Abstract
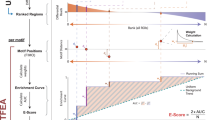
Using 62 probe-level datasets obtained with a custom-designed Caulobacter crescentus microarray chip, we identify transcriptional start sites of 769 genes, 53 of which are transcribed from multiple start sites. Transcriptional start sites are identified by analyzing probe signal cross-correlation matrices created from probe pairs tiled every 5 bp upstream of the genes. Signals from probes binding the same message are correlated. The contribution of each promoter for genes transcribed from multiple promoters is identified. Knowing the transcription start site enables targeted searching for regulatory-protein binding motifs in the promoter regions of genes with similar expression patterns. We identified 27 motifs, 17 of which share no similarity to the characterized motifs of other C. crescentus transcriptional regulators. Using these motifs, we predict coregulated genes. We verified novel promoter motifs that regulate stress-response genes, including those responding to uranium challenge, a stress-response sigma factor and a stress-response noncoding RNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
McAdams, H.H. & Shapiro, L. A bacterial cell-cycle regulatory network operating in time and space. Science 301, 1874–1877 (2003).
Ausmees, N. & Jacobs-Wagner, C. Spatial and temporal control of differentiation and cell cycle progression in Caulobacter crescentus. Annu. Rev. Microbiol. 57, 225–247 (2003).
Skerker, J.M. & Laub, M.T. Cell-cycle progression and the generation of asymmetry in Caulobacter crescentus. Nat. Rev. Microbiol. 2, 325–337 (2004).
Viollier, P.H. & Shapiro, L. Spatial complexity of mechanisms controlling a bacterial cell cycle. Curr. Opin. Microbiol. 7, 572–578 (2004).
Laub, M.T., McAdams, H.H., Feldblyum, T., Fraser, C.M. & Shapiro, L. Global analysis of the genetic network controlling a bacterial cell cycle. Science 290, 2144–2148 (2000).
Collier, J., Murray, S.R. & Shapiro, L. DnaA couples DNA replication and the expression of two cell cycle master regulators. EMBO J. 25, 346–356 (2006).
Hottes, A.K., Shapiro, L. & McAdams, H.H. DnaA coordinates replication initiation and cell cycle transcription in Caulobacter crescentus. Mol. Microbiol. 58, 1340–1353 (2005).
Holtzendorff, J. et al. Oscillating global regulators control the genetic circuit driving a bacterial cell cycle. Science 304, 983–987 (2004).
Quon, K.C., Yang, B., Domian, I.J., Shapiro, L. & Marczynski, G.T. Negative control of bacterial DNA replication by a cell cycle regulatory protein that binds at the chromosome origin. Proc. Natl. Acad. Sci. USA 95, 120–125 (1998).
Laub, M.T., Chen, S.L., Shapiro, L. & McAdams, H.H. Genes directly controlled by CtrA, a master regulator of the Caulobacter cell cycle. Proc. Natl. Acad. Sci. USA 99, 4632–4637 (2002).
Gomes, S.L., Gober, J.W. & Shapiro, L. Expression of the Caulobacter heat shock gene dnaK is developmentally controlled during growth at normal temperatures. J. Bacteriol. 172, 3051–3059 (1990).
Crosson, S., McGrath, P.T., Stephens, C., McAdams, H.H. & Shapiro, L. Conserved modular design of an oxygen sensory/signaling network with species-specific output. Proc. Natl. Acad. Sci. USA 102, 8018–8023 (2005).
Hu, P., Brodie, E.L., Suzuki, Y., McAdams, H.H. & Andersen, G.L. Whole Genome Transcriptional Analysis of Heavy Metal Stresses in Caulobacter crescentus. J. Bacteriol. 187, 8437–8449 (2005).
Tavazoie, S., Hughes, J.D., Campbell, M.J., Cho, R.J. & Church, G.M. Systematic determination of genetic network architecture. Nat. Genet. 22, 281–285 (1999).
Tjaden, B., Haynor, D.R., Stolyar, S., Rosenow, C. & Kolker, E. Identifying operons and untranslated regions of transcripts using Escherichia coli RNA expression analysis. Bioinformatics 18 Suppl 1, S337–S344 (2002).
Kelly, A.J., Sackett, M.J., Din, N., Quardokus, E. & Brun, Y.V. Cell cycle-dependent transcriptional and proteolytic regulation of FtsZ in Caulobacter. Genes Dev. 12, 880–893 (1998).
Mullin, D.A. & Newton, A. Ntr-like promoters and upstream regulatory sequence ftr are required for transcription of a developmentally regulated Caulobacter crescentus flagellar gene. J. Bacteriol. 171, 3218–3227 (1989).
McGrath, P.T., Iniesta, A.A., Ryan, K.R., Shapiro, L. & McAdams, H.H. A dynamically localized protease complex and a polar specificity factor control a cell cycle master regulator. Cell 124, 535–547 (2006).
