Key Points
-
Segmental duplications are a class of repetitive DNA element in the human genome.
-
Segmental duplications have various sizes and organizational configurations.
-
The different copies of a segmental duplication can share 96–99% nucleotide sequence identity with each other.
-
Segmental duplications have been implicated in the aetiology of chromosomal rearrangements that are associated with several genomic disorders.
-
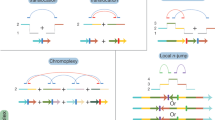
The chromosomal rearrangements include deletions, interstitial duplications, inversions, supernumerary marker chromosomes and translocations.
-
The genomic disorders include DiGeorge and velocardiofacial syndromes, cat eye syndrome, Prader–Willi and Angelman syndromes, Williams–Beuren syndrome and many more.
-
Many models are proposed to explain the mechanisms involved in segmental-duplication-mediated rearrangements.
-
All of these models are based on misalignment between non-allelic segmental duplications, followed by recombination.
-
Segmental duplications represent an under-appreciated source of genetic change in the human genome.
Abstract
The knowledge that specific genetic diseases are caused by recurrent chromosomal aberrations has indicated that genomic instability might be directly related to the structure of the regions involved. The sequencing of the human genome has directed significant attention towards understanding the molecular basis of such recombination 'hot spots'. Segmental duplications have emerged as a significant factor in the aetiology of disorders that are caused by abnormal gene dosage. These observations bring us closer to understanding the mechanisms and consequences of genomic rearrangement.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mazzarella, R. & Schlessinger, D. Pathological consequences of sequence duplications in the human genome. Genome Res. 8, 1007–1021 (1998).
Lupski, J. R. Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 14, 417–422 (1998).
Ji, Y., Eichler, E. E., Schwartz, S. & Nicholls, R. D. Structure of chromosomal duplicons and their role in mediating human genomic disorders. Genome Res. 10, 597–610 (2000).References 1–3 are excellent early reviews that discuss segmental duplications in the human genome and their involvement in genomic disorders.
IHGSC (International Human Genome Sequencing Consortium). Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).Landmark paper that reports the draft sequence of the human genome and also presents a comprehensive analysis of repetitive and duplicated DNA in the entire human genome.
Shaikh, T. H., Kurahashi, H. & Emanuel, B. S. Evolutionarily conserved duplications in 22q11 mediate deletions, duplications, translocations and genomic instability. Genet. Med. 3, 6–13 (2001).
Eichler, E. E. et al. Duplication of a gene rich cluster between 16p11.1 and Xq28: a novel pericentromeric-directed mechanism for paralogous genome evolution. Hum. Mol. Genet. 5, 899–912 (1996).
Eichler, E. E. et al. Interchromosomal duplications of the adrenoleukodystrophy locus: a phenomenon of pericentromeric plasticity. Hum. Mol. Genet. 6, 991–1002 (1997).
Regnier, V. et al. Emergence and scattering of multiple neurofibromatosis (NF-1)-related sequences during hominoid evolution suggest a process of pericentromeric interchromosomal transposition. Hum. Mol. Genet. 6, 9–16 (1997).References 7 and 8 were among the earliest papers to recognize the pericentromeric plasticity of chromosomes in higher primates.
Trask, B. et al. Members of the olfactory receptor gene family are contained in large blocks of DNA duplicated polymorphically near the ends of human chromosomes. Hum. Mol. Genet. 7, 13–26 (1998).
Trask, B. J. et al. Large multi-chromosomal duplications encompass many members of the olfactory receptor gene family in the human genome. Hum. Mol. Genet. 7, 2007–2020 (1998).
Purandare, S. M. & Patel, P. I. Recombination hot spots and human disease. Genome Res. 7, 773–786 (1997).
Shaffer, L. G. & Lupski, J. R. Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu. Rev. Genet. 34, 297–329 (2000).
Chance, P. F. et al. Two autosomal dominant neuropathies result from reciprocal DNA duplication/deletion of a region on chromosome 17. Hum. Mol. Genet. 3, 223–228 (1994).
