Abstract
Molecular pathophysiology of facioscapulohumeral muscular dystrophy (FSHD) involves the heterozygous contraction of the number of tandemly repeated D4Z4 units at chromosome 4q35.2. FSHD is associated with a range of 1–10 D4Z4 units instead of 11–150 in normal controls. Several factors complicate FSHD molecular diagnosis, especially the cis-segregation of D4Z4 contraction with a 4qA allele, whereas D4Z4 shortening is silent both on alleles 4qB and 10q. Discrimination of pathogenic 4q-D4Z4 alleles from highly homologous 10q-D4Z4 arrays requires the use of the conventional Southern blot, which is not suitable at the single-cell level. Preimplantation genetic diagnosis (PGD) is a frequent request from FSHD families with several affected relatives. We aimed to develop a rapid and sensitive PCR-based multiplex approach on single cells to perform an indirect familial segregation study of pathogenic alleles. Among several available polymorphic markers at 4q35.2, the four most proximal (D4S2390, D4S1652, D4S2930 and D4S1523, <1.23 Mb) showing the highest heterozygote frequencies (67–91%) were selected. Five recombination events in the D4S2390-D4S1523 interval were observed among 144 meioses. In the D4S2390-D4Z4 interval, no recombination event occurred among 28 FSHD meioses. Instead, a particular haplotype segregated with both clinical and molecular status, allowing the characterization of an at-risk allele in each tested FSHD family (maximal LOD score 2.98 for θ=0.0). This indirect protocol can easily complement conventional techniques in prenatal diagnosis. Although our multiplex PCR-based approach technically fulfils guidelines for single-cell analysis, the relatively high recombination risk hampers its application to PGD.
Similar content being viewed by others
Introduction
Facioscapulohumeral muscular dystrophy (FSHD1A, OMIM#158900) is the third most common hereditary muscle disease after Duchenne muscular dystrophy and myotonic dystrophy, with an estimated prevalence of 1 in 15 000. FSHD usually begins in adulthood with asymmetrical weakness and wasting of specific muscles of the face, shoulder girdle, upper and lower limbs. Extra-muscular symptoms such as mild hearing loss and asymptomatic retinal telangiectasia, as well as infantile and severe forms, can be observed. This dominantly inherited disorder manifests almost complete penetrance (>95%) after 20 years of age, but at least 5% of carriers remain asymptomatic.1, 2
Several linkage analysis studies have mapped the FSHD1A locus at the 4q35.2 subtelomeric region, which contains a polymorphic macrosatellite D4Z4 repeat consisting of 3.3-kb KpnI units varying in numbers between 11 and 150 copies. FSHD is commonly associated with the heterozygous contraction of 4q-D4Z4 arrays between 1 and 11 repeats.3, 4, 5, 6, 7, 8 The smallest D4Z4 arrays are roughly correlated with the earliest and the most severe disease forms.9, 10, 11, 12, 13 The subtelomere of chromosome 10q also contains a D4Z4-like repeat; however, despite the 98% homology between the D4Z4 repeats at 4q35 and 10q26, and the equal frequency of translocations observed between these two regions, FSHD has been uniquely associated with contractions on chromosome 4.14, 15 The significant level of somatic mosaicism (>40% in de novo cases) and the high incidence of new mutations (up to 30% of cases) result from these interchromosomal repeat interactions occurring during early embryogenesis. In addition, the large variability of the severity and age of onset even within families12, 16 might be related to the proportion of mosaic cells.17, 18, 19
As monosomy of 4qter does not lead to FSHD, the etiopathogenic mechanism is not likely to be haploinsufficiency for the repeats or for a nearby gene.20 D4Z4 contraction may lead to FSHD through complex association of epigenetic modifications (hypomethylation, histone acetylation and interaction with nuclear matrix) and/or transcriptional modulation of cis- or trans-acting genes.21, 22, 23, 24
Facioscapulohumeral muscular dystrophy-sized D4Z4 repeats are exclusively associated with the 4qA allele of a 4qA/4qB biallelic variation distal to D4Z4.25, 26, 27 Further complexity comes from additional polymorphisms specifying several haplotypes, and a recent study showed that FSHD is restricted to haplotype 4qA161.28
D4Z4 repeat counting and chromosome assignment are obtained through the conventional heavy and tricky Southern blot analysis. The digested genomic DNAs are separated on linear (LGE) or pulsed field gel electrophoresis (PFGE), and then hybridized with specific probes (p13E-11, D4Z4 or the 4qA plus 4qB).29, 30 Non-pathogenic deletions (carried by non-4qA161 alleles), somatic mosaicism or complex 4q-10q D4Z4 array exchanges (hybrid arrays) are at risk of leading to false-negative results.29 Hence, the molecular diagnosis usually implies very cautious interpretation of data and is offered by a few numbers of specialized laboratories. Moreover, as this reference method requires large amounts of genomic DNA and up to several weeks of diagnosis delay, it is not suitable for the preimplantation genetic diagnosis (PGD) and even for some cases of prenatal diagnosis (PND).
