Abstract
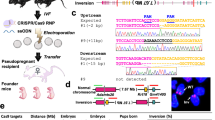
Homologous recombination in Escherichia coli simplifies the generation of gene targeting constructs for transduction into mouse embryonic stem (ES) cells1,2,3,4,5,6,7. Taking advantage of the extensive homology provided by intact bacterial artificial chromosomes (BACs), we have developed an efficient method for preparing targeted gene disruptions in ES cells. Correctly integrated clones were identified by a simple screening procedure based on chromosomal fluorescence in situ hybridization (FISH). To date, five mutant lines have been generated and bred to homozygosity by this approach.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tsuzuki, T. & Rancourt, D.E. Embryonic stem cell gene targeting using bacteriophage lambda vectors generated by phage-plasmid recombination. Nucleic Acids Res. 26, 988–993 (1998).
Yang, X.W., Model, P. & Heintz, N. Homologous recombination based modification in Escherichia coli and germline transmission in transgenic mice of a bacterial artificial chromosome. Nat. Biotechnol. 15, 859–865 (1997).
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 5978–5983 (2000).
Zhang, Y., Buchholz, F., Muyrers, J.P. & Stewart, A.F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Zhang, P., Li, M.Z. & Elledge, S.J. Towards genetic genome projects: genomic library screening and gene-targeting vector construction in a single step. Nat. Genet. 30, 31–39 (2002).
Murphy, K.C. Use of bacteriophage lambda recombination functions to promote gene replacement in Escherichia coli. J. Bacteriol. 180, 2063–2071 (1998).
Angrand, P.O., Daigle, N., van der Hoeven, F., Scholer, H.R. & Stewart, A.F. Simplified generation of targeting constructs using ET recombination. Nucleic Acids Res. 27, e16 (1999).
Koller, B.H. & Smithies, O. Altering genes in animals by gene targeting. Annu. Rev. Immunol. 10, 705–730 (1992).
Soriano, P. Gene targeting in ES cells. Annu. Rev. Neurosci. 18, 1–18 (1995).
Deng, C. & Capecchi, M.R. Reexamination of gene targeting frequency as a function of the extent of homology between the targeting vector and the target locus. Mol. Cell Biol. 12, 3365–3371 (1992).
Hasty, P., Rivera-Perez, J. & Bradley, A. The length of homology required for gene targeting in embryonic stem cells. Mol. Cell Biol. 11, 5586–5591 (1991).
Poteete, A.R. & Fenton, A.C. Lambda red-dependent growth and recombination of phage P22. Virology 134, 161–167 (1984).
El Karoui, M., Amundsen, S.K., Dabert, P. & Gruss, A. Gene replacement with linear DNA in electroporated wild-type Escherichia coli. Nucleic Acids Res. 27, 1296–1299 (1999).
Deichmann, M., Bentz, M. & Haas, R. Ultra-sensitive FISH is a useful tool for studying chronic HIV-1 infection. J. Virol. Methods 65, 19–25 (1997).
Reid, L.H., Shesely, E.G., Kim, H.S. & Smithies, O. Cotransformation and gene targeting in mouse embryonic stem cells. Mol. Cell Biol. 11, 2769–2777 (1991).
Bollag, R.J., Waldman, A.S. & Liskay, R.M. Homologous recombination in mammalian cells. Annu. Rev. Genet. 23, 199–225 (1989).
Capecchi, M.R. Altering the genome by homologous recombination. Science 244, 1288–1292 (1989).
Thyagarajan, B., McCormick-Graham, M., Romero, D.P. & Campbell, C. Characterization of homologous DNA recombination activity in normal and immortal mammalian cells. Nucleic Acids Res. 24, 4084–4091 (1996).
Yang, Y. et al. Targeted disruption of the murine Fanconi anemia gene, Fancg/Xrcc9. Blood 98, 3435–3440 (2001).
Acknowledgements
We thank Naifang Lu and Jeannie T. Lee for help in developing the FISH protocol, Naifang Lu for blastocyst injections, and Vidya Kunjathoor, Yanhong Ma, and Amy Stirman for assistance. This work was supported by grants from the US National Institutes of Health (AI27849, AI46731, and HL66678 to B.S.) and a postdoctoral fellowship from the Cancer Research Institute (to Y.Y.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yang, Y., Seed, B. Site-specific gene targeting in mouse embryonic stem cells with intact bacterial artificial chromosomes. Nat Biotechnol 21, 447–451 (2003). https://doi.org/10.1038/nbt803
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt803
This article is cited by
-
Beyond editing to writing large genomes
Nature Reviews Genetics (2017)
-
A novel transchromosomic system: stable maintenance of an engineered Mb-sized human genomic fragment translocated to a mouse chromosome terminal region
Transgenic Research (2014)
-
Mice lacking Dok-1, Dok-2, and Dok-3 succumb to aggressive histiocytic sarcoma
Laboratory Investigation (2010)
-
Integration of functional bacterial artificial chromosomes into human cord blood-derived multipotent stem cells
Gene Therapy (2009)
-
A caveat in mouse genetic engineering: ectopic gene targeting in ES cells by bidirectional extension of the homology arms of a gene replacement vector carrying human PARP-1
Transgenic Research (2009)