Abstract
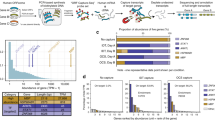
The genome sequences of Caenorhabditis elegans, Drosophila melanogaster and Arabidopsis thaliana have been predicted to contain 19,000, 13,600 and 25,500 genes, respectively1,2,3. Before this information can be fully used for evolutionary and functional studies, several issues need to be addressed. First, the gene number estimates obtained in silico and not yet supported by any experimental data need to be verified. For example, it seems biologically paradoxical that C. elegans would have 50% more genes than Drosophilia. Second, intron/exon predictions need to be tested experimentally. Third, complete sets of open reading frames (ORFs), or “ORFeomes,”4 need to be cloned into various expression vectors. To address these issues simultaneously, we have designed and applied to C. elegans the following strategy. Predicted ORFs are amplified by PCR from a highly representative cDNA library4 using ORF-specific primers, cloned by Gateway recombination cloning4,5,6 and then sequenced to generate ORF sequence tags (OSTs) as a way to verify identity and splicing. In a sample (n=1,222) of the nearly 10,000 genes predicted ab initio (that is, for which no expressed sequence tag (EST) is available so far), at least 70% were verified by OSTs. We also observed that 27% of these experimentally confirmed genes have a structure different from that predicted by GeneFinder. We now have experimental evidence that supports the existence of at least 17,300 genes in C. elegans. Hence we suggest that gene counts based primarily on ESTs may underestimate the number of genes in human and in other organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Adams, M.D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Walhout, A.J.M. et al. Gateway recombinational cloning: application to the cloning of large numbers of open reading frames, or ORFeomes. Methods Enzymol. 328, 575–592 (2000).
Walhout, A.J.M. et al. Protein interaction mapping in C. elegans using proteins involved in vulval development. Science 287, 116–122 (2000).
Hartley, J.L., Temple, F.T. & Brasch, M.A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
Hill, A.A., Hunter, C.P., Tsung, B.T., Tucker-Kellogg, G. & Brown, E.L. Genomic analysis of gene expression in C. elegans. Science 290, 809–812 (2000).
Gopal, S. et al. Homology-based annotation yields 1,042 new candidate genes in the Drosophila melanogaster genome. Nature Genet. 27, 337–340 (2001).
Ewing, B. & Green, P. Analysis of expressed sequence tags indicates 35,000 human genes. Nature Genet. 25, 232–234 (2000).
Dunham, I. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Roest Crollius, H. et al. Estimate of human gene number provided by genome-wide analysis using Tetraodon nigroviridis DNA sequence. Nature Genet. 25, 235–238 (2000).
Liang, F. et al. Gene Index analysis of the human genome estimates approximately 120,000 genes. Nature Genet. 25, 239–240 (2000); correction: 26, 501 (2000).
Acknowledgements
We thank S. Boulton, L. Matthews, J. Polanowska, M. Tewari and A.J.M. Walhout for comments on the manuscript and discussions; and L. Hillier and P. Green for the primer design program (OSP). This work was supported by grants from CREST, Japan Science and Technology Corporation and Grant-in-Aid for Scientific Research on Priority Areas from the Ministry of Education, Science, Sports and Culture of Japan (to Y.K.), and by grants 1 RO1 HG01715-01 from the National Human Genome Research Institute, 1 R21 CA81658 A 01 from the National Cancer Institute and 128 from the Merck Genome Research Institute (to M.V.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Reboul, J., Vaglio, P., Tzellas, N. et al. Open-reading-frame sequence tags (OSTs) support the existence of at least 17,300 genes in C. elegans. Nat Genet 27, 332–336 (2001). https://doi.org/10.1038/85913
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/85913
This article is cited by
-
Live imaging of cellular dynamics during Caenorhabditis elegans postembryonic development
Nature Protocols (2012)
-
Genome-wide functional annotation and structural verification of metabolic ORFeome of Chlamydomonas reinhardtii
BMC Genomics (2011)
-
A genome-wide study of PDZ-domain interactions in C. elegans reveals a high frequency of non-canonical binding
BMC Genomics (2010)
-
Metabolic network analysis integrated with transcript verification for sequenced genomes
Nature Methods (2009)
-
Characterizing the transcriptional regulation of let-721, a Caenorhabditis elegans homolog of human electron flavoprotein dehydrogenase
Molecular Genetics and Genomics (2009)