On the basis of a rapid one-step multiplex PCR protocol allowing robust segregation analysis of highly informative markers proximal to the D4Z4 locus, we propose a newly designed strategy for indirect FSHD molecular diagnosis. Such an approach could be useful in some complex PND situations. This study was primarily intended to test the accuracy of this PCR-based approach at the single-cell level and to estimate its feasibility in PGD.
Materials and methods
Subjects and families
Facioscapulohumeral muscular dystrophy-affected families were recruited through genetic consultation in the ‘Centre de réference des Maladies Neuromusculaires et de la sclérose latérale amyotrophique’, Marseille, France, according to the following inclusion criteria: a pedigree corresponding to two or three generations, at least one affected proband, and confirmed clinical and molecular status of all the tested individuals. Diagnosis of FSHD was based on the identification and sizing of the 4qD4Z4 repeats by Southern blot. Sporadic and mosaic cases, individuals under 18 and pregnant women were not included. Two blood samples per FSHD family member were collected: one with ethylenediamine tetraacetic acid for genomic DNA extraction and the second with sodium heparinate or sodium citrate (ACD, containing sodium citrate, citric acid and dextrose, BD Diagnostics, Le Pont-de-Claix, France) for single lymphocyte isolation. Informed consent was obtained from each individual included in this study.
Furthermore, 46 unrelated non-affected controls and 80 members of 6 large control families from the laboratory collection registered at the French Research Ministry (# DC-2008-417) were analysed anonymously. Genomic DNA was obtained with the FlexiGene DNA Kit (Qiagen, Courtaboeuf, France) according to the manufacturer's recommendations.
The genotypes of the selected markers available at the CEPH Fondation Jean Dausset database (http://www.cephb.fr/fr/index.php) for members of four CEPH pedigrees (52 meioses) were included in the study. Our study fulfilled the ethical rules of all the institutions involved.
Single lymphocytes
(In France, the 2004 Bioethic Law does not authorize pre-clinical research on blastomeres.) Set-up and validation steps of PCR-based molecular approaches at the single-cell level were performed on isolated lymphocytes obtained as previously described.31 Briefly, lymphocytes were separated from fresh blood samples using a separation medium (LSM, Eurobio, Les Ulis, France) and resuspended in an appropriate cell medium. Single lymphocytes were sorted manually under an inverted microscope and heated for 10 min at 65°C in 3 μl of lysis solution (200 mM KOH, 50 mM DTT),32 and then used immediately for PCR or stored at −20°C.
