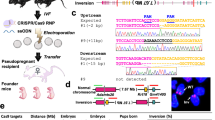
Abstract
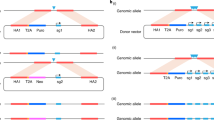
Balancer chromosomes are genetic reagents that are used in Drosophila melanogaster for stock maintenance and mutagenesis screens1. Despite their utility, balancer chromosomes are rarely used in mice because they are difficult to generate using conventional methods. Here we describe the engineering of a mouse balancer chromosome with the Cre-loxP recombination system. The chromosome features a 24-centiMorgan (cM) inversion between Trp53 (also known as p53) and Wnt3 on mouse chromosome 11 that is recessive lethal and dominantly marked with a K14-Agouti transgene2. When allelic to a wild-type chromosome, the inversion suppresses crossing over in the inversion interval, accompanied by elevated recombination in the flanking regions. The inversion functions as a balancer chromosome because it can be used to maintain a lethal mutation in the inversion interval as a self-sustaining trans-heterozygous stock. This strategy can be used to generate similar genetic reagents throughout the mouse genome. Engineering of visibly marked inversions and deficiencies is an important step toward functional analyses of the mouse genome and will facilitate large-scale mutagenesis programs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ashburner, M. Drosophila: A Laboratory Handbook (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, 1989).
Kucera, G.T., Bortner, D.M. & Rosenberg, M.P. Overexpression of an Agouti cDNA in the skin of transgenic mice recapitulates dominant coat color phenotypes of spontaneous mutants. Dev. Biol. 173, 162–173 (1996).
Muller, H.J. Genetic variability, twin hybrids and constant hybrids, in a case of balanced lethal factors. Genetics 3, 422–499 (1918).
Strickberger, M.W. Genetics 435–438 (Macmillan, New York, 1985).
Brown, S.D. & Nolan, P.M. Mouse mutagenesis-systematic studies of mammalian gene function. Hum. Mol. Genet. 7, 1627–1633 (1998).
Hrabe de Angelis, M. & Balling, R. Large scale ENU screens in the mouse: genetics meets genomics. Mutat. Res. 400, 25–32 (1998).
Takahashi, J.S., Pinto, L.H. & Vitaterna, M.H. Forward and reverse genetic approaches to behavior in the mouse. Science 264, 1724–1733 (1994).
Dietrich, W.F. et al. A comprehensive genetic map of the mouse genome. Nature 380, 149–152 (1996).
Talbot, C.J. et al. High-resolution mapping of quantitative trait loci in outbred mice. Nature Genet. 21, 305–308 (1999).
Holdener-Kenny, B., Sharan, S.K. & Magnuson, T. Mouse albino-deletions: from genetics to genes in development. Bioessays 14, 831–839 (1992).
Rinchik, E.M. & Russell, L.B. Germ-line deletion mutations in the mouse: tools for intensive functional and physical mapping of regions of the mammalian genome. in Genome Analysis Volume 1: Genetic and Physical Mapping (eds Davies, K.E. & Tilghman, S.M.) 121–158 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1990).
Schumacher, A., Faust, C. & Magnuson, T. Positional cloning of a global regulator of anterior-posterior patterning in mice. Nature 383, 250–253 (1996).
Schuler, G.D. et al. A gene map of the human genome. Science 274, 540–546 (1996).
Liu. P. et al. Requirement for Wnt3 in vertebrate axis formation. Nature Genet. 22, 361–365 (1999).
Ramirez-Solis, R., Liu, P. & Bradley, A. Chromosome engineering in mice. Nature 378, 720–724 (1995).
Zheng, B., Mills, A.A. & Bradley, A. A system for rapid generation of coat color-tagged knockouts and defined chromosomal rearrangements in mice. Nucleic Acids Res. 27, 2354–2360 (1999).
Yokoyama, T. et al. Conserved cysteine to serine mutation in tyrosinase is responsible for the classical albino mutation in laboratory mice. Nucleic Acids Res. 18, 7293–7298 (1990).
