Abstract
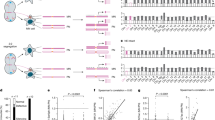
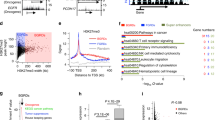
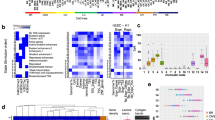
Silencing of individual genes can occur by genetic and epigenetic processes during carcinogenesis, but the underlying mechanisms remain unclear. By creating an integrated prostate cancer epigenome map using tiling arrays, we show that contiguous regions of gene suppression commonly occur through long-range epigenetic silencing (LRES). We identified 47 LRES regions in prostate cancer, typically spanning about 2 Mb and harbouring approximately 12 genes, with a prevalence of tumour suppressor and miRNA genes. Our data reveal that LRES is associated with regional histone deacetylation combined with subdomains of different epigenetic remodelling patterns, which include re-enforcement, gain or exchange of repressive histone, and DNA methylation marks. The transcriptional and epigenetic state of genes in normal prostate epithelial and human embryonic stem cells can play a critical part in defining the mode of cancer-associated epigenetic remodelling. We propose that consolidation or effective reduction of the cancer genome commonly occurs in domains through a combination of LRES and LOH or genomic deletion, resulting in reduced transcriptional plasticity within these regions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Baylin, S. B. et al. Aberrant patterns of DNA methylation, chromatin formation and gene expression in cancer. Hum. Mol. Genet. 10, 687–692 (2001).
Bird, A. P. & Wolffe, A. P. Methylation-induced repression—belts, braces, and chromatin. Cell 99, 451–454 (1999).
Jones, P. A. & Laird, P. W. Cancer epigenetics comes of age. Nature Genet. 21, 163–167 (1999).
Lund, A. H. & van Lohuizen, M. Epigenetics and cancer. Genes Dev. 18, 2315–2335 (2004).
Bracken, A. P., Dietrich, N., Pasini, D., Hansen, K. H. & Helin, K. Genome-wide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 20, 1123–1136 (2006).
Sparmann, A. & van Lohuizen, M. Polycomb silencers control cell fate, development and cancer. Nature Rev. Cancer 6, 846–856 (2006).
Schlesinger, Y. et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nature Genet. 39, 232–236 (2007).
Widschwendter, M. et al. Epigenetic stem cell signature in cancer. Nature Genet. 39, 157–158 (2007).
Santos-Reboucas, C. B. & Pimentel, M. M. Implication of abnormal epigenetic patterns for human diseases. Eur. J. Hum. Genet. 15, 10–17 (2007).
Ohm, J. E. et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nature Genet. 39, 237–242 (2007).
Ting, A. H., McGarvey, K. M. & Baylin, S. B. The cancer epigenome—components and functional correlates. Genes Dev. 20, 3215–3231 (2006).
Frigola, J. et al. Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nature Genet. 38, 540–549 (2006).
Smith, J. S. & Costello, J. F. A broad band of silence. Nature Genet. 38, 504–506 (2006).
Nie, Y. et al. DNA hypermethylation is a mechanism for loss of expression of the HLA class I genes in human esophageal squamous cell carcinomas. Carcinogenesis 22, 1615–1623 (2001).
van Noesel, M. M. et al. Clustering of hypermethylated genes in neuroblastoma. Genes, Chrom. Cancer 38, 226–233 (2003).
Palmisano, W. A. et al. Aberrant promoter methylation of the transcription factor genes PAX5 alpha and beta in human cancers. Cancer Res. 63, 4620–4625 (2003).
Novak, P. et al. Epigenetic inactivation of the HOXA gene cluster in breast cancer. Cancer Res. 66, 10664–10670 (2006).
Stransky, N. et al. Regional copy number-independent deregulation of transcription in cancer. Nature Genet. 38, 1386–1396 (2006).
