Abstract
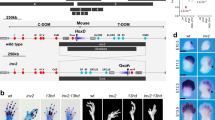
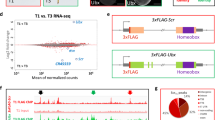
THE Hox genes encode transcription factors which mediate the formation of the mammalian body plan along the anteroposterior and appendicular axes1–15. Paralogous Hox genes within the separate linkage groups are closely related with respect to DNA sequence and expression16,17, suggesting that they could have at least partially redundant functions. We showed previously that mice homozygous for independent targeted disruptions in the paralogous genes hoxa-3 and hoxd-3 had no defects in common1,8. But our current analysis of double mutants has revealed strong, dosage-dependent interactions between these genes. We report here that in hoxd-3- homozygotes the first cervical vertebra, the atlas, is homeotically transformed to the adjacent anterior structure. Unexpectedly, in double mutants, rather than observing a more extensive homeotic transformation, the entire atlas is deleted. These observations are interpreted in terms of a model in which these Hox genes differentially regulate the proliferation rates of the appropriate sets of precursor cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chisaka, O. & Capecchi, M. R. Nature 350, 473–479 (1991).
Lufkin, T., Dierich, A., LeMeur, M., Mark, M. & Chambon, P. Cell 66, 1105–1119 (1991).
Chisaka, O., Musci, T. S. & Capecchi, M. Nature 355, 516–520 (1992).
LeMouellic, H., Lallemand, Y. & Brûlet, P. Cell 69, 251–264 (1992).
Ramirez-Solis, R., Zheng, H., Whiting, J., Krumlauf, R. & Bradley, A. Cell 73, 279–294 (1993).
Carpenter, E. M., Goddard, J. M., Chiska, O., Manley, N. R. & Capecchi, M. R. Development 118, 1063–1075 (1993).
Mark, M. et al. Development 119, 319–338 (1993).
Condie, B. G. & Capecchi, M. R. Development 119, 579–595 (1993).
Jeannotte, L., Lemieux, M., Charron, J., Poirier, F. & Robertson, E. J. Genes Dev. 7, 2085–2096 (1993).
Dollé, P. et al. Cell 75, 431–441 (1993).
Small, K. M. & Potter, S. S. Genes Dev. 7, 2318–2328 (1993).
Gendron-Maguire, M., Mallo, M., Zhang, M. & Gridley, T. Cell 75, 1317–1331 (1993).
Rijli, F. M. et al. Cell 75, 1333–1349 (1993).
Kostic, D. & Capecchi, M. R. Mech. Dev. (in the press).
Davis, A. P. & Capecchi, M. R. Development (in the press).
Hunt, P. et al. Nature 353, 861–864 (1991).
McGinnis, W. & Krumlauf, R. Cell 68, 283–302 (1992).
Aberdam, D., Negreanu, V., Sachs, L. & Blatt, C. Molec. cell. Biol. 11, 554–557 (1991).
Dollé, P. et al. Proc. natn. Acad. Sci. U.S.A. 90, 7666–7670 (1993).
Kessel, M. & Gruss, P. Cell 67, 89–104 (1991).
Mansour, S. L., Goddard, J. M. & Capecchi, M. R. Development 117, 13–28 (1993).
Jegalian, B. G. & DeRobertis, E. M. Cell 71, 901–910 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Condie, B., Capecchi, M. Mice with targeted disruptions in the paralogous genes hoxa-3 and hoxd-3 reveal synergistic interactions. Nature 370, 304–307 (1994). https://doi.org/10.1038/370304a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/370304a0
This article is cited by
-
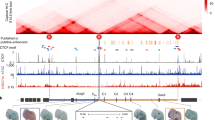
A multiple super-enhancer region establishes inter-TAD interactions and controls Hoxa function in cranial neural crest
Nature Communications (2023)
-
Inadvertent nucleotide sequence alterations during mutagenesis: highlighting the vulnerabilities in mouse transgenic technology
Journal of Genetic Engineering and Biotechnology (2021)
-
Loss of Hox5 function results in myofibroblast mislocalization and distal lung matrix defects during postnatal development
Science China Life Sciences (2018)
-
Hox Genes and Limb Musculoskeletal Development
Current Osteoporosis Reports (2014)
-
Hox genes and kidney development
Pediatric Nephrology (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.