998). Molecular systematics of plants ii: DNA sequencing. Dordrecht: Kluwer Academic Publishers.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). Molecular evolutionary genetics analysis (mega) software version 4.0. Molecular Biology and Evolution, 24, 1596–1599.
Tormo, R., Recio, D., Silva, I., & Muñoz, A. F. (2001). A quantitative investigation of airborne algae and liche soredia obtained from pollen traps in South-West Spain. European Journal of Phycology, 36, 385–390.
Weisburg, W. G., Barns, S. M., Pelletier, D. A., & Lane, D. J. (1991). 16s ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173(2), 697–703.
Whyte, W., Green, G., & Albisu, A. (2007). Collection efficiency and design of microbial air samplers. Journal of Aerosol Science, 38, 101–114.
Wittmaack, K., Whnes, H., Heinzmann, U., & Agerer, R. (2005). An overview on bioaerosols viewed by scanning electron microscopy. Science of the Total Environment, 346, 244–255.
Acknowledgments
Hilde Kristiansen and Sabina Arnautovic are thanked for helping with sample collection. This work was financed by the Swedish Research Council for Environment, Agricultural Science and Spatial Planning (FORMAS), grant no. 214-2008-1113, by EU grant no. SEC6-PR-214400 (AEROBACTICS), and by the Research Council of Norway, grant no. 177802/V40.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fahlgren, C., Bratbak, G., Sandaa, RA. et al. Diversity of airborne bacteria in samples collected using different devices for aerosol collection. Aerobiologia 27, 107–120 (2011). https://doi.org/10.1007/s10453-010-9181-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10453-010-9181-z