Abstract
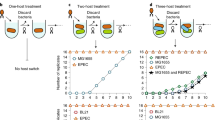
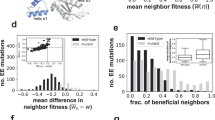
The ubiquity of mechanisms that generate genetic variation has spurred arguments that evolvability, the ability to generate adaptive variation, has itself evolved in response to natural selection1,2. The high mutation rate of RNA viruses is postulated to be an adaptation for evolvability3,4, but the paradox is that whereas some RNA viruses evolve at high rates4,5, others are highly stable5,6. Here we show that evolvability in the RNA bacteriophage φ6 is also determined by the accessibility of advantageous genotypes within the mutational neighbourhood (the set of mutants one or a few mutational steps away). We found that two φ6 populations that were derived from a single ancestral phage repeatedly evolved at different rates and toward different fitness maxima. Fitness measurements of individual phages showed that the fitness distribution of mutants differed between the two populations. Whereas population A, which evolved toward a higher maximum, had a distribution that contained many advantageous mutants, population B, which evolved toward a lower maximum, had a distribution that contained only deleterious mutants. We interpret these distributions to measure the fitness effects of genotypes that are mutationally available to the two populations. Thus, the /evolvability of φ6 is constrained by the distribution of its mutational neighbours, despite the fact that this phage has the characteristic high mutation rate of RNA viruses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Radman, M., Matic, I. & Taddei, F. Evolution of Evolvability. Ann. N. Y. Acad. Sci. 870, 146–155 ( 1999).
Wagner, G. P., Chiu, C. & Hansen, T. F. Is Hsp90 a regulator of evolvability? J. Exp. Zool. 285, 116–118 (1999).
Holland, J. et al. Rapid evolution of RNA genomes. Science 215, 1577–1585 (1982).
Domingo, E. & Holland, J. J. in The Evolutionary Biology of Viruses (ed. Morse, S. S.) 161–184 (Raven, New York, 1994).
Fraile, A. et al. A century of tobramovirus evolution in an Australian population of Nicotiana glauca. J. Virol. 71, 8316–8320 (1997).
Chao, L. Evolution of sex and the molecular clock in RNA viruses. Gene 205, 301–308 (1997).
Burch, C. L. & Chao, L. Evolution by small steps and rugged landscapes in the RNA virus φ6. Genetics 151, 921–927 (1999).
Fenster, C. B., Galloway, L. F. & Chao, L. Epistasis and its consequences for the evolution of natural populations. Trends Ecol. Evol. 12, 282–286 (1997).
Wright, S. The roles of mutation, inbreeding, crossbreeding and selection in evolution. Proc. Sixth Int. Congress Genetics 1, 356 –366 (1932).
Eigen, M. New concepts for dealing with the evolution of nucleic acids. Cold Spring Harbor Symp. Quant. Biol. 52, 307– 320 (1987).
Kauffman, S. A. The Origins of Order. (Oxford Univ. Press, New York, 1993).
Nijhuis, M. et al. Increased fitness of drug resistant HIV-1 protease as a result of acquisition of compensatory mutations during suboptimal therapy. AIDS 13, 2349–2359 ( 1999).
Eigen, M. On the nature of viral quasispecies. Trends. Microbiol. 4, 212–214 (1996).
Schuster, P. & Swetina, J. Stationary mutant distributions and evolutionary optimization. Bull. Math. Biol. 50 , 636–660 (1988).
Vidaver, A. K., Kolski, R. K. & van Etten, J. L. Bacteriophage φ6: a lipid-containing virus of Pseudomonas phaseolicola. J. Virol. 11, 799–805 (1973).
Mindich, L., Sinclair, J. F., Levine, D. & Cohen, J. Genetic studies of temperature-sensitive and nonsense mutants of bacteriophage φ6. Virology 75, 218–223 (1976).
Chao, L., Trang, T. T. & Trang, T. T. The advantage of sex in the RNA virus φ6. Evolution 46, 289–299 ( 1997).
Chao, L. Fitness of RNA virus decreased by Muller's ratchet. Nature 348, 454–455 (1990).
Sokal, R. R. & Rohlf, F. J. Biometry 3rd edn (Freeman, San Francisco, 1995).
Acknowledgements
We thank our laboratory group for feedback, D. Metzgar for comments on manuscript and L. Mindich for discussion and stocks. Support was provided to C.L.B. by a predoctoral fellowship from the Howard Hughes Medical Institute and a Dissertation Improvement Grant from the National Science Foundation. Support was provided to L.C. by the University of California San Diego and the National Institutes of Health (NIGMS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Burch, C., Chao, L. Evolvability of an RNA virus is determined by its mutational neighbourhood . Nature 406, 625–628 (2000). https://doi.org/10.1038/35020564
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35020564
This article is cited by
-
Host diversity slows bacteriophage adaptation by selecting generalists over specialists
Nature Ecology & Evolution (2021)
-
Virus Disinfection and Population Genetics: Toward the Control of Waterborne Virus Diseases by Water Engineering
Current Pollution Reports (2021)
-
Diminishing-returns epistasis decreases adaptability along an evolutionary trajectory
Nature Ecology & Evolution (2017)
-
Attenuation of RNA viruses by redirecting their evolution in sequence space
Nature Microbiology (2017)
-
Contrasting effects of historical contingency on phenotypic and genomic trajectories during a two-step evolution experiment with bacteria
BMC Evolutionary Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.