Multiplex PCR
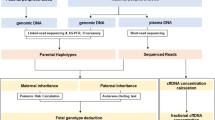
Fourteen microsatellite markers were initially selected according to their position on the physical map from the Human Genome Browser Gateway at UCSC (http://genome.ucsc.edu/cgi-bin/hgGateway) (Figure 1). Primers suitable for multiplex PCR were carefully designed using Primer3 software (http://frodo.wi.mit.edu/), and the reverse primers were 5′-labelled with the 6-FAM fluorochrome (MWG BIOTECH, Ebersberg, Germany). For the four markers further retained for single-cell analysis, the primer sequences, product sizes and amplification conditions are described in Table 1. PCR amplifications were carried out according to the manufacturer's instructions (Qiagen, Courtaboeuf, France), using the QIAGEN Multiplex PCR Kit, plus 3–15 pmoles of each primer and 100 ng of DNA, without Q-solution, in an Applied Biosystems 2720 Thermal Cycler or a GeneAmp PCR system 9700 (Applied Biosystems, Foster City, CA, USA) at 60°C for 35 cycles (standard DNA) or 40 cycles (single lymphocyte DNA).
Schematic representation of the 4qter region. For each marker, we have reported (1) the physical distance to the D4Z4 locus in Mb (megabases) (according to http://genome.ucsc.edu/cgi-bin/hgGateway), (2) the positions in cM (according to the Marshfield genetic map), (3) the estimated heterozygote frequencies for the four selected markers (bold underlined) and (4) the heterozygote frequencies from the CEPH database (http://www.cephb.fr/fr/cephdb/). The estimated heterozygote frequencies were low for the D4S2283 and near zero for the newly described markers (BF992518, 17xTG, BG167386, BC087857, D4F104S1 and 22xTG). (*) The D4F104S1 that overlaps the SSLP described by Lemmers et al.28 displayed complex electrophoresis patterns (not shown) with more than two alleles. Het, heterozygote frequencies; CEN, centromer; TEL, telomer.
The validation of the one-step quadruplex PCR protocol was performed on 720 lymphocytes including 202 from control subjects and 518 from the members of five FSHD families. Negative controls with water and blank samples (3 μl lysis solution without cell) were included in each PCR batch to assess the contamination rate.
Genotyping analysis
Sample mixes (0.5–1 μl PCR products plus 18.5 μl deionized formamide and 0.7 μl of LIZ-labelled size standard GS600) were injected during 5–10 s in an ABI 3130xl Genetic Analyser (Applied Biosystems) and electrophoresed at 55°C, 15 kV, during 2800 s using a POP6 polymer (Applied Biosystems). PCR product sizing and allele identification and assignment were performed using GeneMapper 4.0 software (Applied Biosystems).
The most likely haplotypes were determined by the ‘GNAlogique’ software, developed in our laboratory by Christophe Béroud and Dalil Hamroun (23 May 2007) and freely available at http://www.genetique-iurc-u827.fr/.
Linkage analysis
LOD scores and recombination fractions were determined using the LINKAGE Analysis Package (http://linkage.rockefeller.edu/ott/linkhelp.htm, New York, NY, USA).33
Results
Preliminary study of microsatellite markers on DNA extracted from peripheral blood
A first set of 14 microsatellite markers was studied, all mapped proximally to the D4Z4 locus (0.12 Mb for the 22xTG up to 1.88 Mb for the D4S2688 and D4S426) (Figure 1). Eight markers (D4S2688, D4S426, D4S2299, D4S2390, D4S1652, D4S2930, D4S2283 and D4S1523) are already described microsatellites (GeneBank, GDB, UCSC…). Six other putative dinucleotide repeats (BF992518, 17xTG, BG167386, BC087857, 22xTG and D4F104S1) were detected by the systematic exploration of all sequences available in the public databases. Unfortunately, there is no polymorphic microsatellite within the CHLC.GCT10C02 STS marker, 7 kb distal to the D4Z4 locus. Amplifications were set up on 100 ng control DNA and allowed an initial estimation of heterozygosity rates for the 14 markers (data not shown). Four microsatellites (D4S2688, D4S426, D4S2299 and D4S2283) and the six new dinucleotide repeats (BF992518, 17xTG, BG167386, BC087857, 22xTG and D4F104S1) were discarded for at least one of the following reasons: distance to the disease locus (>1.23 Mb), poor amplification and electropherogram profile in multiplex conditions or low informativity (<60%) (data not shown). Finally, the heterozygote frequencies were estimated after quadruplex amplification of the selected locus (D4S2390, D4S1652, D4S2930 and D4S1523) from an additional set of 46 unrelated controls. The high heterozygote frequencies observed (from 67 up to 91%) were quite similar to those obtained in CEPH families (http://www.cephb.fr/fr/index.php, Figure 1).
