Key Points
-
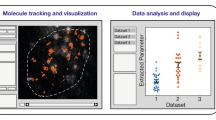
In vivo imaging is routinely used to probe the dynamic behaviour of proteins and cellular compartments. These methods generate large, kinetically complex data sets, which often cannot be intuitively interpreted.
-
High-density data sets contain information that can be used to probe complex systems. Cellular processes that make up complex systems can be displayed as diagrams.
-
Translating cell-biological diagrams into kinetic models is based on standard principles of chemical kinetics. Quantitative models allow simulation of the diagram's response to a given experimental protocol and comparison of its predictions to experimental data.
-
Comparison of model predictions and experimental data permits quantitative hypothesis testing for even very complex hypotheses. An abundance of software tools and databases is available to support this work.
-
The numerical analysis techniques, optimization and parameter estimation, can be used to give each hypothesis its best chance to simultaneously account for all the available experimental data.
-
Alternative hypotheses can be quickly tested. Parameter values can be extracted from experimental data to quantify the three classes of biological processes: transformation, translocation, and binding. Biophysical properties such as diffusion coefficients, rate constants, steady-state distribution ratios, fluxes and residence times are among the most useful quantities that can be obtained.
-
Data collected using GFP-labelled proteins in living cells are particularly well suited to kinetic analysis. This is especially true in the context of the various photobleaching protocols, such as FRAP and FLIP, and will become even more profound as new developments in GFP technology become available.
-
As cellular process diagrams become more and more complex, it becomes essential to build databases for models as well as for experimental data. The new fields of integrative bioinformatics and pathway databases lie at the intersection of kinetic modelling and database technology.
-
The burgeoning fields of biological modelling, computational cell biology and systems biology will need a standard language for exchange of models among software tools. Nascent standards have been proposed in the form of the XML-based tools, CellML and SBML.
Abstract
The ability to visualize protein dynamics and biological processes by in vivo microscopy is revolutionizing many areas of biology. These methods generate large, kinetically complex data sets, which often cannot be intuitively interpreted. The combination of dynamic imaging and computational modelling is emerging as a powerful tool for the quantitation of biophysical properties of molecules and processes. The new discipline of computational cell biology will be essential in uncovering the pathways, mechanisms and controls of biological processes and systems as they occur in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lippincott-Schwartz, J., Snapp, E. & Kenworthy, A. Studying protein dynamics in living cells. Nature Rev. Mol. Cell Biol. 2, 444–456 (2001).A comprehensive review on kinetic imaging methods.
Periasamy, A. & Day, R. N. Visualizing protein interactions in living cells using digitized GFP imaging and FRET microscopy. Methods Cell Biol. 58, 293–314 (1999).
Wouters, F. S., Verveer, P. J. & Bastiaens, P. I. Imaging biochemistry inside cells. Trends Cell Biol. 11, 203–211 (2001).
Misteli, T. & Spector, D. L. Applications of the green fluorescent protein in cell biology and biotechnology. Nature Biotechnol. 15, 961–964 (1997).
Taylor, D. L. & Wang, Y. L. Molecular cytochemistry: incorporation of fluorescently labeled actin into living cells. Proc. Natl Acad. Sci. USA 75, 857–861 (1978).
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Patterson, G., Day, R. N. & Piston, D. Fluorescent protein spectra. J. Cell Sci. 114, 837–838 (2001).
Verkhusha, V. V. et al. An enhanced mutant of red fluorescent protein DsRed for double labeling and developmental timer of neural fiber bundle formation. J. Biol. Chem. 276, 29621–29624 (2001).
Griffin, B. A., Adams, S. R. & Tsien, R. Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Lippincott-Schwartz, J., Roberts, T. & Hirschberg, K. Secretory protein trafficking and organelle dynamics in living cells. Annu. Rev. Cell Dev. Biol. 16, 557–589 (2000).
Shima, D. T., Haldar, K., Pepperkok, R., Watson, R. & Warren, G. Partitioning of the Golgi apparatus during mitosis in living HeLa cells. J. Cell Biol. 137, 1211–1228 (1997).
Rudolf, R., Salm, T., Rustom, A. & Gerdes, H. H. Dynamics of immature secretory granules: role of cytoskeletal elements during transport, cortical restriction, and f-actin-dependent tethering. Mol. Biol. Cell 12, 1353–1365 (2001).
Ellenberg, J. et al. Nuclear membrane dynamics and reassembly in living cells: targeting of an inner nuclear membrane protein in interphase and mitosis. J. Cell Biol. 138, 1193–1206 (1997).