Liu, P., Zhang, H., McLellan, A., Vogel, H. & Bradley, A. Embryonic lethality and tumorigenesis caused by segmental aneuploidy on mouse chromosome 11. Genetics 150, 1155–1168 (1998).
Ramirez-Solis, R., Davis, A.C. & Bradley, A. Gene targeting in embryonic stem cells. Methods Enzymol. 225, 855–878 (1993).
Singleton, J.R. A mechanism intrinsic to heterozygous inversions affecting observed recombination frequencies in adjacent regions. Genetics 49, 541–560 (1964).
Ramirez-Solis, R., Zheng, H., Whiting, J., Krumlauf, R. & Bradley, A. Hoxb-4 (Hox-2.6) mutant mice show homeotic transformation of a cervical vertebra and defects in the closure of the sternal rudiments. Cell 73, 279–294 (1993).
Justice, M.J. Mutagenesis of the mouse germline. in Mouse Genetics and Transgenics: A Practical Approach (eds Jackson, I.J. & Abbott, C.A.) (Oxford University Press, Oxford, in press).
Lee, G. & Saito, I. Role of nucleotide sequences of loxP spacer region in Cre-mediated recombination. Gene 216, 55–65 (1998).
Araki, K., Araki, M. & Yamamura, K. Targeted integration of DNA using mutant lox sites in embryonic stem cells. Nucleic Acids Res. 25, 868–872 (1997).
Buchholz, F., Angrand, P.O. & Stewart, A.F. Improved properties of FLP recombinase evolved by cycling mutagenesis. Nature Biotechnol. 16, 657–662 (1998).
Dymecki, S.M. Flp recombinase promotes site-specific DNA recombination in embryonic stem cells and transgenic mice. Proc. Natl Acad. Sci. USA 93, 6191–6196 (1996).
Zakut-Houri, R. et al. A single gene and a pseudogene for the cellular tumour antigen p53. Nature 306, 594–597 (1983).
Bienz, B., Zakut-Houri, R., Givol, D. & Oren, M. Analysis of the gene coding for the murine cellular tumour antigen p53. EMBO J. 3, 2179–2183 (1984).
Matzuk, M.M., Finegold, M.J., Su, J.G., Hsueh, A.J. & Bradley, A. α-inhibin is a tumour-suppressor gene with gonadal specificity in mice. Nature 360, 313–319 (1992).
Robertson, E.J. Embryo-derived stem cell lines. in Teratocarcinomas and Embryonic Stem Cells: A Practical Approach (ed. Robertson, E.J.) 71–112 (IRL, Oxford, 1987).
Baldini, A. & Lindsay, E.A. Mapping human YAC clones by fluorescence in situ hybridization using Alu-PCR from single yeast colonies. in In Situ Hybridization Protocols (ed. Choo, K.H.A.) 75–84 (Humana Press, Totowa, New Jersey, 1994).
Acknowledgements
We thank H. Su for information on polymorphic markers used in assessing recombination frequencies; M. Suraokar for the Hoxb4r mouse line; S. Rivera and S. Vaishnav for preparing feeder cells; J. Weber for helpful discussions; and H. Bellen and M. Justice for helpful comments on the manuscript. This work was supported partly by grants from the National Institutes of Health and the U.S. Department of Defense. A.B. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zheng, B., Sage, M., Cai, WW. et al. Engineering a mouse balancer chromosome. Nat Genet 22, 375–378 (1999). https://doi.org/10.1038/11949
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/11949
This article is cited by
-
Fast and efficient generation of a full-length balancer chromosome by a single Cre/loxP recombination event
Mammalian Genome (2022)
-
GRIBCG: a software for selection of sgRNAs in the design of balancer chromosomes
BMC Bioinformatics (2019)
-
A method using electroporation for the protein delivery of Cre recombinase into cultured Arabidopsis cells with an intact cell wall
Scientific Reports (2019)
-
QTL analysis of modifiers for pigmentary disorder in rats carrying Ednrbsl mutations
Scientific Reports (2016)
-
Understanding the DNA damage response in order to achieve desired gene editing outcomes in mosquitoes
Chromosome Research (2015)