Abstract
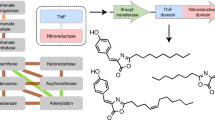
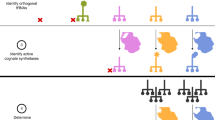
In all genome-sequencing projects completed to date, a considerable number of 'gaps' have been found in the biochemical pathways of the respective species. In many instances, missing enzymes are displaced by analogs, functionally equivalent proteins that have evolved independently and lack sequence and structural similarity. Here we fill such gaps by analyzing anticorrelating occurrences of genes across species. Our approach, applied to the thiamin biosynthesis pathway comprising approximately 15 catalytic steps, predicts seven instances in which known enzymes have been displaced by analogous proteins. So far we have verified four predictions by genetic complementation, including three proteins for which there was no previous experimental evidence of a role in the thiamin biosynthesis pathway. For one hypothetical protein, biochemical characterization confirmed the predicted thiamin phosphate synthase (ThiE) activity. The results demonstrate the ability of our computational approach to predict specific functions without taking into account sequence similarity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Burdick, D. in Kirk-Othmer Encyclopedia of Chemical Technology, vol. 25 (ed. Howe-Grant, M.) 152–171 (Wiley, New York, 1998).
Fenster, R. Feed Additives: A Global Market Study (PJB Publications, Richmond, Surrey, UK, 2001).
Begley, T.P. et al. Thiamin biosynthesis in prokaryotes. Arch. Microbiol. 171, 293–300 (1999).
Hohmann, S. & Meacock, P.A. Thiamin metabolism and thiamin diphosphate-dependent enzymes in the yeast Saccharomyces cerevisiae: genetic regulation. Biochim. Biophys. Acta 1385, 201–219 (1998).
White, R.L. & Spenser, I.D. in Escherichia coli and Salmonella. Cellular and Molecular Biology, edn. 2, vol. 1 (ed. Neidhardt, F.C.) 680–686 (ASM Press, Washington, DC, 1996).
Xi, J., Ge, Y., Kinsland, C., McLafferty, F.W. & Begley, T.P. Biosynthesis of the thiazole moiety of thiamin in Escherichia coli: identification of an acyldisulfide-linked protein-protein conjugate that is functionally analogous to the ubiquitin/E1 complex. Proc. Natl. Acad. Sci. USA 98, 8513–8518 (2001).
Allen, S., Zilles, J.L. & Downs, D.M. Metabolic flux in both the purine mononucleotide and histidine biosynthetic pathways can influence synthesis of the hydroxymethyl pyrimidine moiety of thiamine in Salmonella enterica. J. Bacteriol. 184, 6130–6137 (2002).
Fitch, W.M. Distinguishing homologous from analogous proteins. Syst. Zool. 19, 99–113 (1970).
Koonin, E.V., Mushegian, A.R. & Bork, P. Non-orthologous gene displacement. Trends Genet. 12, 334–336 (1996).
Enright, A.J., Iliopoulos, I., Kyrpides, N.C. & Ouzounis, C.A. Protein interaction maps for complete genomes based on gene fusion events. Nature 402, 86–90 (1999).
Marcotte, E.M. et al. Detecting protein function and protein-protein interactions from genome sequences. Science 285, 751–753 (1999).
Dandekar, T., Snel, B., Huynen, M. & Bork, P. Conservation of gene order: a fingerprint of proteins that physically interact. Trends Biochem. Sci. 23, 324–328 (1998).
Overbeek, R., Fonstein, M., D'Souza, M., Pusch, G.D. & Maltsev, N. The use of gene clusters to infer functional coupling. Proc. Natl. Acad. Sci. USA 96, 2896–2901 (1999).
Pellegrini, M., Marcotte, E.M., Thompson, M.J., Eisenberg, D. & Yeates, T.O. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc. Natl. Acad. Sci. USA 96, 4285–4288 (1999).
Galperin, M.Y. & Koonin, E.V. Who's your neighbor? New computational approaches for functional genomics. Nat. Biotechnol. 18, 609–613 (2000).
Daugherty, M., Vonstein, V., Overbeek, R. & Osterman, A. Archaeal shikimate kinase, a new member of the GHMP-kinase family. J. Bacteriol. 183, 292–300 (2001).
Dynes, J.L. & Firtel, R.A. Molecular complementation of a genetic marker in Dictyostelium using a genomic DNA library. Proc. Natl. Acad. Sci. USA 86, 7966–7970 (1989).
Myllykallio, H. et al. An alternative flavin-dependent mechanism for thymidylate synthesis. Science 297, 105–107 (2002).
Vander Horn, P.B., Backstrom, A.D., Stewart, V. & Begley, T.P. Structural genes for thiamine biosynthetic enzymes (thiCEFGH) in Escherichia coli K-12. J. Bacteriol. 175, 982–992 (1993).
Webb, E. & Downs, D. Characterization of thiL, encoding thiamin-monophosphate kinase, in Salmonella typhimurium. J. Biol. Chem. 272, 15702–15707 (1997).