Hitchins, M. P. et al. Epigenetic inactivation of a cluster of genes flanking MLH1 in microsatellite-unstable colorectal cancer. Cancer Res. 67, 9107–9116 (2007).
Novak, P. et al. Agglomerative epigenetic aberrations are a common event in human breast cancer. Cancer Res. 68, 8616–8625 (2008).
Kim, H. et al. The retinoic acid synthesis gene ALDH1a2 is a candidate tumor suppressor in prostate cancer. Cancer Res. 65, 8118–8124 (2005).
Dhanasekaran, S. M. et al. Delineation of prognostic biomarkers in prostate cancer. Nature 412, 822–826 (2001).
Dhanasekaran, S. M. et al. Molecular profiling of human prostate tissues: insights into gene expression patterns of prostate development during puberty. FASEB J. 19, 243–245 (2005).
Lapointe, J. et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc. Natl Acad. Sci. USA 101, 811–816 (2004).
Luo, J. et al. Human prostate cancer and benign prostatic hyperplasia: molecular dissection by gene expression profiling. Cancer Res. 61, 4683–4688 (2001).
Singh, D. et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1, 203–209 (2002).
Tomlins, S. A. et al. Integrative molecular concept modeling of prostate cancer progression. Nature Genet. 39, 41–51 (2007).
Vanaja, D. K., Cheville, J. C., Iturria, S. J. & Young, C. Y. Transcriptional silencing of zinc finger protein 185 identified by expression profiling is associated with prostate cancer progression. Cancer Res. 63, 3877–3882 (2003).
Welsh, J. B. et al. Analysis of gene expression identifies candidate markers and pharmacological targets in prostate cancer. Cancer Res. 61, 5974–5978 (2001).
Yu, Y. P. et al. Gene expression alterations in prostate cancer predicting tumor aggression and preceding development of malignancy. J. Clin. Oncol. 22, 2790–2799 (2004).
Feltus, F. A., Lee, E. K., Costello, J. F., Plass, C. & Vertino, P. M. DNA motifs associated with aberrant CpG island methylation. Genomics 87, 572–579 (2006).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Engstrom, P. G., Fredman, D. & Lenhard, B. Ancora: a web resource for exploring highly conserved noncoding elements and their association with developmental regulatory genes. Genome Biol. 9, R34 (2008).
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nature Genet. 40, 499–507 (2008).
Gupta, P. B., Chaffer, C. L. & Weinberg, R. A. Cancer stem cells: mirage or reality? Nature Med. 15, 1010–1012 (2009).
Cher, M. L. et al. Genetic alterations in untreated metastases and androgen-independent prostate cancer detected by comparative genomic hybridization and allelotyping. Cancer Res. 56, 3091–3102 (1996).
Dumur, C. I. et al. Genome-wide detection of LOH in prostate cancer using human SNP microarray technology. Genomics 81, 260–269 (2003).
Latil, A., Cussenot, O., Fournier, G., Baron, J. C. & Lidereau, R. Loss of heterozygosity at 7q31 is a frequent and early event in prostate cancer. Clin. Cancer Res. 1, 1385–1389 (1995).
Henrique, R. et al. High promoter methylation levels of APC predict poor prognosis in sextant biopsies from prostate cancer patients. Clin. Cancer Res. 13, 6122–6129 (2007).
Kohonen-Corish, M. R. et al. Promoter methylation of the mutated in colorectal cancer gene is a frequent early event in colorectal cancer. Oncogene 26, 4435–4441 (2007).
Guan, M., Zhou, X., Soulitzis, N., Spandidos, D. A. & Popescu, N. C. Aberrant methylation and deacetylation of deleted in liver cancer-1 gene in prostate cancer: potential clinical applications. Clin. Cancer Res. 12, 1412–1419 (2006).
Chene, L. et al. Extensive analysis of the 7q31 region in human prostate tumors supports TES as the best candidate tumor suppressor gene. Int. J. Cancer 111, 798–804 (2004).