Moir, R. D., Yoon, M., Khuon, S. & Goldman, R. D. Nuclear lamins A and B1: different pathways of assembly during nuclear envelope formation in living cells. J. Cell Biol. 151, 1155–1168 (2000).
Dundr, M., Misteli, T. & Olson, M. O. J. The dynamics of postmitotic reassembly of the nucleolus. J. Cell Biol. 150, 433–446 (2000).
Misteli, T., Cáceres, J. F. & Spector, D. L. The dynamics of a pre-mRNA splicing factor in living cells. Nature 387, 523–527 (1997).
Platani, M., Goldberg, I., Swedlow, J. & Lamond, A. I. In vivo analysis of cajal body movement, separation, and joining in live human cells. J. Cell Biol. 151, 1561–1574 (2000).
Savino, T. M., Gebrane-Younes, J., De Mey, J., Sibarita, J. B. & Hernandez-Verdun, D. Nucleolar assembly of the rRNA processing machinery in living cells. J. Cell Biol. 153, 1097–1110 (2001).
Manders, E. M., Kimura, H. & Cook, P. R. Direct imaging of DNA in living cells reveals the dynamics of chromosome formation. J. Cell Biol. 144, 813–821 (1999).
McNally, J. G., Muller, W. G., Walker, D., Wolford, R. & Hager, G. L. The glucocorticoid receptor: rapid exchange with regulatory sites in living cells. Science 287, 1262–1265 (2000).
Robinett, C. et al. In vivo localization of DNA sequences and visualisation of large-scale chromatin organisation using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996). The first description of an experimental system to study a chromatin region in living cells.
Zink, D. et al. Structure and dynamics of human interphase chromosome territories in vivo. Hum. Genet. 102, 241–251 (1998).
Tsukamoto, T. et al. Visualisation of gene activity in living cells. Nature Cell Biol. 2, 871–878 (2000).
Thomas, C. F. & White, J. G. Four-dimensional imaging: the exploration of space and time. Trends Biotechnol. 16, 175–182 (1998).
Bornfleth, H., Edelmann, P., Zink, D., Cremer, T. & Cremer, C. Quantitative motion analysis of subchromosomal foci in living cells using four-dimensional microscopy. Biophys. J. 77, 2871–2886 (1999).
Bergsma, C. B., Streekstra, G. J., Smeulders, A. W. & Manders, E. M. Velocity estimation of spots in three-dimensional confocal image sequences of living cells. Cytometry 43, 261–272 (2001).
Tvarusko, W. et al. Time-resolved analysis and visualisation of dynamic processes in living cells. Proc. Natl Acad. Sci. USA 96, 7950–7955 (1999).
Gehrlich, D., Beaudouin, J., Gebhard, M., Ellenberg, J. & Eils, R. Four-dimensional imaging and quantitative reconstruction to analyse complex spatiotemporal processes in live cells. Nature Cell Biol. 3, 852–855 (2001).
Edidin, M., Zagyansky, Y. & Lardner, T. J. Measurement of membrane protein lateral diffusion in single cells. Science 191, 466–468 (1976).
Axelrod, D., Koppel, D. E., Schlessinger, J., Elson, E. & Webb, W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J. 16, 1055–1069 (1976).This is the classic paper on the quantitative analysis of FRAP data for cases in which the recovery is dominated by diffusion.
Reits, E. A. & Neefjes, J. J. From fixed to FRAP: measuring protein mobility and activity in living cells. Nature Cell Biol. 3, 145–147 (2001).
Cole, N. B. et al. Diffusion mobility of Golgi proteins in membranes of living cells. Science 273, 797–801 (1996).
Dittrich, P., Malvezzi-Campeggi, F., Jahnz, M. & Schwille, P. Accessing molecular dynamics in cells by fluorescence correlation spectroscopy. Biol. Chem. 382, 491–494 (2001).
Schwille, P., Haupts, U., Maiti, S. & Webb, W. W. Molecular dynamics in living cells observed by fluorescence correlation spectroscopy with one- and two-photon excitation. Biophys. J. 77, 2251–2265 (1999).
Wachsmuth, M., Waldeck, W. & Langowski, J. Anomalous diffusion of fluorescent probes inside living cell nuclei investigated by spatially-resolved fluorescence correlation spectroscopy. J. Mol. Biol. 298, 677–689 (2000).
Brock, R., Vamosi, G., Vereb, G. & Jovin, T. M. Rapid characterization of green fluorescent protein fusion proteins on the molecular and cellular level by fluorescence correlation microscopy. Proc. Natl Acad. Sci. USA 96, 10123–10128 (1999).
Rigler, R. et al. Specific binding of proinsulin C-peptide to human cell membranes. Proc. Natl Acad. Sci. USA 96, 13318–13323 (1999).
