Abstract
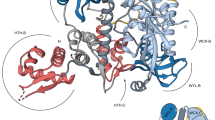
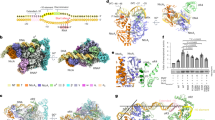
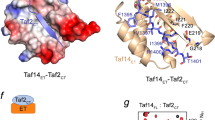
HAP1 is a member of a family of fungal transcription factors that contain a Zn2Cys6 binuclear cluster domain and bind as homodimers to sequences containing two DNA half sites. We have determined the 2.5 Å crystal structure of HAP1 bound to a cognate upstream activation sequence from the CYC7 gene. The structure reveals that HAP1 is bound in a dramatically asymmetric manner to the DNA target. This asymmetry aligns the Zn2Cys6 domains in a tandem head-to-tail fashion to contact two DNA half sites, positions an N-terminal arm of one of the protein subunits to interact with the inter-half site base pairs in the DNA minor groove, and suggests a mechanism by which DNA-binding facilitates asymmetric dimerization by HAP1. Comparisons with the DNA complexes of the related GAL4, PPR1 and PUT3 proteins illustrate how a conserved protein domain can be reoriented to recognize DNA half sites of different polarities and how homodimeric proteins adopt dramatically asymmetric structures to recognize cognate DNA targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Lalonde, B., Arcangioli, B. & Guarente, L. A single Saccharomyces Cerevisiae upstream activation site (UAS1) has two distinct regions essential for its activity Mol. Cell. Biol. 6, 4690 –4696 (1986).
Pfeifer, K., Arcangioli, B. & Guarente, L. Yeast HAP1 activator competes with factor RC2 for binding for the upstream activation site UAS1 of the CYC1 gene. Cell 49, 9–18 (1987).
Pfeifer, K., Prezant, T. & Guarente, L. Yeast HAP1 activator binds two upstream activation sites of different sequence Cell 49, 19–27 (1987).
Zitomer, R.S. et al. Elements involved in oxygen regulation of the Saccharomyces cerevisiae CYC7 gene. Mol. Cell. Biol. 7, 2212–2220 (1987).
Winkler, H. et al. C0-ordinate control of synthesis of mitochondrial and non-mitochondrial hemoproteins: a binding site for the HAP1(CYP1) protein in the UAS region of the yeat catalase T gene (CTT1). EMBO J. 7, 1799–1804 (1988).
Lodi, T. & Guiard, B. Complex transcriptional regulation of the Saccharomyces cerevisiae CYB2 gene encoding cytochrome b2: CYP1 (HAP1) activator binds to the CYB2 upstream activation site UAS1-B2. Mol. Cell. Biol. 11, 3762–3772 (1991).
Pfeifer, K., Kim, K.S., Kogan, S. & Guarente, L. Functional dissection and sequence of yeast HAP1 activator. Nature 342, 200–203 (1989).
Schjerling, P. & Holmberg, S. Comparative amino acid sequence analysis of the C6 zinc cluster family of transcriptional regulators. Nucleic Acids Res. 24, 4599–4607 (1996).
Gardner, K.H., Anderson, S.F. & Coleman, J.E. Solution structure of the Kluyveromyces lactis LAC9 Cd2Cys6 DNA-binding domain. Nature Struct. Biol. 2, 898–905 (1995).
Johnston, M. A model fungal gene regulatory mechanism: the GAL genes of Saccharomyces cerevisiae. Microbiol. Rev. 51, 458–476 (1987).
Marmorstein, R. & Harrison, S.C. Crystal structure of a PPR1-DNA complex: DNA recognition by proteins containing a Zn2Cys6 binuclear cluster. Genes Devel. 8, 2504–12 (1994).
Liang, S.D., Marmorstein, R., Harrison, S.C. & Ptashne, M. DNA sequence preferences of GAL4 and PPR1: how a subset of Zn2Cys6 binuclear cluster proteins recognize DNA. Mol. Cell. Biol. 16, 3773–3780 (1996).
Hellauer, K., Rochon, M.-H. & Turcotte, B. A novel DNA binding motif for yeast zinc cluster proteins: the Leu3p and Pdr3p transcriptional activators recognize everted repeats. Mol. Cell. Biol. 16, 6096–6102 (1996).
