Abstract
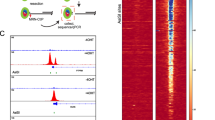
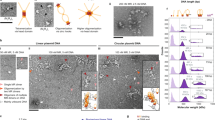
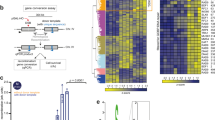
The tumour-suppressor gene ATM, mutations in which cause the human genetic disease ataxia telangiectasia (A-T), encodes a key protein kinase that controls the cellular response to DNA double-strand breaks (DSBs)1,2. DNA DSBs caused by ionizing radiation or chemicals result in rapid ATM autophosphorylation, leading to checkpoint activation and phosphorylation of substrates that regulate cell-cycle progression, DNA repair, transcription and cell death3. However, the precise mechanism by which damaged DNA induces ATM and checkpoint activation remains unclear. Here, we demonstrate that linear DNA fragments added to Xenopus egg extracts mimic DSBs in genomic DNA and provide a platform for ATM autophosphorylation and activation. ATM autophosphorylation and phosphorylation of its substrate NBS1 are dependent on DNA fragment length and the concentration of DNA ends. The minimal DNA length required for efficient ATM autophosphorylation is ∼200 base pairs, with cooperative autophosphorylation induced by DNA fragments of at least 400 base pairs. Importantly, full ATM activation requires it to bind to DNA regions flanking DSB ends. These findings reveal a direct role for DNA flanking DSB ends in ATM activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Abraham, R. T. Cell cycle checkpoint signaling through the ATM and ATR kinases. Genes Dev. 15, 2177–2196 (2001).
Shiloh, Y. ATM and related protein kinases: safeguarding genome integrity. Nature Rev. Cancer 3, 155–168 (2003).
Bakkenist, C. J. & Kastan, M. B. Initiating cellular stress responses. Cell 118, 9–17 (2004).
Costanzo, V., Robertson, K. & Gautier, J. Xenopus cell-free extracts to study the DNA damage response. Methods Mol. Biol. 280, 213–227 (2004).
Petersen, P. et al. Protein phosphatase 2A antagonizes ATM and ATR in a Cdk2- and Cdc7-independent DNA damage checkpoint. Mol. Cell. Biol. 26, 1997–2011 (2006).
You, Z., Chahwan, C., Bailis, J., Hunter, T. & Russell, P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol. Cell. Biol. 25, 5363–5379 (2005).
Almouzni, G. & Mechali, M. Assembly of spaced chromatin involvement of ATP and DNA topoisomerase activity. EMBO J. 7, 4355–4365 (1988).
Ladoux, B. et al. Fast kinetics of chromatin assembly revealed by single-molecule videomicroscopy and scanning force microscopy. Proc. Natl Acad. Sci. USA 97, 14251–14256 (2000).
Bakkenist, C. J. & Kastan, M. B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421, 499–506 (2003).
Pellegrini, M. et al. Autophosphorylation at serine 1987 is dispensable for murine Atm activation in vivo. Nature 443, 222–225 (2006).
Kastan, M. B. & Lim, D. S. The many substrates and functions of ATM. Nature Rev. Mol. Cell. Biol. 1, 179–186 (2000).
Johnson, S. A., You, Z. & Hunter, T. Monitoring ATM kinase activity in living cells. DNA Repair 6, 1277–1284 (2007).
Goodarzi, A. A. et al. Autophosphorylation of ataxia-telangiectasia mutated is regulated by protein phosphatase 2A. EMBO J. 23, 4451–4461 (2004).
Lukas, C., Falck, J., Bartkova, J., Bartek, J. & Lukas, J. Distinct spatiotemporal dynamics of mammalian checkpoint regulators induced by DNA damage. Nature Cell Biol. 5, 255–260 (2003).
McSherry, T. D. & Mueller, P. R. Xenopus Cds1 is regulated by DNA-dependent protein kinase and ATR during the cell cycle checkpoint response to double-stranded DNA ends. Mol. Cell. Biol. 24, 9968–9985 (2004).
Lou, Z. et al. MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals. Mol. Cell 21, 187–200 (2006).
Stucki, M. & Jackson, S. P. γH2AX and MDC1: anchoring the DNA-damage-response machinery to broken chromosomes. DNA Repair 5, 534–543 (2006).
Cerosaletti, K., Wright, J. & Concannon, P. Active role for nibrin in the kinetics of atm activation. Mol. Cell. Biol. 26, 1691–1699 (2006).
Difilippantonio, S. et al. Role of Nbs1 in the activation of the Atm kinase revealed in humanized mouse models. Nature Cell Biol. 7, 675–685 (2005).
Berkovich, E., Monnat, R. J. Jr. & Kastan, M. B. Roles of ATM and NBS1 in chromatin structure modulation and DNA double-strand break repair. Nature Cell Biol. 9, 683–690 (2007).