Politz, J. C., Browne, E. S., Wolf, D. E. & Pederson, T. Intranuclear diffusion and hybridization state of oligonucleotides measured by fluorescence correlation spectroscopy in living cells. Proc. Natl Acad. Sci. USA 95, 6043–6048 (1998).
Pramanik, A., Olsson, M., Langel, U., Bartfai, T. & Rigler, R. Fluorescence correlation spectroscopy detects galanin receptor diversity on insulinoma cells. Biochemistry 40, 10839–10845 (2001).
Widengren, J. & Rigler, R. Fluorescence correlation spectroscopy as a tool to investigate chemical reactions in solutions and on cell surfaces. Cell. Mol. Biol. (Noisy-le-grand) 44, 857–879 (1998).
Misteli, T. Protein dynamics: implications for nuclear architecture and gene expression. Science 291, 843–847 (2001).
Kimura, H. & Cook, P. R. Kinetics of core histones in living human cells: little exchange of H3 and H4 and some rapid exchange of H2B. J. Cell Biol. 153, 1341–1353 (2001).
Phair, R. D. & Misteli, T. High mobility of proteins in the mammalian cell nucleus. Nature 404, 604–60 (2000).An application of FRAP, FLIP and kinetic modelling to obtain quantitative measures of the mobility of several functionally distinct nuclear proteins.
Nehls, S. et al. Dynamics and retention of misfolded proteins in native ER membranes. Nature Cell Biol. 2, 288–295 (2000).
Adams, C. L., Chen, Y. T., Smith, S. J. & Nelson, W. J. Mechanisms of epithelial cell–cell adhesion and cell compaction revealed by high-resolution tracking of E-cadherin–green fluorescent protein. J. Cell Biol. 142, 1105–1119 (1998).
Pena, D., Tiao, G. C. & Tsay, R. S. A Course in Time Series Analysis (Wiley, New York, 2000).
Eriksson, K., Estep, D., Hansbro, P. & Johnson, C. Computational Differential Equations (Cambridge University, Cambridge, 1996).
Newton, I. The Method of Fluxions and Infinite Series. (Henry Woodfall, London, 1736).
McAdams, H. H. & Shapiro, L. Circuit simulation of genetic networks. Science 269, 650–656 (1995).
Brenan, K. E., Campbell, S. L. & Petzold, L. R. Numerical Solution of Initial-value Problems in Differential Algebraic Equations (Society for Industrial and Applied Mathematics, 1996).
Bhalla, U. S. & Iyengar, R. Emergent properties of networks of biological signaling pathways. Science 283, 381–387 (1999).
Firth, C. A. J. M. & Bray, D. Computational Modeling of Genetic and Biochemical Networks (eds Bower, J. M. & Bolouri, H.) 263–286 (MIT Press, Cambridge, 2001).
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Bassingthwaighte, J. B., Liebovitch, L. S. & West, B. J. Fractal physiology (Oxford University Press, New York, 1994).
Phair, R. D. Development of kinetic models in the non-linear world of molecular cell biology. Metabolism 46, 1489–1495 (1997).
Hedley, W. J., Nelson, M. R., Bullivant, D. P. & Nielsen, P. F. A short introduction to CellML. Phil. Trans. R. Soc. Lond. A 359, 1073–1089 (2001).
Hucka, M. et al. Foundations of Systems Biology (ed. Kitano, H.) (MIT Press, Cambridge, 2001).
Schaff, J. C., Slepchenko, B. M. & Loew, L. M. Physiological modeling with virtual cell framework. Methods Enzymol. 321, 1–23 (2000).
Fink, C. C. et al. Morphological control of inositol-1,4,5-trisphosphate-dependent signals. J. Cell Biol. 147, 929–936 (1999).A clear and compelling examination of the importance of partial differential equation models when studying large cells or cells with long processes in which one must account for simultaneous diffusion and spatially distributed chemical reactions.
Nocedal, J. & Wright, S. J. Numerical Optimisation (Springer, New York, 1999).
Hirschberg, K. et al. Kinetic analysis of secretory protein traffic and characterization of golgi to plasma membrane transport intermediates in living cells. J. Cell Biol. 143, 1485–1503 (1998).This was among the first studies to combine the power of green fluorescent protein chimaeras, photobleaching techniques and kinetic analysis to answer questions about protein transport in living cells.
Bagowski, C. P. & Ferrell, J. E. Jr Bistability in the JNK cascade. Curr. Biol. 11, 1–20 (2001).
Karp, P. D. Pathway databases: a case study in computational symbolic theories. Science 293, 2040–2044 (2001).