Lupski, J. R. Charcot–Marie–Tooth disease: lessons in genetic mechanisms. Mol. Med. 4, 3–11 (1998).
Dorschner, M. O., Sybert, V. P., Weaver, M., Pletcher, B. A. & Stephens, K. NF1 microdeletion breakpoints are clustered at flanking repetitive sequences. Hum. Mol. Genet. 9, 35–46 (2000).
Christian, S. L., Fantes, J. A., Mewborn, S. K., Huang, B. & Ledbetter, D. H. Large genomic duplicons map to sites of instability in Prader–Willi/Angelman syndrome chromosome region (15q11–q13). Hum. Mol. Genet. 8, 1025–1037 (1999).
Amos-Landgraf, J. M. et al. Chromosome breakage in the Prader–Willi and Angelman syndromes involves recombination between large, transcribed repeats at proximal and distal breakpoints. Am. J. Hum. Genet. 65, 370–386 (1999).
Perez-Jurado, L. A. et al. A duplicated gene in the breakpoint regions of the 7q11.23 Williams–Beuren syndrome deletion encodes the initiator binding protein TFII-I and BAP-135, a phosphorylation target of BTK. Hum. Mol. Genet. 7, 325–334 (1998).
Peoples, R. et al. A physical map, including a BAC/PAC clone contig, of the Williams–Beuren syndrome –deletion region at 7q11.23. Am. J. Hum. Genet. 66, 47–68 (2000).
Chen, K. S. et al. Homologous recombination of a flanking repeat gene cluster is a mechanism for a common contiguous gene deletion syndrome. Nature Genet. 17, 154–163 (1997).
Potocki, L. et al. Molecular mechanism for duplication 17p11.2 – the homologous recombination reciprocal of the Smith–Magenis microdeletion. Nature Genet. 24, 84–87 (2000).
McTaggart, K. E. et al. Cat eye syndrome chromosome breakpoint clustering: identification of two intervals also associated with 22q11 deletion syndrome breakpoints. Cytogenet. Cell Genet. 81, 222–228 (1998).
Edelmann, L., Pandita, R. K. & Morrow, B. E. Low-copy repeats mediate the common 3-Mb deletion in patients with velo-cardio-facial syndrome. Am. J. Hum. Genet. 64, 1076–1086 (1999).
Edelmann, L. et al. A common molecular basis for rearrangement disorders on chromosome 22q11. Hum. Mol. Genet. 8, 1157–1167 (1999).
Shaikh, T. H. et al. Chromosome 22-specific low copy repeats and the 22q11.2 deletion syndrome: genomic organization and deletion endpoint analysis. Hum. Mol. Genet. 9, 489–501 (2000).
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).References 25 and 26 present a detailed, sequence-based analysis of segmental duplications on 22q11.
Emanuel, B. S. et al. in Etiology and Morphogenesis of Congenital Heart Disease (eds Clark, E., Nakazawa, M. & Takao, A.) 335–339 (Futura Publishing Co., Inc., Armonk, New York, 1999).
Shaikh, T. H., Budarf, M. L., Celle, L., Zackai, E. H. & Emanuel, B. S. Clustered 11q23 and 22q11 breakpoints and 3:1 meiotic malsegregation in multiple unrelated t(11;22) families. Am. J. Hum. Genet. 65, 1595–1607 (1999).
Edelmann, L. et al. A common breakpoint on 11q23 in carriers of the constitutional t(11;22) translocation. Am. J. Hum. Genet. 65, 1608–1616 (1999).
Kurahashi, H. et al. Regions of genomic instability on 22q11 and 11q23 as the etiology for the recurrent constitutional t(11;22). Hum. Mol. Genet. 9, 1665–1670 (2000).
Kurahashi, H. et al. Tightly clustered 11q23 and 22q11 breakpoints permit PCR-based detection of the recurrent constitutional t(11;22). Am. J. Hum. Genet. 67, 763–768 (2000).References 30 and 31 present extensive research in mapping the breakpoints of the constitutional t(11; 22). Reference 30 was the first to show that the t(11; 22) breakpoints localize to palindromic (A+T)-rich repeats. Reference 31 describes identical sequences of t(11; 22) breakpoints in several, unrelated translocation carriers.
