Abstract
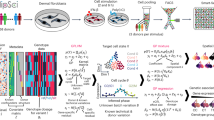
Individual genetic variation affects gene responsiveness to stimuli, often by influencing complex molecular circuits. Here we combine genomic and intermediate-scale transcriptional profiling with computational methods to identify variants that affect the responsiveness of genes to stimuli (responsiveness quantitative trait loci or reQTLs) and to position these variants in molecular circuit diagrams. We apply this approach to study variation in transcriptional responsiveness to pathogen components in dendritic cells from recombinant inbred mouse strains. We identify reQTLs that correlate with particular stimuli and position them in known pathways. For example, in response to a virus-like stimulus, a trans-acting variant responds as an activator of the antiviral response; using RNA interference, we identify Rgs16 as the likely causal gene. Our approach charts an experimental and analytic path to decipher the mechanisms underlying genetic variation in circuits that control responses to stimuli.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Smith, E.N. & Kruglyak, L. Gene-environment interaction in yeast gene expression. PLoS Biol. 6, e83 (2008).
Heinig, M. et al. A trans-acting locus regulates an antiviral expression network and type 1 diabetes risk. Nature 467, 460–464 (2010).
Zhong, H. et al. Liver and adipose expression associated SNPs are enriched for association to type 2 diabetes. PLoS Genet. 6, e1000932 (2010).
Barreiro, L.B. et al. Deciphering the genetic architecture of variation in the immune response to Mycobacterium tuberculosis infection. Proc. Natl. Acad. Sci. USA 109, 1204–1209 (2012).
Gargalovic, P.S. et al. Identification of inflammatory gene modules based on variations of human endothelial cell responses to oxidized lipids. Proc. Natl. Acad. Sci. USA 103, 12741–12746 (2006).
Smirnov, D.A., Morley, M., Shin, E., Spielman, R.S. & Cheung, V.G. Genetic analysis of radiation-induced changes in human gene expression. Nature 459, 587–591 (2009).
Yang, I.V. et al. Identification of novel genes that mediate innate immunity using inbred mice. Genetics 183, 1535–1544 (2009).
Dombroski, B.A. et al. Gene expression and genetic variation in response to endoplasmic reticulum stress in human cells. Am. J. Hum. Genet. 86, 719–729 (2010).
Romanoski, C.E. et al. Systems genetics analysis of gene-by-environment interactions in human cells. Am. J. Hum. Genet. 86, 399–410 (2010).
Maranville, J.C. et al. Interactions between glucocorticoid treatment and cis-regulatory polymorphisms contribute to cellular response phenotypes. PLoS Genet. 7, e1002162 (2011).
Peirce, J.L., Lu, L., Gu, J., Silver, L.M. & Williams, R.W. A new set of BXD recombinant inbred lines from advanced intercross populations in mice. BMC Genet. 5, 7 (2004).
Takeda, K. & Akira, S. Toll-like receptors in innate immunity. Int. Immunol. 17, 1–14 (2005).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Wang, J., Williams, R.W. & Manly, K.F. WebQTL: web-based complex trait analysis. Neuroinformatics 1, 299–308 (2003).
Chesler, E.J. et al. Complex trait analysis of gene expression uncovers polygenic and pleiotropic networks that modulate nervous system function. Nat. Genet. 37, 233–242 (2005).
Bystrykh, L. et al. Uncovering regulatory pathways that affect hematopoietic stem cell function using 'genetical genomics'. Nat. Genet. 37, 225–232 (2005).
Fairfax, B.P. et al. Genetics of gene expression in primary immune cells identifies cell type-specific master regulators and roles of HLA alleles. Nat. Genet. 44, 502–510 (2012).
Gerrits, A. et al. Expression quantitative trait loci are highly sensitive to cellular differentiation state. PLoS Genet. 5, e1000692 (2009).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type-dependent manner. Science 325, 1246–1250 (2009).
Mackay, T.F., Stone, E.A. & Ayroles, J.F. The genetics of quantitative traits: challenges and prospects. Nat. Rev. Genet. 10, 565–577 (2009).
Hubner, N. et al. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nat. Genet. 37, 243–253 (2005).
Amit, I. et al. Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science 326, 257–263 (2009).
Garber, M. et al. A high-throughput chromatin immunoprecipitation approach reveals principles of dynamic gene regulation in mammals. Mol. Cell 47, 810–822 (2012).
O'Neill, L.A. When signaling pathways collide: positive and negative regulation of toll-like receptor signal transduction. Immunity 29, 12–20 (2008).
Barbalat, R., Lau, L., Locksley, R.M. & Barton, G.M. Toll-like receptor 2 on inflammatory monocytes induces type I interferon in response to viral but not bacterial ligands. Nat. Immunol. 10, 1200–1207 (2009).
Ulitsky, I. et al. Expander: from expression microarrays to networks and functions. Nat. Protoc. 5, 303–322 (2010).
Chevrier, N. et al. Systematic discovery of TLR signaling components delineates viral-sensing circuits. Cell 147, 853–867 (2011).
Baum, A. & Garcia-Sastre, A. Differential recognition of viral RNA by RIG-I. Virulence 2, 166–169 (2011).
Diebold, S.S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Frazer, K.A. et al. Segmental phylogenetic relationships of inbred mouse strains revealed by fine-scale analysis of sequence variation across 4.6 mb of mouse genome. Genome Res. 14, 1493–1500 (2004).