Onsager, L., Hemmer, P. C., Holden, H. & Ratkje, S. K. The Collected Works of Lars Onsager with Commentary (World Scientific, Singapore, 1996).
Segel, I. H. Enzyme Kinetics Behavior and Analysis of Rapid Equilibrium and Steady State Enzyme Systems (Wiley, New York, 1975).
Mayr, B. & Montminy, M. Transcriptional regulation by the phosphorylation–dependent factor CREB. Nature Rev. Mol. Cell Biol. 2, 599–609 (2001).
Author information
Authors and Affiliations
Related links
Related links
DATABASES
FURTHER READING
Simulation and Modelling Software: SAAM Institute
Online textbook
Databases
Standards
Glossary
- CONFOCAL MICROSCOPY
-
A microscopy method used to obtain a thin optical section through a specimen.
- MULTI-PHOTON MICROSCOPY
-
A microscopy method that uses the simultaneous absorbance of several low-energy electrons to generate an optical section through a specimen.
- FLUOROPHORE
-
A small molecule or a part of a larger molecule that can be excited by light to emit fluorescence.
- STEADY STATE
-
An open system, the content of which is held constant by a continuous input. Here, the output equals the input.
- DIFFUSION COEFFICIENT
-
A measure to characterize the speed with which a particular molecule moves in a particular medium when driven by random thermal agitation.
- PARAMETER
-
The numerical constant that determines the absolute speed of a process. A first-order process is characterized by a single parameter, the rate constant. A process that is governed by a Michaelis–Menten equation is characterized by two parameters, Vmax and Km.
- STOCHASTIC SYSTEMS
-
A dynamic system, the processes of which are characterized by a probability distribution. The stochastic system theory is particularly important when the abundance of molecules in a particular state falls below the deterministic limit, about 100 molecules per cell.
- FRACTALS
-
These are objects that provide more and more features as the resolution of the observation increases. These finer features show statistical self-similarity as seen in biological branching patterns, ion-channel currents and heart rate.
- CHAOS
-
A deterministic system (for example, some systems of nonlinear differential equations), the output of which seems random, but is not. Such systems show a surprising sensitivity to initial conditions.
- ANALYSIS OF VARIANCE
-
A statistical procedure for testing for differences among the means of several populations. It partitions the total sample variance among several specific sources to carry out the test on means.
- SUM OF EXPONENTIALS
-
An algebraic expression that is made up of exponentials. In a first-order system, the time-course solution for every state can be precisely mimicked by the sum of exponentials that correspond to the number of states in the system.
- POLYNOMIAL
-
Algebraic expressions that are made up of more than one term — for example, mx + b.
- SUM OF GAUSSIANS
-
An approximation by weighted sums of normal distributions, or Gaussians, each characterized by two parameters — a mean and a variance — to describe a data set.
- STATE
-
The generic name used here to identify those variables that change with time and for which differential equations are written.
- ALLOSTERIC REGULATION
-
A modification of a process by a molecule that binds to an enzyme or a transporter or another protein at a site other than its active, or catalytic, site.
- RATE LAWS
-
Algebraic expressions for the flux through a given pathway.
- PROCESS
-
The generic name for events that bring about changes in one or more states.
- EXTENSIBLE MARKUP LANGUAGE
-
A method for putting structured data in a text file so that applications receive not only unambiguous data but also unambiguous context. XML documents are not meant to be read, except by software.
- INTEGRATIVE BIOINFORMATICS
-
The intersection of kinetic modelling and database technology, a combination that becomes essential as cell biologists move to analyse larger and more complex molecular genetic control systems.
Rights and permissions
About this article
Cite this article
Phair, R., Misteli, T. Kinetic modelling approaches to in vivo imaging. Nat Rev Mol Cell Biol 2, 898–907 (2001). https://doi.org/10.1038/35103000
Issue Date:
DOI: https://doi.org/10.1038/35103000
This article is cited by
-
Retinal pigment epithelium degeneration caused by aggregation of PRPF31 and the role of HSP70 family of proteins
Molecular Medicine (2020)
-
N-glycosylation enables high lateral mobility of GPI-anchored proteins at a molecular crowding threshold
Nature Communications (2016)
-
Measurement of velocity fluctuations in microfluidics with simultaneously ultrahigh spatial and temporal resolution
Experiments in Fluids (2016)
-
The molecular size of the extra-membrane domain influences the diffusion of the GPI-anchored VSG on the trypanosome plasma membrane
Scientific Reports (2015)
-
Spatial heterogeneity of dynamics of H1 linker histone
European Biophysics Journal (2014)