Nosaka, K., Kaneko, Y., Nishimura, H. & Iwashima, A. Isolation and characterization of a thiamin pyrophosphokinase gene, THI80, from Saccharomyces cerevisiae. J. Biol. Chem. 268, 17440–17447 (1993).
Praekelt, U.M., Byrne, K.L. & Meacock, P.A. Regulation of THI4 (MOL1), a thiamine-biosynthetic gene of Saccharomyces cerevisiae. Yeast 10, 481–490 (1994).
Pang, A.S., Nathoo, S. & Wong, S.L. Cloning and characterization of a pair of novel genes that regulate production of extracellular enzymes in Bacillus subtilis. J. Bacteriol. 173, 46–54 (1991).
Miranda-Rios, J. et al. Expression of thiamin biosynthetic genes (thiCOGE) and production of symbiotic terminal oxidase cbb3 in Rhizobium etli. J. Bacteriol. 179, 6887–6893 (1997).
Rodionov, D.A., Vitreschak, A.G., Mironov, A.A. & Gelfand, M.S. Comparative genomics of thiamin biosynthesis in procaryotes. New genes and regulatory mechanisms. J. Biol. Chem. 277, 48949–48959 (2002).
Petersen, L.A. & Downs, D.M. Identification and characterization of an operon in Salmonella typhimurium involved in thiamine biosynthesis. J. Bacteriol. 179, 4894–4900 (1997).
Chiu, H.J., Reddick, J.J., Begley, T.P. & Ealick, S.E. Crystal structure of thiamin phosphate synthase from Bacillus subtilis at 1.25 A resolution. Biochemistry 38, 6460–6470 (1999).
Baker, L.J., Dorocke, J.A., Harris, R.A. & Timm, D.E. The crystal structure of yeast thiamin pyrophosphokinase. Structure (Camb) 9, 539–546 (2001).
Machado, C.R. et al. Dual role for the yeast THI4 gene in thiamine biosynthesis and DNA damage tolerance. J. Mol. Biol. 273, 114–121 (1997).
Kim, Y.S. et al. A Brassica cDNA clone encoding a bifunctional hydroxymethylpyrimidine kinase/thiamin-phosphate pyrophosphorylase involved in thiamin biosynthesis. Plant Mol. Biol. 37, 955–966 (1998).
Gourley, D.G. et al. The two types of 3-dehydroquinase have distinct structures but catalyze the same overall reaction. Nat. Struct. Biol. 6, 521–525 (1999).
Snel, B., Lehmann, G., Bork, P. & Huynen, M.A. STRING: a web-server to retrieve and display the repeatedly occurring neighbourhood of a gene. Nucleic Acids Res. 28, 3442–3444 (2000).
von Mering, C. et al. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 31, 258–261 (2003).
Vandeyar, M.A. & Zahler, S.A. Chromosomal insertions of Tn917 in Bacillus subtilis. J. Bacteriol. 167, 530–534 (1986).
Tatusov, R.L., Koonin, E.V. & Lipman, D.J. A genomic perspective on protein families. Science 278, 631–637 (1997).
Schwartz, C.J., Djaman, O., Imlay, J.A. & Kiley, P.J. The cysteine desulfurase, IscS, has a major role in in vivo Fe-S cluster formation in Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 9009–9014 (2000).
Huynen, M.A. & Bork, P. Measuring genome evolution. Proc. Natl. Acad. Sci. USA 95, 5849–5856 (1998).
Karp, P.D. Pathway databases: a case study in computational symbolic theories. Science 293, 2040–2044 (2001).
Acknowledgements
This work was supported in part by grants from the Consejo Nacional de Ciencia y Tecnología (Mexico), Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México (Mexico), Netherlands Organization for Scientific Research, the Deutsche Forschungsgemeinschaft (Germany) and Bundesministerium für Forschung und Bildung (Germany). E.M. thanks the Alexander von Humboldt Stiftung. We thank R. Hernandez and P. Gaytan for technical assistance. We are indebted to T.P. Begley for providing HMP and a plasmid with the thiD gene from E. coli and to D.M. Downs, M. Soberón, J. Miranda and R. Russell for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Morett, E., Korbel, J., Rajan, E. et al. Systematic discovery of analogous enzymes in thiamin biosynthesis. Nat Biotechnol 21, 790–795 (2003). https://doi.org/10.1038/nbt834
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt834
This article is cited by
-
Discovery of an ene-reductase for initiating flavone and flavonol catabolism in gut bacteria
Nature Communications (2021)
-
The identification of an integral membrane, cytochrome c urate oxidase completes the catalytic repertoire of a therapeutic enzyme
Scientific Reports (2015)
-
Inferring functional modules of protein families with probabilistic topic models
BMC Bioinformatics (2011)
-
Non-homologous isofunctional enzymes: A systematic analysis of alternative solutions in enzyme evolution
Biology Direct (2010)
-
Inferring modules of functionally interacting proteins using the Bond Energy Algorithm
BMC Bioinformatics (2008)