Segregation analysis of the four polymorphic markers was performed in FSHD families without any a priori knowledge of the FSHD clinical and molecular status. In each family, a specific haplotype segregated with both clinical and D4Z4 short alleles, previously sized by conventional Southern blot, and allowed the indirect identification of the at-risk chromosome. The D4S1523 marker was linked to the D4Z4 locus with a maximum LOD score of 2.98 for θ=0.00.
Validation of the quadruplex PCR approach on 720 single cells
The one-step multiplex amplification of the four selected loci (D4S2390, D4S1652, D4S2930 and D4S1523) was adapted to DNA from single cells. Positive amplifications for all 4 loci were obtained in 97.6% of cells (703/720; Table 2) from which the number of ADOs (corresponding to the random non-amplification of one of the alleles present in a heterozygous sample) observed in at least one locus was 7 among 703 cells (1%). Complete genotypes at the 4 loci were reached for 696 cells among 703 positive amplifications (99%) (Table 2).
Evaluation of recombination rates
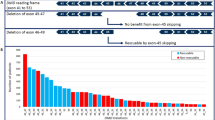
Recombination rates were estimated from 144 meioses: 28 meioses from the 12 FSHD families, 64 from the 6 control families and 52 from 4 CEPH families (Table 3). Using the linkage package, a recombination fraction of 0.021 was found between the D4S1523 and the D4S2390 loci. No recombination event was observed in the FSHD families. Five recombination events were scored, and surprisingly, all of them occurred in a single control family where 12 meioses were analysed (Figure 2, individuals II:5, II:6 and III:3). The subject II:6 carried a paternal and a maternal recombination (Figure 2). A double recombination would explain a homozygous 139 bp allele at the D4S1652 locus for the subject II:5. Sample mix-up or genotyping errors were excluded after re-genotyping from new dilutions of the genomic DNA. Moreover, no evidence of non-paternity was detected in this family following segregation analysis of 15 additional markers, distributed over 13 autosomes, as well as the amelogenin gene from the sexual chromosomes (data not shown). Third, gene dosage performed by semiquantitative PCR showed two alleles at this locus and thus excluded loss of heterozygosity (data not shown).
Discussion
Besides the large variation of clinical severity and age of onset, FSHD manifests an almost complete penetrance after the age of 20 years and prognosis in terms of mobility loss, pain and fatigue might be pejorative. Moreover, there is no specific therapeutic and some FSHD patients experience a very poor quality of life, which often worsens with age. With a 50% chance of having affected offspring, D4Z4 contraction carriers and affected parents may request PGD as an alternative to termination of pregnancy after PND. Standard molecular testing for FSHD (Southern blot and PFGE) is not applicable to PGD, which is usually performed on single blastomere DNA sampled from a 6- to 8-cell embryo, and in a limited time frame of 24 h at day 3 after intracytoplasmic sperm injection. Thus, PCR-based methods are mandatory for single-cell genetic analysis.
Although counting D4Z4 repeats by long PCR seems promising in terms of cost, specificity and rapidity (14 h), it suffers from a number of weaknesses. In particular, it is unsuitable for PGD because it requires large amounts of genomic DNA (>500 ng).34 Moreover, the absence of PCR amplicons might be observed in various situations: amplification failure, large FSHD-sized alleles (>18.4 kb, ie, >5 repeats) frequent in Caucasian patients, non-carrier individuals (>11 repeats), or p13E-11 deletion including the upstream primer (3% of FSHD cases).11, 30, 35 In addition, false-positive amplification of the short allele occurs in asymptomatic mosaic carriers.36 Thus, this approach constitutes a first-step screening method and Southern blotting is still required in at least 10% of FSHD cases.34 In addition, the UCSC in Silico PCR gave no match with the long-range primers and The Human BLAT Search mapped the reverse primer on 10q but not on 4qter (http://genome.ucsc.edu/).