Chen, C., Wang, H., Fong, C.W. & Lin, S.C. Multiple phosphorylation sites in RGS16 differentially modulate its GAP activity. FEBS Lett. 504, 16–22 (2001).
Hackett, C.A., Meyer, R.C. & Thomas, W.T. Multi-trait QTL mapping in barley using multivariate regression. Genet. Res. 77, 95–106 (2001).
Xu, C., Li, Z. & Xu, S. Joint mapping of quantitative trait Loci for multiple binary characters. Genetics 169, 1045–1059 (2005).
Banerjee, S., Yandell, B.S. & Yi, N. Bayesian quantitative trait loci mapping for multiple traits. Genetics 179, 2275–2289 (2008).
Gilbert, H. & Le Roy, P. Methods for the detection of multiple linked QTL applied to a mixture of full and half sib families. Genet. Sel. Evol. 39, 139–158 (2007).
Lee, S.I., Pe'er, D., Dudley, A.M., Church, G.M. & Koller, D. Identifying regulatory mechanisms using individual variation reveals key role for chromatin modification. Proc. Natl. Acad. Sci. USA 103, 14062–14067 (2006).
Xie, S. et al. IL-17 activates the canonical NF-kappaB signaling pathway in autoimmune B cells of BXD2 mice to upregulate the expression of regulators of G-protein signaling 16. J. Immunol. 184, 2289–2296 (2010).
Shankar, S.P. et al. RGS16 Attenuates Pulmonary Th2/Th17 Inflammatory Responses. J. Immunol. 188, 6347–6356 (2012).
Shi, G.X., Harrison, K., Han, S.B., Moratz, C. & Kehrl, J.H. Toll-like receptor signaling alters the expression of regulator of G protein signaling proteins in dendritic cells: implications for G protein-coupled receptor signaling. J. Immunol. 172, 5175–5184 (2004).
Park, H. et al. Discovery of common Asian copy number variants using integrated high-resolution array CGH and massively parallel DNA sequencing. Nat. Genet. 42, 400–405 (2010).
Addona, T.A. et al. Multi-site assessment of the precision and reproducibility of multiple reaction monitoring-based measurements of proteins in plasma. Nat. Biotechnol. 27, 633–641 (2009).
Bendall, S.C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Irizarry, R.A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Wilks, S.S. The large-sample distribution of the likelihood ratio for testing composite hypotheses. Ann. Math. Stat. 9, 60–62 (1938).
Geiss, G.K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Gaglani, S.M., Lu, L., Williams, R.W. & Rosen, G.D. The genetic control of neocortex volume and covariation with neocortical gene expression in mice. BMC Neurosci. 10, 44 (2009).
Scheffe, H. The Analysis of Variance (John Wiley and Sons, Inc., 1959).
Falconer, D. & Mackay, T.F.C. Introduction to Quantitative Genetics (Pearson, 1996).
Root, D.E., Hacohen, N., Hahn, W.C., Lander, E.S. & Sabatini, D.M. Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat. Methods 3, 715–719 (2006).
Acknowledgements
We thank C. (Jimmie) Ye, J. Pickerel, M. Daly and E. Lander for comments and discussions. I.G.-V. and I.A. were supported by Human Frontiers Science Program postdoctoral fellowships. Work was supported by Howard Hughes Medical Institute, a US National Institutes of Health PIONEER award, a Burroughs-Wellcome Fund Career Award at the Scientific Interface (A.R.), a Center for Excellence in Genome Science grant 5P50HG006193-02 from the National Human Genome Research Institute (N.H. and A.R.), the Klarman Cell Observatory at the Broad Institute (A.R.), the New England Regional Center for Excellence/Biodefense and Emerging Infectious Disease U54 AI057159 (N.H.), the Israeli Centers of Research Excellence (I-CORE) Gene Regulation in Complex Human Disease, Center No. 41/11 (I.G.-V., R.W. and Y.S.), the Human Frontiers Science Program Career Development Award and an Israeli Science Foundation Bikura Institutional Research Grant Program (I.A.) and the Edmond J. Safra Center for Bioinformatics at Tel-Aviv University (R.W. and Y.S.). A.R. is a fellow of the Merkin Foundation for Stem Cell Research at the Broad Institute. I.G.-V. is a Faculty Fellow of the Edmond J. Safra Center for Bioinformatics at Tel Aviv University and an Alon Fellow.
Author information
Authors and Affiliations
Contributions
I.G.-V., I.A. and A.R. conceived and designed the study. N.C., T.E., R.R., A.S. and I.A. conducted the experiments. I.G.-V. and A.R. conceived computational methods. I.G.-V., R.W. and Y.S. conceived, developed and implemented the computational methods. N.H. participated in study design and interpretation. I.G.-V., I.A. and A.R. wrote the manuscript with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1–6 and Supplementary Notes 1–3 (PDF 8508 kb)
Supplementary Table 7
Dataset 1 (XLS 761 kb)
Supplementary Table 8
Dataset 2 (XLS 739 kb)
Supplementary Table 9
Dataset 3 (XLS 746 kb)
Supplementary Table 10
Dataset 4 (XLS 761 kb)
Rights and permissions
About this article
Cite this article
Gat-Viks, I., Chevrier, N., Wilentzik, R. et al. Deciphering molecular circuits from genetic variation underlying transcriptional responsiveness to stimuli. Nat Biotechnol 31, 342–349 (2013). https://doi.org/10.1038/nbt.2519
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2519