Abstract
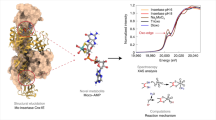
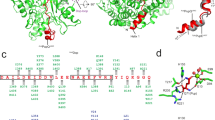
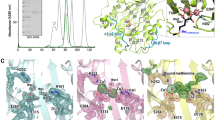
Molybdenum cofactor (Moco) biosynthesis is an evolutionarily conserved pathway present in eubacteria, archaea and eukaryotes, including humans. Genetic deficiencies of enzymes involved in Moco biosynthesis in humans lead to a severe and usually fatal disease. Moco contains a tricyclic pyranopterin, termed molybdopterin (MPT), that bears the cis-dithiolene group responsible for molybdenum ligation. The dithiolene group of MPT is generated by MPT synthase, which consists of a large and small subunits. The 1.45 Å resolution crystal structure of MPT synthase reveals a heterotetrameric protein in which the C-terminus of each small subunit is inserted into a large subunit to form the active site. In the activated form of the enzyme this C-terminus is present as a thiocarboxylate. In the structure of a covalent complex of MPT synthase, an isopeptide bond is present between the C-terminus of the small subunit and a Lys side chain in the large subunit. The strong structural similarity between the small subunit of MPT synthase and ubiquitin provides evidence for the evolutionary antecedence of the Moco biosynthetic pathway to the ubiquitin dependent protein degradation pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rajagopalan, K.V. Adv. Enzymol. 64, 215–290 (1991).
Rajagopalan, K.V. & Johnson, J.L. J. Biol. Chem. 267, 10199–10202 (1992).
Kisker, C., Schindelin, H. & Rees, D.C. Annu. Rev. Biochem. 66, 233–267 (1997).
Rajagopalan, K.V. Biochem. Soc. Trans. 25, 757–761 (1997).
Rajagopalan, K.V. In Escherichia coli and Salmonella typhimurium — molecular and cellular biology, Vol. 1 (ed. Neidhardt, F.C.) 674–679 (ASM Press, Washington DC; 1996).
Reiss, J. Christensen, E., Kurlemann, G., Zabot, M.-T. & Dorche, C. Hum. Genet. 103, 639–644 (1998).
Reiss, J. et al. Nature Genet. 20, 51–53 (1998).
Reiss, J. et al. Am. J. Hum. Genet. 64, 706–711 (1999).
Wuebbens, M.M. & Rajagopalan, K.V. J. Biol. Chem. 270, 1082–1087 (1995).
Rieder, C. et al. Eur. J. Biochem. 255, 24–36 (1998).
Pitterle, D.M. & Rajagopalan, K.V. J. Biol. Chem. 268, 13499–13505 (1993).
Pitterle, D.M., Johnson, J.L. & Rajagopalan, K.V. J. Biol. Chem. 268, 13506–13509 (1993).
Joshi, M.S., Johnson, J.L. & Rajagopalan, K.V. J. Bacteriol. 178, 4310–4312 (1996).
Hasona, A., Ray, R.M. & Shanmugam, K.T. J. Bacteriol. 180, 1466–1472 (1998).
Holm, L. & Sander, C. Trends Biochem. Sci. 20, 478–480 (1995).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281–296 (1991).
Begley, T.P., Xi, J., Kinsland, C., Taylor, S. & McLafferty, F. Curr. Opin. Chem. Biol. 3, 623–629 (1999).
Palenchar, P.M., Buck, C.J., Cheng, H., Larson, T.J. & Mueller, E.G. J. Biol. Chem. 275, 8283–8286 (2000).
Wang, C., Xi, J., Begley, T.P. & Nicholson, L.K. Nature Struct. Biol. 8, 47–51 (2000).
Rivers, S.L., McNairn, E., Blasco, F., Giordano, G. & Boxer, D.H. Mol. Microbiol. 8, 1071–1081 (1993).
Nohno, T., Kasai, Y. & Saito, T. J. Bacteriol. 170, 4097–4102 (1988).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Bailey, S. Acta Crystallogr.D 50, 760–763 (1994).
DeLaFortelle, E. & Bricogne, G. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G.W. Acta Crystallogr. D 52, 30–42 (1996).
Perrakis, A., Morris, R. & Lamzin, V.S. Nature. Struct. Biol. 6, 458–463 (1999).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Murshudov, G., Vagin, A. & Dodson, E. Acta Crystallogr. D 53, 240–255 (1997).
Hutchinson, E.G. & Thornton, J.M. Protein Sci. 3, 2207–2216 (1994).
Barton, G.J. Protein Eng. 6, 37–40 (1993).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 276, 505–524 (1997).
Acknowledgements
Supported by National Institutes of Health Grants to H.S. and K.V.R. M.W. would like to thank H. Sage and R. Stevens (Duke University) for performing sedimentation equilibrium centrifugation and mass spectrometry experiments. The National Synchrotron Light Source at Brookhaven is supported by the United States Department of Energy and National Institutes of Health, and beamline X26C is supported in part by the State University of New York at Stony Brook and its Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rudolph, M., Wuebbens, M., Rajagopalan, K. et al. Crystal structure of molybdopterin synthase and its evolutionary relationship to ubiquitin activation. Nat Struct Mol Biol 8, 42–46 (2001). https://doi.org/10.1038/83034
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/83034
This article is cited by
-
SAXS-guided Enhanced Unbiased Sampling for Structure Determination of Proteins and Complexes
Scientific Reports (2018)
-
Evolution of the archaeal and mammalian information processing systems: towards an archaeal model for human disease
Cellular and Molecular Life Sciences (2017)
-
Cleavage of the moaX-encoded fused molybdopterin synthase from Mycobacterium tuberculosis is necessary for activity
BMC Microbiology (2015)
-
Functional Identification of a Novel Gene, moaE, for 3-Succinoylpyridine Degradation in Pseudomonas putida S16
Scientific Reports (2015)
-
Nmag_2608, an extracellular ubiquitin-like domain-containing protein from the haloalkaliphilic archaeon Natrialba magadii
Extremophiles (2012)