Abstract
Fragile X-associated disorders are caused by a CGG trinucleotide repeat expansion in the 5′-untranslated region of the FMR1 gene. Expansion of the CGG trinucleotide repeats to >200 copies (that is, a full mutation) induces methylation of the FMR1 gene, with transcriptional silencing being the eventual outcome. Previous data have shown that FMR1 premutation carriers (individuals with 55–199 repeats) have increased FMR1 mRNA levels with decreased protein (fragile X mental retardation protein (FMRP)) levels. However, the point at which this translational inefficiency occurs, given the increased transcription mechanism, has not yet been explored and remains to be elucidated. We examined the repeat length group, FMR1 transcript and FMRP levels in 74 males with a wide range of repeat lengths using analysis of covariance to better characterize this association. Results showed that the mean FMRP level among carriers with 80–89 repeats was significantly higher than the mean levels among lower (54–79) and higher (90–120) premutation carriers, in spite of the increasing transcript level with repeat length. Taken together, these results suggest that the 80–89-repeat group may lead to different properties that increase the efficiency of translation compared with other premutation repeat size groups.
Similar content being viewed by others
Introduction
Fragile X-associated disorders are caused by the expansion of a CGG trinucleotide repeats located in the 5′-untranslated region of the FMR1 gene. Classification and severity of disorders are based on the length of the trinucleotide repeat, with normal individuals having <45 CGG repeats, intermediates having 45–54 repeats, premutation carriers having 55–199 repeats and full-mutation carriers having >200 repeats.1 The fragile X syndrome occurs in the presence of full mutation in which the FMR1 locus is hypermethylated and transcriptionally silenced, leading to lack of the fragile X mental retardation protein (FMRP).2
Interestingly, premutation carriers have increased levels of FMR1 transcript and decreased FMRP levels.3, 4, 5 The increased transcript levels are due to increased transcription, and not due to increased transcript stability or decreased mRNA turnover.3, 5, 6, 7 The decreased levels of FMRP are thought to be a result of inefficient translation caused by secondary mRNA (hairpin loops) structures that disrupt proper ribosomal scanning, causing stalling at the expanded CGG repeat of FMR1.5, 7, 8, 9 The threshold at which the inverse relationship of increased transcription and decreased FMRP translation occurs is unknown. In this study, we test the hypothesis that this inverse relationship, representing translational inefficiency, is linear with repeat size.
Materials and methods
Subjects
The Emory University institutional review board approved the protocols for this study. Blood samples were obtained from individuals with a wide FMR1 repeat range. Samples were processed and stored at −70 °C for protein analysis and in RNALater (Applied Biosystems, Foster City, CA, USA) for transcript analysis. The study sample consisted of 74 males, with 12 samples previously reported for transcript levels.10
Molecular investigation
The CGG repeat size was determined as previously described.11 On a subset of samples, the AGG interruption pattern was determined by sequencing the CGG repeat region.12 Quantitation of FMR1 levels was previously described using real-time PCR with hypoxanthine–guanine phosphoribosyl transferase 1 (HPRT) as reference.10 The FMRP analysis previously described4 was modified for sensitive quantification using an Enhanced Chemi-Fluorescence (ECF) Kit (GE Healthcare, Waukesha, WI, USA), with anti-FMRP and anti-eIF4E (anti-eukaryotic translation initiation factor 4E) (Bethyl Laboratories, Montgomery, TX, USA) detected by Typhoon 9200 (Amersham Bioscience, Fairfield, CT, USA).
Statistical methods
To determine the effect of CGG repeat size on transcript and protein levels, we used regression analysis and analysis of covariance (ANCOVA). In the post hoc tests to identify group differences, we used Tukey's studentized range test. Transcript level was measured as a ratio of FMR1 and HPRT genes to a relative control with 29 repeats. The FMRP level was represented by a ratio of FMRP and eIF4e. The FMRP/transcript ratio was used as a measure of translational efficiency. To achieve normality, a square root transformation was used for transcript levels and a log transformation for FMRP levels and the FMRP/transcript ratio. SAS System for Windows, Release 8.02 was used for analyses (http://www.sas.com/index.html).
Results
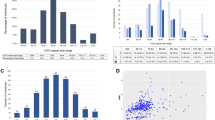
To confirm the effect of repeat size on transcript level, we initially examined transcript levels using regression analysis including days in storage (‘days’) as a covariate. In this model, ‘days’ was statistically significant (β=−0.0003, P<0.02), repeat length as a continuous variable was marginally significant (β=0.003, P=0.08) and together explained 11% of the variation observed in the transcript level. To avoid the assumption of linearity, we used ANCOVA and found similar results: the repeat size group was marginally significant (β=1.13, P=0.09) adjusted for ‘days’ (β=0.76, P=0.03) and, overall, the model explained about 15% of the variation (F=2.75, P<0.04) (Table 1). Results were the same when the two outliers (defined as two standard deviations outside the mean FMR1 transcript) were excluded.