Shroff, R. et al. Distribution and dynamics of chromatin modification induced by a defined DNA double-strand break. Curr. Biol. 14, 1703–1711 (2004).
Meek, K., Gupta, S., Ramsden, D. A. & Lees-Miller, S. P. The DNA-dependent protein kinase: the director at the end. Immunol. Rev. 200, 132–141 (2004).
Pazin, M. J., Bhargava, P., Geiduschek, E. P. & Kadonaga, J. T. Nucleosome mobility and the maintenance of nucleosome positioning. Science 276, 809–812 (1997).
Dupré, A., Boyer-Chatenet, L. & Gautier, J. Two-step activation of ATM by DNA and the Mre11-Rad50-Nbs1 complex. Nature Struct. Mol. Biol. 13, 451–457 (2006).
Robertson, K., Hensey, C. & Gautier, J. Isolation and characterization of Xenopus ATM (X-ATM): expression, localization, and complex formation during oogenesis and early development. Oncogene 18, 7070–7079 (1999).
Lee, J. H. & Paull, T. T. ATM activation by DNA double-strand breaks through the Mre11–Rad50–Nbs1 complex. Science 308, 551–554 (2005).
Stucki, M. et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123, 1213–1226 (2005).
You, Z., Kong, L. & Newport, J. The role of single-stranded DNA and polymerase alpha in establishing the ATR, Hus1 DNA replication checkpoint. J. Biol. Chem. 277, 27088–27093 (2002).
Hekmat-Nejad, M., You, Z., Yee, M. C., Newport, J. W. & Cimprich, K. A. Xenopus ATR is a replication-dependent chromatin-binding protein required for the DNA replication checkpoint. Curr. Biol. 10, 1565–1573 (2000).
Dilworth, S. M., Black, S. J. & Laskey, R. A. Two complexes that contain histones are required for nucleosome assembly in vitro: role of nucleoplasmin and N1 in Xenopus egg extracts. Cell 51, 1009–1018 (1987).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Loayza, D. & De Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 423, 1013–1018 (2003).
Blow, J. J., Gillespie, P. J., Francis, D. & Jackson, D. A. Replication origins in Xenopus egg extract are 5–15 kilobases apart and are activated in clusters that fire at different times. J. Cell Biol. 152, 15–25 (2001).
Jackson, D. A. & Pombo, A. Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J. Cell Biol. 140, 1285–1295 (1998).
Sullivan, B. & Karpen, G. Centromere identity in Drosophila is not determined in vivo by replication timing. J. Cell Biol. 154, 683–690 (2001).
Acknowledgements
This article is dedicated to the memory of John Newport. We thank Matthew Weitzman, Walter Eckhart, Paul Russell and Robert Abraham for critical discussions, James Kadonaga, Dominique Ray-Gallet, Geneviève Almouzni, Paul Labhart, Takeo Kiskimoto, Graeme Smith, Mark O'Connor, Aaron Straight, Andrew Murray and Beth Baber for providing valuable reagents, Ramiro Verdun for technical assistance in dot-blotting, and Andrew Dillin for use of his microscopes. Z.Y. was supported by a Pioneer Fund Postdoctoral Fellowship. T.H. is a Frank and Else Schilling American Cancer Society Research Professor. This work was supported by Public Health Service Grants CA14195 and CA80100 from the National Cancer Institute (T.H.).
Author information
Authors and Affiliations
Contributions
Z.Y. designed and performed the majority of the experiments with contributions from T.H., J.B. and S.J. S.J. provided the Chk2 substrate used in Fig. 2a, and J.B. performed the chromatin fibre immunostaining experiments in Fig. 3c. S.D. provided critical reagents. Z.Y., J.B. and T.H. wrote the paper.
Corresponding author
Supplementary information
Supplementary Information
Supplementary figures S1, S2, S3, S4, S5, S6 and S7 (PDF 670 kb)
Rights and permissions
About this article
Cite this article
You, Z., Bailis, J., Johnson, S. et al. Rapid activation of ATM on DNA flanking double-strand breaks. Nat Cell Biol 9, 1311–1318 (2007). https://doi.org/10.1038/ncb1651
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1651
This article is cited by
-
Structure-function analysis of TOPBP1’s role in ATR signaling using the DSB-mediated ATR activation in Xenopus egg extracts (DMAX) system
Scientific Reports (2021)
-
HDAC1 and HDAC2 integrate checkpoint kinase phosphorylation and cell fate through the phosphatase-2A subunit PR130
Nature Communications (2018)
-
Destabilization of linker histone H1.2 is essential for ATM activation and DNA damage repair
Cell Research (2018)
-
Chromosome missegregation during anaphase triggers p53 cell cycle arrest through histone H3.3 Ser31 phosphorylation
Nature Cell Biology (2016)
-
Intracellular calcium regulates nonsense-mediated mRNA decay
Nature Medicine (2014)