Reiter, L. T., Murakami, T., Koeuth, T., Gibbs, R. A. & Lupski, J. R. The human COX10 gene is disrupted during homologous recombination between the 24 kb proximal and distal CMT1A-REPs. Hum. Mol. Genet. 6, 1595–1603 (1997).
Reiter, L. T. et al. Human meiotic recombination products revealed by sequencing a hotspot for homologous strand exchange in multiple HNPP deletion patients. Am. J. Hum. Genet. 62, 1023–1033 (1998).
Inoue, K. et al. The 1.4-Mb CMT1A duplication/HNPP deletion genomic region reveals unique genome architectural features and provides insights into the recent evolution of new genes. Genome Res. 11, 1018–1033 (2001).
Yen, P. H. et al. Frequent deletions of the human X chromosome distal short arm result from recombination between low copy repetitive elements. Cell 61, 603–610 (1990).
Li, X. M., Yen, P. H. & Shapiro, L. J. Characterization of a low copy repetitive element S232 involved in the generation of frequent deletions of the distal short arm of the human X chromosome. Nucleic Acids Res. 20, 1117–1122 (1992).
Lahn, B. T. & Page, D. C. A human sex-chromosomal gene family expressed in male germ cells and encoding variably charged proteins. Hum. Mol. Genet. 9, 311–319 (2000).
Small, K., Iber, J. & Warren, S. T. Emerin deletion reveals common X-chromosome inversion mediated by inverted repeats. Nature Genet. 16, 96–99 (1997).
Naylor, J. A. et al. Investigation of the factor VIII intron 22 repeated region (int22h) and the associated inversion junctions. Hum. Mol. Genet. 4, 1217–1224 (1995).
Naylor, J. A. et al. A novel DNA inversion causing severe hemophilia A. Blood 87, 3255–3261 (1996).
Kehrer-Sawatzki, H., Schwickardt, T., Assum, G., Rocchi, M. & Krone, W. A third neurofibromatosis type 1 (NF1) pseudogene at chromosome 15q11.2. Hum. Genet. 100, 595–600 (1997).
Edelmann, L. et al. AT-rich palindromes mediate the constitutional t(11;22) translocation. Am. J. Hum. Genet. 68, 1–13 (2001).
Ji, Y. et al. The ancestral gene for transcribed, low-copy repeats in the Prader–Willi/Angelman region encodes a large protein implicated in protein trafficking, which is deficient in mice with neuromuscular and spermiogenic abnormalities. Hum. Mol. Genet. 8, 533–542 (1999).
Ji, Y. et al. Structure of the highly conserved HERC2 gene and of multiple partially duplicated paralogs in human. Genome Res. 10, 319–329 (2000).
Pujana, M. A. et al. Additional complexity on human chromosome 15q: identification of a set of newly recognized duplicons (LCR15) on 15q11–q13, 15q24, and 15q26. Genome Res. 11, 98–111 (2001).
Valero, M. C., de Luis, O., Cruces, J. & Perez Jurado, L. A. Fine-scale comparative mapping of the human 7q11.23 region and the orthologous region on mouse chromosome 5G: the low-copy repeats that flank the Williams–Beuren syndrome deletion arose at breakpoint sites of an evolutionary inversion. Genomics 69, 1–13 (2000).
Kiyosawa, H. & Chance, P. F. Primate origin of the CMT1A-REP repeat and analysis of a putative transposon-associated recombinational hot spot. Hum. Mol. Genet. 8, 745–753 (1996).
Reiter, L. T. et al. A recombination hotspot responsible for two inherited peripheral neuropathies is located near a mariner transposon-like element. Nature Genet. 12, 288–297 (1996).This paper identifies a recombination hot spot in the CMT1A-REP. This report was the first to indicate that there might be recombinogenic hot spots in large segmental duplications.