We designed new primers more proximal to the D4Z4 locus and outside the p13E-11 sequence. Nonetheless, a complex array of PCR products (all below 10 kb) was observed instead of specific products of expected size, which is not surprising according to the GC richness (73%) and the complexity of this region (data not shown).
As the direct D4Z4 array sizing by PCR is not reliable in all situations, we turned towards an indirect molecular strategy by segregation analysis of microsatellite markers. We first evaluated a nested PCR protocol on single lymphocytes.31 After an exhaustive in silico exploration of genomic 4q35.2 sequences, 14 putative polymorphic loci were combined into several multiplex PCRs and evaluated on 600 lymphocytes, followed by about 1500 specific PCRs. The electrophoretic patterns and the estimated amplification rates did not satisfy the recommended criteria for the single-cell level.37 Moreover, although highly sensitive, the nested PCR is costly and time-consuming, requiring a very tedious handling of a large number of PCR microtubes, increasing the risk of contamination and labelling errors.
Among the 14 markers, the D4F104S1 overlaps the recently described SSLP. On the basis of the electrophoresis sizes, it displays four frequent alleles 161, 163, 166 and 168 on 4qA, 4qB and/or 10qA haplotypes.28 As this marker is mapped within the highly homologous 4qter and 10qter regions, all our family members displayed complex electrophoretic patterns with 2–4 alleles. Moreover, the exhaustive 4qA/B haplotyping is unusable for single cell, as it requires Southern blotting for A/B variation and D4Z4 SNP genotyping, and fragment analysis of the SSLP.28 Thus, D4F104S1 proved to be unsuitable for accurate genotyping.
Finally, 10 markers, including D4F104S1, were dismissed and we selected 4 markers (D4S2390, D4S1652, D4S2930 and D4S1523) according to their estimated informativity, amplification efficacies and distance to the D4Z4 locus (1.23–0.55 Mb, Figure 1).
To analyse several microsatellite markers, we evaluated whole genome amplification using multiple displacement amplification (MDA) rapid protocols (1.5 h).38 Locus-specific PCRs on amplified single-cell genomic DNA showed poor efficacies and very high ADOs (data not shown). Similar observations were reported for single-cell applications, and surprisingly, despite these major drawbacks, MDA protocols have been successfully applied in PGD.39, 40, 41, 42, 43, 44, 45
Then, we developed a new one-step multiplex PCR with the four selected markers, which proved to be more rapid (<8 h for 16–32 samples), less expensive and requiring fewer sample handling steps than other protocols. The observed heterozygote rates at the four loci revealed full informativity in our control cohort and corroborated the CEPH data (http://www.cephb.fr/fr/index.php). Haplotype analysis in the D4S2390-D4S1523 interval revealed five crossing-overs, all occurring in the same control family (Figure 2). A double recombination (scored in individual II.5) is a very rare event considering the genetic and physical distances (Figure 1). Therefore, we proposed that the D4S1652-135 bp allele be changed into a 139 bp allele by the addition of a GATA repeat during the replication (Figure 2). Under this hypothesis, recombination events would be 3 out of 144 meioses (2.1%) instead of 5 out of 144 meioses (3.5%), corroborating the recombination fraction θ=0.021, estimated by linkage analysis throughout the genetic interval (D4S2390-D4S1523) in the whole sample of 22 pedigrees. Nonetheless, segregation analysis allowed the identification, in every FSHD family, of a specific haplotype that perfectly segregates with the disease phenotype and with the D4Z4 repeat sizing determined by the Southern blot reference diagnosis. In addition, the D4S1523 showed tight linkage to the D4Z4 locus (maximum LOD score=2.98 at θ=0.0) and the recombination beyond D4S1523 remains low in our FSHD families.