Swaminathan, K., Flynn, P., Reece, R.R. & Marmorstein, R. Crystal structure of a PUT3–DNA complex reveals a novel mechanism for DNA recognition by a protein containing a Zn2Cys6 binuclear cluster. Nature Struct. Biol. 4, 751–759 (1997).
Marmorstein, R., Carey, M., Ptashne, M. & Harrison, S.C. DNA recognition by GAL4: structure of a protein–DNA complex. Nature 356, 408–414 (1992).
Zhang, L. & Guarente, L. The yeast activator HAP1 — a GAL4 family member — binds DNA in a directly repeated orientation. Genes Devel. 8, 2110–2119 (1994).
Turcotte, B. & Guarente, L. HAP1 positive control mutants specific for one of two binding sites. Genes Devel. 6, 2001–2009 (1992).
Ellenberger, T.E., Brandl, C.J., Struhl, K. & Harrison, S.C. The GCN4 basic-region leucine zipper binds DNA as a dimer of uninterrupted α-helices: crystal structure of the protein–DNA complex. Cell 71, 1223–123– (1992).
Cohen, C. & Parry, D.A. Alpha-helical coiled coils and bundles: how to design an alpha-helical protein. Proteins 7, 1–15 (1990).
O'Shea, E.K., Klemm, J.D., Kim, P.S. & Alber, T. X-ray structure of the GCN4 leucine zipper, a two stranded parallel coiled-coil. Science 254, 539–544 (1991).
Zhang, L. & Guarente, L. The C6 zinc cluster dictates asymmetric binding by HAP1. EMBO J. 15, 4676–4681 (1996).
Baleja, J.D., Marmorstein, R., Harrison, S.C. & Wagner, G. Solution structure of the DNA-binding domain of Cd2-GAL4 from S. cerevisiae. Nature 356, 405–453 (1992).
Kraulis, P.J., Raine, A.R.C., Gadhavi, P.L. & Laue, E.D. Structure of the DNA binding domain of zinc GAL4. Nature 356, 448–450 (1992).
Walters, K., Dayie, K.T., Reece, R.J., Ptashne, M. & Wagner, G. Structure and mobility of the PUT3 dimer: a DNA pincer. Nature Struct. Biol. 4, 744–750 (1997).
Creusot, F., Verdiere, J., Gaisne, M. & Slonimski, P.P. CTP1 (HAP1) regulation of oxygen-dependent gene expression in yeast. J. Mol. Biol. 204, 423–432 (1988).
Timmerman, J. et al. 1H, 15N Resonance assignment and three-dimensional structure of CYP1 (HAP1) DNA-binding domain. J. Mol. Biol. 259, 792–804 (1996).
Ha, N., Hellaur, K. & Turcotte, B. Mutations in target DNA elements of yeast HAP1 modulates transcriptional activity without affecting DNA binding. Nucleic Acids Res. 24, 1453–1459 (1996).
Otting, G. et al. Protein–DNA contacts in the structure of a homeodomain–DNA complex determined by nuclear magnetic resonance spectroscopy in solution EMBO J. 9, 3085–3092 (1990).
Kissinger, C., Liu, B., Martin-Blanco, E., Kornberg, T. & Pabo, C. Crystal Structure of an engrailed homeodomain–DNA complex at 2.8 Å resolution: a framework for understanding homeodomain–DNA interactions. Cell 63, 579–590 (1990).
Wolberger, C., Vershon, A., Liu, B., Johnson, A. & Pabo, C. Crystal stucture of a MATα2 homeodomain-operator complex suggests a general model for homeodomain–DNA interactions Cell 67, 517–528 (1991).
Kim, K.-S. & Guarente, L. Mutations that alter transcriptional activation but not DNA binding in the zinc finger of yeast activator HAP1. Nature 343, 200–203 (1989).
Lefstin, J., Thoma, J. & Yamamoto, K. Influence of a steroid receptor DNA-binding domain on transcriptional regulatory functions Genes Devel. 8, 2842–2856 (1994).
Reece, R.J. & Ptashne, M. Determinants of binding site specificity among yeast C6 zinc cluster proteins. Science 261, 909–911 (1993).
Prezant, T., Pfeifer, K. & Guarente, L. Organization of the regulatory region of the yeast CYC7 gene: Multiple factors are involved in regulation. Mol. Cell. Biol. 7, 3252–3259 (1987).