Lopez-Correa, C. et al. Recombination hotspot in NF1 microdeletion patients. Hum. Mol. Genet. 10, 1387–1392 (2001).
Zimonjic, D., Kelley, M., Rubin, J., Aaronson, S. & Popescu, N. Fluorescence in situ hybridization analysis of keratinocyte growth factor gene amplification and dispersion in evolution of great apes and humans. Proc. Natl Acad. Sci. USA 94, 11461–11465 (1997).
Keller, M. P., Seifried, B. A. & Chance, P. F. Molecular evolution of the CMT1A-REP region: a human and chimpanzee-specific repeat. Mol. Biol. Evol. 16, 1019–1026 (1999).
Botta, A., Lindsay, E. A., Jurecic, V. & Baldini, A. Comparative mapping of the DiGeorge syndrome region in mouse shows inconsistent gene order and differential degree of gene conversion. Mamm. Genome 8, 890–895 (1997).
Puech, A. et al. Comparative mapping of the human 22q11 chromosomal region and the orthologous region in mice reveals complex changes in gene organization. Proc. Natl Acad. Sci. USA 94, 14608–14613 (1997).
Sutherland, H. F., Kim, U.-J. & Scambler, P. J. Cloning and comparative mapping of the DiGeorge syndrome critical region in the mouse. Genomics 52, 37–43 (1998).
Lund, J. et al. Sequence-ready physical map of the mouse chromosome 16 region with conserved synteny to the human velocardiofacial syndrome region on 22q11.2. Mamm. Genome 10, 438–443 (1999).
Bonifas, J. M., Morley, B. J., Oakey, R. E., Kan, Y. W. & Epstein, E. H. Jr Cloning of a cDNA for steroid sulfatase: frequent occurrence of gene deletions in patients with recessive X chromosome-linked ichthyosis. Proc. Natl Acad. Sci. USA 84, 9248–9251 (1987).
Conary, J. T. et al. Genetic heterogeneity of steroid sulfatase deficiency revealed with cDNA for human steroid sulfatase. Biochem. Biophys. Res. Commun. 144, 1010–1017 (1987).
Ballabio, A. & Andria, G. Deletions and translocations involving the distal short arm of the human X chromosome: review and hypotheses. Hum. Mol. Genet. 1, 221–227 (1992).
Ewart, A. K. et al. Hemizygosity at the elastin locus in a developmental disorder, Williams syndrome. Nature Genet. 5, 11–16 (1993).
Dutly, F. & Schinzel, A. Unequal interchromosomal rearrangements may result in elastin gene deletions causing the Williams–Beuren syndrome. Hum. Mol. Genet. 5, 1893–1898 (1996).
Urbán, Z. et al. 7q11.23 deletions in Williams syndrome arise as a consequence of unequal meiotic crossover. Am. J. Hum. Genet. 59, 958–962 (1996).
Baumer, A. et al. High level of unequal meiotic crossovers at the origin of the 22q11.2 and 7q11.23 deletions. Hum. Mol. Genet. 7, 887–894 (1998).
Perez Jurado, L. A., Peoples, R., Kaplan, P., Hamel, B. C. & Francke, U. Molecular definition of the chromosome 7 deletion in Williams syndrome and parent-of-origin effects on growth. Am. J. Hum. Genet. 59, 781–792 (1996).
Robinson, W. P. et al. Delineation of 7q11.2 deletions associated with Williams–Beuren syndrome and mapping of a repetitive sequence to within and to either side of the common deletion. Genomics 34, 17–23 (1996).
Carrozzo, R. et al. Inter- and intrachromosomal rearrangements are both involved in the origin of 15q11–q13 deletions in Prader–Willi syndrome. Am. J. Hum. Genet. 61, 228–231 (1997).
Robinson, W. P. et al. The mechanisms involved in formation of deletions and duplications of 15q11–q13. J. Med. Genet. 35, 130–136 (1998).