Thus, the rapid, simple and sensitive protocol described in this paper allowed a robust indirect diagnosis that could easily solve or complement standard diagnosis in complex and urgent situations, particularly in the context of PND.
Finally, knowing the hope of young couples suffering from FSHD to benefit from a PGD, we evaluated the quadruplex PCR protocol on a large number of lymphocytes. The specificity and sensitivity of our strategy largely fulfils the recommended criteria for PGD (97.6% amplification rates, ADO and contamination rates both <1%).37
In conclusion, we have designed a new one-step multiplex PCR protocol that is technically efficient and suitable for the indirect diagnosis of FSHD at the single-cell level. This protocol could be theoretically offered to a very limited number of couples persistently requesting a PGD, who are fully informed regarding the selected markers and after careful exclusion of frequent sporadic cases and mosaicism in de novo FSHD families. Owing to the risk of recombination and the absence of flanking distal polymorphic markers, PGD for FSHD should not be recommended. Nevertheless, our improved protocol can be useful to some centres in the world that offer this diagnosis.
References
Lunt PW, Compston DA, Harper PS : Estimation of age dependent penetrance in facioscapulohumeral muscular dystrophy by minimising ascertainment bias. J Med Genet 1989; 26: 755–760.
Tonini MM, Passos-Bueno MR, Cerqueira A, Matioli SR, Pavanello R, Zatz M : Asymptomatic carriers and gender differences in facioscapulohumeral muscular dystrophy (FSHD). Neuromuscul Disord 2004; 14: 33–38.
Gilbert JR, Stajich JM, Wall S et al: Evidence for heterogeneity in facioscapulohumeral muscular dystrophy (FSHD). Am J Hum Genet 1993; 53: 401–408.
Mills KA, Buetow KH, Xu Y et al: Genetic and physical mapping on chromosome 4 narrows the localization of the gene for facioscapulohumeral muscular dystrophy (FSHD). Am J Hum Genet 1992; 51: 432–439.
Sarfarazi M, Wijmenga C, Upadhyaya M et al: Regional mapping of facioscapulohumeral muscular dystrophy gene on 4q35: combined analysis of an international consortium. Am J Hum Genet 1992; 51: 396–403.
van Deutekom JC, Wijmenga C, van Tienhoven EA et al: FSHD associated DNA rearrangements are due to deletions of integral copies of a 3.2 kb tandemly repeated unit. Hum Mol Genet 1993; 2: 2037–2042.
Weiffenbach B, Bagley R, Falls K et al: Linkage analyses of five chromosome 4 markers localizes the facioscapulohumeral muscular dystrophy (FSHD) gene to distal 4q35. Am J Hum Genet 1992; 51: 416–423.
Wijmenga C, Frants RR, Hewitt JE et al: Molecular genetics of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 1993; 3: 487–491.
Butz M, Koch MC, Muller-Felber W, Lemmers RJ, van der Maarel SM, Schreiber H : Facioscapulohumeral muscular dystrophy. Phenotype-genotype correlation in patients with borderline D4Z4 repeat numbers. J Neurol 2003; 250: 932–937.
Klinge L, Eagle M, Haggerty ID, Roberts CE, Straub V, Bushby KM : Severe phenotype in infantile facioscapulohumeral muscular dystrophy. Neuromuscul Disord 2006; 16: 553–558.
Lemmers RJ, Osborn M, Haaf T et al: D4F104S1 deletion in facioscapulohumeral muscular dystrophy: phenotype, size, and detection. Neurology 2003; 61: 178–183.
Lunt PW, Jardine PE, Koch M et al: Phenotypic-genotypic correlation will assist genetic counseling in 4q35-facioscapulohumeral muscular dystrophy. Muscle Nerve 1995; 2: S103–S109.