Roberts, R.C. & Shapiro, L. Transcription of genes encoding DNA replication proteins is coincident with cell cycle control of DNA replication in Caulobacter crescentus. J. Bacteriol. 179, 2319–2330 (1997).
Osteras, M., Stotz, A., Schmid Nuoffer, S. & Jenal, U. Identification and transcriptional control of the genes encoding the Caulobacter crescentus ClpXP protease. J. Bacteriol. 181, 3039–3050 (1999).
Bailey, T.L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Liu, X., Brutlag, D.L. & Liu, J.S. BioProspector: discovering conserved DNA motifs in upstream regulatory regions of co-expressed genes. Pac. Symp. Biocomput, 127–138 (2001).
Bailey, T.L. & Gribskov, M. Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14, 48–54 (1998).
Herring, C.D. et al. Immobilization of Escherichia coli RNA polymerase and location of binding sites by use of chromatin immunoprecipitation and microarrays. J. Bacteriol. 187, 6166–6174 (2005).
Reisenauer, A., Mohr, C.D. & Shapiro, L. Regulation of a heat shock sigma32 homolog in Caulobacter crescentus. J. Bacteriol. 178, 1919–1927 (1996).
Craven, M., Page, D., Shavlik, J., Bockhorst, J. & Glasner, J. A probabilistic learning approach to whole-genome operon prediction. Proc. Int. Conf. Intell. Syst. Mol. Biol. 8, 116–127 (2000).
Gober, J.W. & Shapiro, L. A developmentally regulated Caulobacter flagellar promoter is activated by 3′ enhancer and IHF binding elements. Mol. Biol. Cell 3, 913–926 (1992).
Helmann, J.D. The extracytoplasmic function (ECF) sigma factors. Adv. Microb. Physiol. 46, 47–110 (2002).
Brun, Y.V. & Shapiro, L. A temporally controlled sigma-factor is required for polar morphogenesis and normal cell division in Caulobacter. Genes Dev. 6, 2395–2408 (1992).
Malakooti, J., Wang, S.P. & Ely, B. A consensus promoter sequence for Caulobacter crescentus genes involved in biosynthetic and housekeeping functions. J. Bacteriol. 177, 4372–4376 (1995).
Acknowledgements
We thank Esteban Toro for comments on the manuscript and Michael Laub and Michael Mittman for assistance in designing the CauloHI1 chip. This work was supported by National Institute of Health (NIH) grants GM32506 (L.S.) and 5R24GM73011-2 (H.H.M., L.S.), Department of Energy Office of Science grant DE-FG02-01ER63219 (H.H.M., L.S.), NIH training grant T32 HG00044 (P.T.M.) and Damon Runyon Cancer Research Foundation Fellowship DRG-1880-05 (N.J.H.). P.H. was funded by the DOE Genomes to Life program under the auspices of the University of California, Lawrence Berkeley National Laboratory, Contract no. DE-AC02-05CH11231. H.L. was supported by a Stanford Graduate Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Correlation Tables for dnaN (CC0156) and clpX (CC1961) (PDF 239 kb)
Supplementary Fig. 2
Correlation tables for CC2510 (PDF 19 kb)
Supplementary Fig. 3
Cluster diagram of co-expressed genes in cell cycle microarray experiments (PDF 43 kb)
Supplementary Fig. 4
Cluster diagram of coexpressed genes from heavy metal response data (PDF 33 kb)
Supplementary Fig. 5
cc_2 promoter site (PDF 11 kb)
Supplementary Fig. 6
cc_3 is a FixK binding site (PDF 13 kb)
Supplementary Fig. 7
cc_7 is a SigU binding site (PDF 15 kb)
Supplementary Fig. 8
m_5 in the CC1891 promoter (PDF 14 kb)
Supplementary Fig. 9
Motif cc_5 is the RNAP sigma 54 binding motif (PDF 12 kb)
Supplementary Table 1
Motif Summary (PDF 13 kb)
Rights and permissions
About this article
Cite this article
McGrath, P., Lee, H., Zhang, L. et al. High-throughput identification of transcription start sites, conserved promoter motifs and predicted regulons. Nat Biotechnol 25, 584–592 (2007). https://doi.org/10.1038/nbt1294
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1294
This article is cited by
-
Role and regulation of ferritin-like proteins in iron homeostasis and oxidative stress survival of Caulobacter crescentus
BioMetals (2016)
-
Analysis of the Caulobacter crescentus Zur regulon reveals novel insights in zinc acquisition by TonB-dependent outer membrane proteins
BMC Genomics (2014)
-
Cell cycle transition from S-phase to G1 in Caulobacter is mediated by ancestral virulence regulators
Nature Communications (2014)
-
Genome organization in Mycoplasma hyopneumoniae: identification of promoter-like sequences
Molecular Biology Reports (2014)