Pentao, L., Wise, C. A., Chinault A. C., Patel, P. I. & Lupski, J. R. Charcot–Marie–Tooth type 1A duplication appears to arise from recombination at repeat sequences flanking the 1.5 Mb monomer unit. Nature Genet. 2, 292–300 (1992).
Lupski, J. R. et al. Gene dosage is a mechanism for Charcot–Marie–Tooth disease type 1A. Nature Genet. 1, 29–33 (1992).
Krawczak, M. & Cooper, D. N. Gene deletions causing human genetic disease: mechanisms of mutagenesis and the role of the local DNA environment. Hum. Genet. 86, 425–441 (1991).
Lakich, D., Kazazian, H. H., Antonarakis, S. E. & Gitschier, J. Inversions disrupting the factor VIII gene are a common cause of severe haemophilia A. Nature Genet. 5, 236–241 (1993).
Rossiter, J. P. et al. Factor VIII gene inversions causing severe hemophilia A originate almost exclusively in male germ cells. Hum. Mol. Genet. 3, 1035–1039 (1994).
Blennow, E. et al. Fifty probands with extra structurally abnormal chromosomes characterized by fluorescence in situ hybridization. Am. J. Med. Genet. 55, 85–94 (1995).
Cheng, E. Y. & Gartler, S. M. A fluorescent in situ hybridization analysis of X chromosome pairing in early human female meiosis. Hum. Genet. 94, 389–394 (1994).
Huang, B. et al. Refined molecular characterization of the breakpoints in small inv dup(15) chromosomes. Hum. Genet. 99, 11–17 (1997).
Wandstrat, A. E., Leana-Cox, J., Jenkins, L. & Schwartz, S. Molecular cytogenetic evidence for a common breakpoint in the largest inverted duplications of chromosome 15. Am. J. Hum. Genet. 62, 925–936 (1998).
Wandstrat, A. E. & Schwartz, S. Isolation and molecular analysis of inv dup(15) and construction of a physical map of a common breakpoint in order to elucidate their mechanism of formation. Chromosoma 109, 498–505 (2000).
Christian, S. L. et al. Molecular characterization of two proximal deletion breakpoint regions in both Prader–Willi and Angelman syndrome patients. Am. J. Hum. Genet. 57, 40–48 (1995).
Footz, T. K. et al. Analysis of the cat eye syndrome critical region in humans and the region of conserved synteny in mice: a search for candidate genes at or near the human chromosome 22 pericentromere. Genome Res. 11, 1053–1070 (2001).
Page, S. L., Shin, J. C., Han, J. Y., Choo, K. H. & Shaffer, L. G. Breakpoint diversity illustrates distinct mechanisms for Robertsonian translocation formation. Hum. Mol. Genet. 5, 1279–1288 (1996).
De La Chapelle, A., Herva, R., Koivisto, M. & Aula, P. A deletion in chromosome 22 can cause DiGeorge syndrome. Hum. Genet. 57, 253–256 (1981).
Li, M. et al. Clustering of DiGeorge/velocardiofacial-associated translocations suggestive of a translocation 'hot spot'. Am. J. Hum. Genet. 57, A119 (1995).
Rhodes, C. H. et al. Molecular studies of an ependymoma-associated constitutional t(1;22)(p22;q11.2). Cytogenet. Cell Genet. 78, 247–252 (1997).
Lopes, J. et al. Sex-dependent rearrangements resulting in CMT1A and HNPP. Nature Genet. 17, 136–137 (1997).
Palau, F. et al. Origin of the de novo duplication in Charcot–Marie–Tooth disease type 1A: unequal nonsister chromatid exchange during spermatogenesis. Hum. Mol. Genet. 2, 2031–2035 (1993).
Lopes, J. et al. Fine mapping of de novo CMT1A and HNPP rearrangements within CMT1A-REPs evidences two distinct sex-dependent mechanisms and candidate sequences involved in recombination. Hum. Mol. Genet. 7, 141–148 (1998).
Saitta, S. C. et al. The 22q11.2 deletion: parental origin and meiotic mechanism. Am. J. Hum. Genet. 67, S163 (2000).