Ricci E, Galluzzi G, Deidda G et al: Progress in the molecular diagnosis of facioscapulohumeral muscular dystrophy and correlation between the number of KpnI repeats at the 4q35 locus and clinical phenotype. Ann Neurol 1999; 45: 751–757.
Bakker E, Wijmenga C, Vossen RH et al: The FSHD-linked locus D4F104S1 (p13E-11) on 4q35 has a homologue on 10qter. Muscle Nerve 1995; 2: S39–S44.
Deidda G, Cacurri S, Grisanti P, Vigneti E, Piazzo N, Felicetti L : Physical mapping evidence for a duplicated region on chromosome 10qter showing high homology with the facioscapulohumeral muscular dystrophy locus on chromosome 4qter. Eur J Hum Genet 1995; 3: 155–167.
Felice KJ, Jones JM, Conway SR : Facioscapulohumeral dystrophy presenting as infantile facial diplegia and late-onset limb-girdle myopathy in members of the same family. Muscle Nerve 2005; 32: 368–372.
Lemmers RJ, van der Maarel SM, van Deutekom JC et al: Inter- and intrachromosomal sub-telomeric rearrangements on 4q35: implications for facioscapulohumeral muscular dystrophy (FSHD) aetiology and diagnosis. Hum Mol Genet 1998; 7: 1207–1214.
Lemmers RJ, Van Overveld PG, Sandkuijl LA et al: Mechanism and timing of mitotic rearrangements in the subtelomeric D4Z4 repeat involved in facioscapulohumeral muscular dystrophy. Am J Hum Genet 2004; 75: 44–53.
van der Maarel SM, Deidda G, Lemmers RJ et al: De novo facioscapulohumeral muscular dystrophy: frequent somatic mosaicism, sex-dependent phenotype, and the role of mitotic transchromosomal repeat interaction between chromosomes 4 and 10. Am J Hum Genet 2000; 66: 26–35.
Tupler R, Berardinelli A, Barbierato L et al: Monosomy of distal 4q does not cause facioscapulohumeral muscular dystrophy. J Med Genet 1996; 33: 366–370.
Petrov A, Pirozhkova I, Carnac G, Laoudj D, Lipinski M, Vassetzky YS : Chromatin loop domain organization within the 4q35 locus in facioscapulohumeral dystrophy patients versus normal human myoblasts. Proc Natl Acad Sci USA 2006; 103: 6982–6987.
de Greef JC, Frants RR, van der Maarel SM : Epigenetic mechanisms of facioscapulohumeral muscular dystrophy. Mutat Res 2008; 647: 94–102.
Dmitriev P, Lipinski M, Vassetzky YS : Pearls in the junk: dissecting the molecular pathogenesis of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 2009; 19: 17–20.
Pirozhkova I, Petrov A, Dmitriev P, Laoudj D, Lipinski M, Vassetzky Y : A functional role for 4qA/B in the structural rearrangement of the 4q35 region and in the regulation of FRG1 and ANT1 in facioscapulohumeral dystrophy. PLoS ONE 2008; 3: e3389.
Lemmers RJ, de Kievit P, Sandkuijl L et al: Facioscapulohumeral muscular dystrophy is uniquely associated with one of the two variants of the 4q subtelomere. Nat Genet 2002; 32: 235–236.
Lemmers RJ, Wohlgemuth M, Frants RR, Padberg GW, Morava E, van der Maarel SM : Contractions of D4Z4 on 4qB subtelomeres do not cause facioscapulohumeral muscular dystrophy. Am J Hum Genet 2004; 75: 1124–1130.
Thomas NS, Wiseman K, Spurlock G, MacDonald M, Ustek D, Upadhyaya M : A large patient study confirming that facioscapulohumeral muscular dystrophy (FSHD) disease expression is almost exclusively associated with an FSHD locus located on a 4qA-defined 4qter subtelomere. J Med Genet 2007; 44: 215–218.