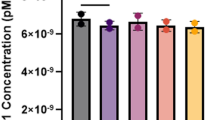
To examine the effect of repeat size on FMRP, we used regression analysis and found that repeat length was marginally significant (P=0.07), whereas ‘days’ was not (P=0.31). Using repeat size groups in an ANCOVA model, only the repeat group was significant (P=0.01) and explained 18% of the variation observed in FMRP levels (F=3.19, P<0.02). Tukey's post hoc group comparisons indicated that the mean FMRP levels of the 80–89 mid-premutation group was significantly higher than that of other premutation groups, but was not different than that in the normal range (Table 1, Figure 1). When data were reanalyzed to exclude four outliers, results were essentially the same.
Mean fragile X mental retardation protein (FMRP) levels (♦) including s.e. for individuals, plotted against their CGG repeat number. For protein analysis, the ratio of FMRP and eukaryotic translation initiation factor 4E (eIF4E), proteins used to assess the translational efficiency of cells, was determined using standard immunodecoration protocols. For protein level determination, each sample was assayed a minimum of three times. The relative FMRP level was determined relative to a control sample. The divided area of the graph represents the normal (shaded), low premutation, mid premutation (shaded) and high premutation range.
To examine the effect of AGG interruptions, we sequenced 43 randomly selected males in our sample. We found that the number of AGG interruptions was not significantly predictive of FMRP and transcript levels or of the translational efficiency ratio (Figure 2), adjusting for repeat size (P>0.3).
Ratio of mean fragile X mental retardation protein (FMRP) level divided by mean FMR1 transcript level for each male, plotted against the CGG repeat number. FMR1 mRNA levels were determined by real-time PCR (Roche, Nutley, NJ, USA) with hypoxanthine–guanine phosphoribosyl transferase 1 (HPRT), a reference gene located on the X-chromosome. The AGG interspersion pattern for each male indicated by symbols: sample with no interruptions (○), ⩾1 AGG interruption (•) or no data available (▵). The divided area of the graph represents the normal (shaded), low premutation, mid premutation (shaded) and high premutation range. The shaded areas indicate the normal and 80–89-repeat range in which FMRP levels are similar.
Discussion
The purpose of this study was to examine the inverse relationship of FMR1 transcript and FMRP levels, representative of translational inefficiency, over a wide range of premutation size repeats. Several reports have clearly shown that transcript levels increase with increasing repeat size, although this effect does not seem to be linear.5, 6, 10, 13, 14 Our data support this observation.
With respect to FMRP levels, reports of decreasing FMRP levels with increasing repeat size have been based primarily on a small series of high premutation carriers (>90 repeats).4, 5, 6 Thus, we examined a wide range of premutation repeat sizes and found a nonlinear association of repeat size and FMRP levels. The mean FMRP level for mid-premutation repeat (80–89) carriers was similar to that in noncarriers and significantly higher than that for low and high premutation carriers. An examination of the ratio of the levels of FMRP/FMR1 transcript suggests greater efficiency of translation in this repeat size range (Figure 2). There are several possible explanations for this finding. First, as there is significant variation in measurements, the results could be due to chance. Although our sample sizes are larger than that of any previous study, they are still relatively small. Second, the increased efficiency could be due to properties related to folding of the transcript on the basis of repeat size range, repeat structure or transcriptional start site.7, 15, 16 We did not observe obvious differences in repeat size structure with respect to the number of AGG interruptions in the mid-premutation group compared with other groups. However, this is only one property that might be considered. Finally, if individuals were related, our observations could be explained by cis elements and other genes that modulate FMRP expression;16, 17, 18, 19 however; the families in our study did not cluster by repeat size.
Recently, a new enzyme-linked immunosorbent assay (ELISA) assay has been developed to quantify FMRP levels.20 This method will allow a larger sample of individuals in the premutation size range to be studied and confirm or refute this interesting preliminary finding of a nonlinear association of repeat size and FMR1 translation efficiency.
References
Sherman, S., Pletcher, B. & Driscoll, D. Fragile X syndrome: diagnositc and carrier testing. Genet. Med. 7, 584–587 (2005).
Gheldof, N., Tabuchi, T. & Dekker, J. The active FMR1 promoter is associated with a large domain of altered chromatin conformation with embedded local histone modification. Proc. Natl Acad. Sci. USA 103, 12463–12468 (2006).
Tassone, F., Hagerman, R. J., Taylor, A. K., Mills, J. B., Harris, S. W., Gane, L. W. et al. Clinical involvement and protein expression in individuals with the FMR1 premutation. Am. J. Med. Genet. 91, 144–152 (2000).
Kenneson, A., Zhang, F., Hagedorn, C. & Warren, S. Reduced FMRP and increased FMR1 transcript is proportionally associated with CGG repeat number in intermediate-length and premutation carriers. Hum. Mol. Genet. 10, 1449–1454 (2001).
Tassone, F., Hagerman, R., Taylor, A., Gane, L., Godfrey, T. & Hagerman, P. Elevated levels of FMR1 mRNA in carrier males: a new mechanism of involvement in the Fragile-X Syndrome. Am. J. Hum. Genet. 66, 6–15 (2000).