Bondeson, M. L. et al. Inversion of the IDS gene resulting from recombination with IDS-related sequences is a common cause of the Hunter syndrome. Hum. Mol. Genet. 4, 615–621 (1995).
Consevage, M. W. et al. Association of a mosaic chromosomal 22q11 deletion with hypoplastic left heart syndrome. Am. J. Cardiol. 77, 1023–1025 (1996).
Kasprzak, L. et al. Deletion of 22q11 in two brothers with different phenotype. Am. J. Med. Genet. 75, 288–291 (1998).
Anguiano, A. et al. Report of a case with manifestations of DiGeorge/velocardiofacial syndrome (VCFS) and microdeletion 22q11.2 mosaicism. Am. J. Hum. Genet. 67, S142 (2000).
Spinner, N. B. & Emanuel, B. S. in Principles and Practice of Medical Genetics Vol. 1 (eds Rimoin, D. L., Conner, J. M., Pyeritz, R. E. & Emery, A. E. H.) 999–1025 (Churchill Livingstone, 1996).
Acknowledgements
The authors acknowledge support for their efforts from the National Institutes of Health. B.S.E. gratefully acknowledges the support provided by the Charles E. H. Upham chair in Pediatrics. T.H.S. is partially supported by funds from the Florence R. C. Murray Foundation.
Author information
Authors and Affiliations
Related links
Glossary
- IMPRINTING
-
A genetic mechanism by which genes are selectively expressed from the maternal or paternal homologue of a chromosome.
- UNIPARENTAL DISOMY
-
A condition whereby an individual or embryo carries two chromosomes inherited from the same parent.
- STENOSIS
-
The blocking of a blood vessel that can be cleared by mechanical disruption.
- CONSTITUTIONAL TRANSLOCATION
-
A rearrangement between two chromosomes that occurs in the parental germ line or very early in embryonic development, such that every cell in the body contains the translocated chromosomes.
- ICHTHYOSIS
-
A genetic disorder that causes the patient to have scaly skin.
- CONSERVED SYNTENY
-
The occurrence of genomic collinearity between homologous genes in different organisms.
- HAPLOTYPE
-
An experimentally determined profile of genetic markers present on a single chromosome of any given individual.
- PULSED-FIELD GEL ELECTROPHORESIS
-
An electrophoretic technique used to separate large fragments of DNA (>20 kb and up to 10 Mb) on an agarose gel by periodically changing the orientation of the electric field applied to the gel.
- BISATELLITED
-
A chromosome that contains two copies of the satellited acrocentric short arm, often as a result of an inverted duplication. It is usually present as a supernumerary marker chromosome in a cell.
- ACROCENTRIC
-
This refers to a chromosome the centromere of which lies very close to one end, such that one arm of the chromosome is much larger than the other.
- ACENTRIC
-
A chromosome or chromosomal fragment that lacks a centromere.
- PARACENTRIC INVERSION
-
An inversion of a chromosomal segment that does not contain the centromere.
Rights and permissions
About this article
Cite this article
Emanuel, B., Shaikh, T. Segmental duplications: an 'expanding' role in genomic instability and disease. Nat Rev Genet 2, 791–800 (2001). https://doi.org/10.1038/35093500
Issue Date:
DOI: https://doi.org/10.1038/35093500
This article is cited by
-
Title-molecular diagnostics of dystrophinopathies in Sri Lanka towards phenotype predictions: an insight from a South Asian resource limited setting
European Journal of Medical Research (2024)
-
Deep psychophysiological phenotyping of adolescents and adults with 22q11.2 deletion syndrome: a multilevel approach to defining core disease processes
BMC Psychiatry (2023)
-
Circular DNA intermediates in the generation of large human segmental duplications
BMC Genomics (2020)
-
Long-read sequence and assembly of segmental duplications
Nature Methods (2019)
-
Autism spectrum disorders: autistic phenotypes and complicated mechanisms
World Journal of Pediatrics (2019)