Towers, T., Luisi, B., Asianov, A. & Freeman, L. DNA target selectivity by the vitamin D3 receptor: mechanism of dimer binding to an asymmetric repeat element. Proc. Natl. Acad. Sc. USA 90, 6310–6314 (1993).
Mader, S. et al. The patterns of binding of RAR, RXR and TR homo- and heterodimers to direct repeats are dictated by the binding specificities of the DNA binding domains EMBO J. 12, 5029–5041 (1993).
Perlmann, T., Rangarajan, P., Umesono, K. & Evans, R. Determinants for selective RAR and TR recognition of direct repeat HREs. Gene. Dev. 7, 1411–1422 (1993).
Kurokawa, R. et al. Differential orientations of the DNA-binding domain and carboxy-terminal dimerization interface regulate binding site selection by nuclear receptor heterodimers. Genes Devel. 7, 1423–1435 (1993).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A50, 157–163 (1994).
Kabsch, W. Evaluation of single-crystal X-ray diffraction data from a position-sensitive detector. J. Appl. Crystallogr. 21, 916–924 (1988b).
Kabsch, W. Automatic indexing of rotation diffraction patterns. J. Appl. Crystallogr. 21, 67–71 (1988a).
King, D.A., Zhang, L., Guarente, L. & Marmorstein, R. Structure of a HAP1-18/DNA complex reveals that protein/DNA interactions can have direct allosteric effects on transcriptional activation. Nature Struct. Biol. 6, 22–27 (1999).
Brünger, A.T. X-PLOR 3.1, A system for X-ray crystallography and NMR. (Yale University Press, New Haven, Connecticut; 1992).
Brünger, A.T. The free R value: a novel statistical quantity assessing the accuracy of crystal structures. Nature 335, 472–474 (1992).
Brünger, A.T. & Krukowski, A. Slow-cooling protocols for crystallographic refinment by simulated annealing. Acta Crystallogr. A46, 585–593 (1990).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallographic R factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Kleywegt, G. & Brünger, A. Checking your imagination: Applications of the free R value Structure 4, 897–904 (1996).
Read, R. Improved Fourier coefficients for maps using phases from partial structures with errors Acta. Crystallgor. A42, 140–149 (1986).
Jones, T.A. A graphics model building and refinement system for macromolecules. J. Appl. Crystallogr. 11, 268–272 (1978).
Rice, L.M. & Brünger, A.T. Torsion angle dynamics: Reduced variable conformational sampling enhances crystallographic structure refinement. Proteins 19, 277–290 (1994).
Jiang, J.S. & Brünger, A.T. Protein hydration observed by X-ray diffraction: solvation properties of penicillopepsin and neuraminidase crystal structures. J. Mol. Biol. 243, 100–115 (1994).
Hodel, A., Kim, S.-H. & Brünger, A.T. Model bias in macromolecular crystal structures. Acta Crystallogr. A48, 851–858 (1992).
Kraulis, P.J. MOLSCRIPT: A program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallgr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. RASTER3D version 2.0: a program for photorealistic molecular graphics. Acta Crystallogr. D50, 869–873 (1994).
Nicholls, A., Sharp, K. & Honig, B. Protein folding and association: insights from interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank A. Lukens, T. Stams, C. Lesburg, X. Li, Y. Mo, S. Benson, R. Venkataramani, and K. Swaminathan for useful discussions. This work was supported by a grant form the NIH and a junior faculty research award from the ACS to R.M.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
King, D., Zhang, L., Guarente, L. et al. Structure of a HAP1–DNA complex reveals dramatically asymmetric DNA binding by a homodimeric protein. Nat Struct Mol Biol 6, 64–71 (1999). https://doi.org/10.1038/4940
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/4940
This article is cited by
-
Similarity regression predicts evolution of transcription factor sequence specificity
Nature Genetics (2019)
-
Zinc regulates ERp44-dependent protein quality control in the early secretory pathway
Nature Communications (2019)
-
Analysis of the HIV-2 protease’s adaptation to various ligands: characterization of backbone asymmetry using a structural alphabet
Scientific Reports (2018)
-
CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
Nature Communications (2017)
-
MlcR, a zinc cluster activator protein, is able to bind to a single (A/T)CGG site of cognate asymmetric motifs in the ML-236B (compactin) biosynthetic gene cluster
Molecular Genetics and Genomics (2009)