Lemmers RJ, Wohlgemuth M, van der Gaag KJ et al: Specific sequence variations within the 4q35 region are associated with facioscapulohumeral muscular dystrophy. Am J Hum Genet 2007; 81: 884–894.
Ehrlich M, Jackson K, Tsumagari K, Camano P, Lemmers RJ : Hybridization analysis of D4Z4 repeat arrays linked to FSHD. Chromosoma 2007; 116: 107–116.
Lemmers RJ, van der Wielen MJ, Bakker E, Frants RR, van der Maarel SM : Rapid and accurate diagnosis of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 2006; 16: 615–617; author reply 617–618.
Girardet A, Hamamah S, Anahory T et al: First preimplantation genetic diagnosis of hereditary retinoblastoma using informative microsatellite markers. Mol Hum Reprod 2003; 9: 111–116.
Cui XF, Li HH, Goradia TM et al: Single-sperm typing: determination of genetic distance between the G gamma-globin and parathyroid hormone loci by using the polymerase chain reaction and allele-specific oligomers. Proc Natl Acad Sci USA 1989; 86: 9389–9393.
Lathrop GM, Lalouel JM, Julier C, Ott J : Multilocus linkage analysis in humans: detection of linkage and estimation of recombination. Am J Hum Genet 1985; 37: 482–498.
Goto K, Nishino I, Hayashi YK : Rapid and accurate diagnosis of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 2006; 16: 256–261.
Ki CS, Lee ST, Kim KS et al: Clinical and genetic analysis of korean patients with facioscapulohumeral muscular dystrophy. J Korean Med Sci 2008; 23: 959–963.
Lemmers RJ, van der Wielen MJ, Bakker E, Padberg GW, Frants RR, van der Maarel SM : Somatic mosaicism in FSHD often goes undetected. Ann Neurol 2004; 55: 845–850.
Thornhill AR, deDie-Smulders CE, Geraedts JP et al: ESHRE PGD Consortium ‘Best practice guidelines for clinical preimplantation genetic diagnosis (PGD) and preimplantation genetic screening (PGS)’. Hum Reprod 2005; 20: 35–48.
Dean FB, Hosono S, Fang L et al: Comprehensive human genome amplification using multiple displacement amplification. Proc Natl Acad Sci USA 2002; 99: 5261–5266.
Coskun S, Alsmadi O : Whole genome amplification from a single cell: a new era for preimplantation genetic diagnosis. Prenat Diagn 2007; 27: 297–302.
Handyside AH, Robinson MD, Simpson RJ et al: Isothermal whole genome amplification from single and small numbers of cells: a new era for preimplantation genetic diagnosis of inherited disease. Mol Hum Reprod 2004; 10: 767–772.
Hellani A, Coskun S, Benkhalifa M et al: Multiple displacement amplification on single cell and possible PGD applications. Mol Hum Reprod 2004; 10: 847–852.
Hellani A, Coskun S, Tbakhi A, Al-Hassan S : Clinical application of multiple displacement amplification in preimplantation genetic diagnosis. Reprod Biomed Online 2005; 10: 376–380.
Jiao Z, Zhou C, Li J et al: Birth of healthy children after preimplantation diagnosis of beta-thalassemia by whole-genome amplification. Prenat Diagn 2003; 23: 646–651.
Renwick P, Ogilvie CM : Preimplantation genetic diagnosis for monogenic diseases: overview and emerging issues. Expert Rev Mol Diagn 2007; 7: 33–43.
Wells D, Sherlock JK : Strategies for preimplantation genetic diagnosis of single gene disorders by DNA amplification. Prenat Diagn 1998; 18: 1389–1401.
Acknowledgements
This research was supported in part by the ‘Association Française contre les Myopathies’ (AFM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Barat-Houari, M., Nguyen, K., Bernard, R. et al. New multiplex PCR-based protocol allowing indirect diagnosis of FSHD on single cells: can PGD be offered despite high risk of recombination?. Eur J Hum Genet 18, 533–538 (2010). https://doi.org/10.1038/ejhg.2009.207
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2009.207