Abstract
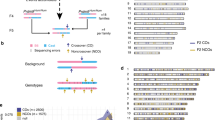
Meiotic recombination predominantly occurs at discrete genomic loci called recombination hotspots, but the features defining these areas are still largely unknown (reviewed in refs 1–5). To allow a comprehensive analysis of hotspot-associated DNA and chromatin characteristics, we developed a direct molecular approach for mapping meiotic DNA double-strand breaks that initiate recombination. Here we present the genome-wide distribution of recombination initiation sites in the mouse genome. Hotspot centres are mapped with approximately 200-nucleotide precision, which allows analysis of the fine structural details of the preferred recombination sites. We determine that hotspots share a centrally distributed consensus motif, possess a nucleotide skew that changes polarity at the centres of hotspots and have an intrinsic preference to be occupied by a nucleosome. Furthermore, we find that the vast majority of recombination initiation sites in mouse males are associated with testis-specific trimethylation of lysine 4 on histone H3 that is distinct from histone H3 lysine 4 trimethylation marks associated with transcription. The recombination map presented here has been derived from a homogeneous mouse population with a defined genetic background and therefore lends itself to extensive future experimental exploration. We note that the mapping technique developed here does not depend on the availability of genetic markers and hence can be easily adapted to other species with complex genomes. Our findings uncover several fundamental features of mammalian recombination hotspots and underline the power of the new recombination map for future studies of genetic recombination, genome stability and evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Arnheim, N., Calabrese, P. & Tiemann-Boege, I. Mammalian meiotic recombination hot spots. Annu. Rev. Genet. 41, 369–399 (2007)
Buard, J. & de Massy, B. Playing hide and seek with mammalian meiotic crossover hotspots. Trends Genet. 23, 301–309 (2007)
Lichten, M. Meiotic chromatin: the substrate for recombination initiation. Genome Dynam. Stabil. 3, 165–193 (2008)
Paigen, K. & Petkov, P. Mammalian recombination hot spots: properties, control and evolution. Nature Rev. Genet. 11, 221–233 (2010)
Clark, A. G., Wang, X. & Matise, T. Contrasting methods of quantifying fine structure of human recombination. Annu. Rev. Genom. Hum. Genet. 11, 45–64 (2010)
Kauppi, L., May, C. A. & Jeffreys, A. J. Analysis of meiotic recombination products from human sperm. Methods Mol. Biol. 557, 323–355 (2009)
The. International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005)
Durbin, R. M. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010)
Myers, S., Bottolo, L., Freeman, C., McVean, G. & Donnelly, P. A fine-scale map of recombination rates and hotspots across the human genome. Science 310, 321–324 (2005)
Neale, M. J. & Keeney, S. Clarifying the mechanics of DNA strand exchange in meiotic recombination. Nature 442, 153–158 (2006)
Petukhova, G. V., Romanienko, P. J. & Camerini-Otero, R. D. The Hop2 protein has a direct role in promoting interhomolog interactions during mouse meiosis. Dev. Cell 5, 927–936 (2003)
Qin, J., Richardson, L. L., Jasin, M., Handel, M. A. & Arnheim, N. Mouse strains with an active H2-Ea meiotic recombination hot spot exhibit increased levels of H2-Ea-specific DNA breaks in testicular germ cells. Mol. Cell. Biol. 24, 1655–1666 (2004)
Paigen, K. et al. The recombinational anatomy of a mouse chromosome. PLoS Genet. 4, e1000119 (2008)
Cox, A. et al. A new standard genetic map for the laboratory mouse. Genetics 182, 1335–1344 (2009)
Burgoyne, P. S. Genetic homology and crossing over in the X and Y chromosomes of mammals. Hum. Genet. 61, 85–90 (1982)
Khambata, S., Mody, J., Modzelewski, A., Heine, D. & Passmore, H. C. Ea recombinational hot spot in the mouse major histocompatibility complex maps to the fourth intron of the Ea gene. Genome Res. 6, 195–201 (1996)
Francino, M. P. & Ochman, H. Strand asymmetries in DNA evolution. Trends Genet. 13, 240–245 (1997)
Duret, L. & Galtier, N. Biased gene conversion and the evolution of mammalian genomic landscapes. Annu. Rev. Genom. Hum. Genet. 10, 285–311 (2009)
Mihola, O., Trachtulec, Z., Vlcek, C., Schimenti, J. C. & Forejt, J. A mouse speciation gene encodes a meiotic histone H3 methyltransferase. Science 323, 373–375 (2009)
Wang, Z., Schones, D. E. & Zhao, K. Characterization of human epigenomes. Curr. Opin. Genet. Dev. 19, 127–134 (2009)
Borde, V. et al. Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J. 28, 99–111 (2008)
Buard, J., Barthes, P., Grey, C. & de Massy, B. Distinct histone modifications define initiation and repair of meiotic recombination in the mouse. EMBO J. 28, 2616–2624 (2009)
Parvanov, E. D., Petkov, P. M. & Paigen, K. Prdm9 controls activation of mammalian recombination hotspots. Science 327, 835 (2010)
Myers, S. et al. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 327, 876–879 (2010)
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010)
Myers, S., Freeman, C., Auton, A., Donnelly, P. & McVean, G. A common sequence motif associated with recombination hot spots and genome instability in humans. Nature Genet. 40, 1124–1129 (2008)
Berg, I. L. et al. PRDM9 variation strongly influences recombination hot-spot activity and meiotic instability in humans. Nature Genet. 42, 859–863 (2010)
Kaplan, N. et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458, 362–366 (2009)
Acknowledgements
We thank M. Lichten (NCI, NIH) and P. Hsieh (NIDDK, NIH) for comments and discussion. We are grateful to S. Sharmeen for her help with high-throughput sequencing. This work was supported in part by Basil O’Connor Starter Scholar Research Award Grant No. 5-FY07-667 from the March of Dimes Foundation (G.V.P.); NIH grant 1R01GM084104-01A1 from NIGMS (G.V.P.); New Investigator Start-up Grants FS71HU, R071HU and CS71HU from USUHS (G.V.P.); and the NIDDK (NIH) Intramural Research Program (R.D.C.-O.).
Author information
Authors and Affiliations
Contributions
F.S. performed all experiments. I.V.G., K.B. and P.K. performed computational data analyses. All authors contributed to experimental design. G.V.P. and R.D.C.-O. designed and supervised the study. G.V.P. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, Supplementary Figures 1-16 with legends, Supplementary Table 1, Supplementary Methods and Materials and additional references. (PDF 2679 kb)
Supplementary Data
This data file contains listings of the DSB hotspots. (XLS 585 kb)
Rights and permissions
About this article
Cite this article
Smagulova, F., Gregoretti, I., Brick, K. et al. Genome-wide analysis reveals novel molecular features of mouse recombination hotspots. Nature 472, 375–378 (2011). https://doi.org/10.1038/nature09869
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09869
This article is cited by
-
Molecular mechanisms and regulation of recombination frequency and distribution in plants
Theoretical and Applied Genetics (2024)
-
FIGNL1 AAA+ ATPase remodels RAD51 and DMC1 filaments in pre-meiotic DNA replication and meiotic recombination
Nature Communications (2023)
-
The effect of DNA polymorphisms and natural variation on crossover hotspot activity in Arabidopsis hybrids
Nature Communications (2023)
-
The histone modification reader ZCWPW1 promotes double-strand break repair by regulating cross-talk of histone modifications and chromatin accessibility at meiotic hotspots
Genome Biology (2022)
-
Achiasmatic meiosis in the unisexual Amazon molly, Poecilia formosa
Chromosome Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.