Primerano, B., Tassone, F., Hagerman, R., Hagerman, P., Amaldi, F. & Bagni, C. Reduced FMR1 mRNA translation efficiency in Fragile X patients with premutations. RNA 8, 1482–1488 (2002).
Feng, Y., Zhang, F., Lokey, L. K., Chastain, J. L., Lakkis, L., Eberhart, D. et al. Translational supperssion by trinucleotide repeat expansion at FMR1. Science 268, 731–734 (1995).
Laggerbauer, B., Ostareck, D., Keidel, E. M., Ostareck-Lederer, A. & Fischer, U. Evidence that fragile X mental retardation protein is a negative regulator of translation. Hum. Mol.Genet. 10, 329–338 (2001).
Fu, Y., Kuhl, D., Pizzuti, A., Pieretti, M., Sutcliffe, J. S., Richards, S. et al. Variation of the CGG repeat at the fragile X site results in genetic instability: resolution of the Sherman paradox 1991. Cell 1047–1058.
Allen, E., He, W., Yadav-Shah, M. & Sherman, S. A study of the distribution characteristics of FMR1 transcript levels in 238 individuals. Hum. Genet. 114, 439–447 (2004).
Meadows, K., Pettay, D., Newman, J., Hersey, J., Ashley, A., Sherman, S. Survey of the fragile X syndrome and the fragile X E syndrome in a special needs population. Am. J. Med. Genet. 64, 428–433 (1996).
Crawford, D., Meadows, K., Newman, J. L., Taft, L. F., Pettay, D. L., Gold, L. B. et al. Prevalence and phenotype consquence of FRAXA and FRAXE alleles in a large, ethnically diverse, special education-needs population. Am. J. Hum. Genet. 64, 495–507 (1999).
Tassone, F., Beilina, A., Carosi, C., Albertosi, S., Bagni, C. & Li, L. et al. Elevated FMR1 mRNA in premutation carriers is due to increased transcription. RNA 13, 555–562 (2007).
Gracia-Alegria, E., Ibanez, B., Minguez, M., Poch, M., Valiente, A. & Sanz-Parra, A. et al. Analysis of FMR1 gene expression in female premutation carriers using robust segmented linear regression models. RNA 13, 756–762 (2007).
Zumwalt, M., Ludwig, A., Hagerman, P. & Dieckmann, T. Secondary structure and dynamics of the r(CGG) repeat in the mRNA of the fragile X mental retardation 1 (FMR1) gene. RNA. Biol. 4, 93–100 (2007).
Beilina, A., Tassone, F., Schwartz, P. H., Sahota, P. & Hagerman, P. J. Redistribution of transcription start sites within the FMR1 promoter region with expansion of the downstream CGG-repeat element. Hum. Mol. Genet. 13, 543–549 (2004).
Singh, K. & Prasad, S. Differential expression of Fmr-1 mRNA and FMRP in female mice brain during aging. Mol. Biol. Rep. 35, 677–684 (2008).
Entezam, A. & Usdin, K. ATR protects the genome against CGG CGG-repeat expansion in Fragile X premutation mice. Nucleic. Acids. Res. 36, 1050–1056 (2008).
Handa, V., Goldwater, D., Stiles, D., Cam, M., Poy, G., Kumari, D. et al. Long CCG-repeat tracts are toxic to human cells: implications for carriers of Fragile X premutation alleles. FEBS Lett. 579, 2702–2708 (2005).
Iwahashi, C., Tassone, F., Hagerman, R. J., Yasui, D., Parrott, G., Nguyen, D. et al. A quantitative ELISA assay for the fragile X mental retardation 1 protein. J. Mol. Diagn. 11, 281–289 (2009).
Acknowledgements
We thank Dr Stephen Warren for kindly providing the anti-FMRP (J1) antibody. This work was supported by the National Institutes of Health grants R01HD029909 and G12GM00680.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Peprah, E., He, W., Allen, E. et al. Examination of FMR1 transcript and protein levels among 74 premutation carriers. J Hum Genet 55, 66–68 (2010). https://doi.org/10.1038/jhg.2009.121
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2009.121
Keywords
This article is cited by
-
Spectrum of Syndromal Disorders Associated with Expansion of CGG Repeats of the FMR1 Gene Promoter: Pathogenetic Mechanisms and Clinical Manifestations
Neuroscience and Behavioral Physiology (2023)
-
AGG interruptions and maternal age affect FMR1 CGG repeat allele stability during transmission
Journal of Neurodevelopmental Disorders (2014)
-
Use of model systems to understand the etiology of fragile X-associated primary ovarian insufficiency (FXPOI)
Journal of Neurodevelopmental Disorders (2014)
-
Genomic studies in fragile X premutation carriers
Journal of Neurodevelopmental Disorders (2014)
-
Intranuclear inclusions in a fragile X mosaic male
Translational Neurodegeneration (2013)