Bachmann, N. et al. Expression changes of CAV1 and EZH2, located on 7q31 approximately q36, are rarely related to genomic alterations in primary prostate carcinoma. Cancer Genet. Cytogenet. 182, 103–110 (2008).
Tatarelli, C., Linnenbach, A., Mimori, K. & Croce, C. M. Characterization of the human TESTIN gene localized in the FRA7G region at 7q31.2. Genomics 68, 1–12 (2000).
Tobias, E. S., Hurlstone, A. F., MacKenzie, E., McFarlane, R. & Black, D. M. The TES gene at 7q31.1 is methylated in tumours and encodes a novel growth-suppressing LIM domain protein. Oncogene 20, 2844–2853 (2001).
Friedman, J. M. et al. The putative tumor suppressor microRNA-101 modulates the cancer epigenome by repressing the polycomb group protein EZH2. Cancer Res. 69, 2623–2629 (2009).
Varambally, S. et al. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 322, 1695–1699 (2008).
Fabbri, M. et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl Acad. Sci. USA 104, 15805–15810 (2007).
Gandellini, P. et al. miR-205 Exerts tumor-suppressive functions in human prostate through down-regulation of protein kinase Cepsilon. Cancer Res. 69, 2287–2295 (2009).
Ke, X. S. et al. Genome-wide profiling of histone h3 lysine 4 and lysine 27 trimethylation reveals an epigenetic signature in prostate carcinogenesis. PLoS ONE 4, e4687 (2009).
O'Hagan, H. M., Mohammad, H. P. & Baylin, S. B. Double strand breaks can initiate gene silencing and SIRT1-dependent onset of DNA methylation in an exogenous promoter CpG island. PLoS Genet. 4, e1000155 (2008).
Hong, Z. et al. A polycomb group protein, PHF1, is involved in the response to DNA double-strand breaks in human cell. Nucleic Acids Res. 36, 2939–2947 (2008).
Schotta, G. et al. A chromatin-wide transition to H4K20 monomethylation impairs genome integrity and programmed DNA rearrangements in the mouse. Genes Dev. 22, 2048–2061 (2008).
Gal-Yam, E. N. et al. Frequent switching of Polycomb repressive marks and DNA hypermethylation in the PC3 prostate cancer cell line. Proc. Natl Acad. Sci. USA 105, 12979–12984 (2008).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R. A. & Feinberg, A. P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nature Genet. 41, 246–250 (2009).
Pauler, F. M. et al. H3K27me3 forms BLOCs over silent genes and intergenic regions and specifies a histone banding pattern on a mouse autosomal chromosome. Genome Res. 19, 221–233 (2009).
Pajerowski, J. D., Dahl, K. N., Zhong, F. L., Sammak, P. J. & Discher, D. E. Physical plasticity of the nucleus in stem cell differentiation. Proc. Natl Acad. Sci. USA 104, 15619–15624 (2007).
Spivakov, M. & Fisher, A. G. Epigenetic signatures of stem-cell identity. Nature Rev. Genet. 8, 263–271 (2007).
Feinberg, A. P. Phenotypic plasticity and the epigenetics of human disease. Nature 447, 433–440 (2007).
Wiblin, A. E., Cui, W., Clark, A. J. & Bickmore, W. A. Distinctive nuclear organisation of centromeres and regions involved in pluripotency in human embryonic stem cells. J. Cell Sci. 118, 3861–3868 (2005).
Liu, P. et al. Sex-determining region Y Box 4 is a transforming oncogene in human prostate cancer cells. Cancer Res. 66, 4011–4019 (2006).
Varambally, S. et al. Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 8, 393–406 (2005).
Song, J. Z., Stirzaker, C., Harrison, J., Melki, J. R. & Clark, S. J. Hypermethylation trigger of the glutathione-S-transferase gene (GSTP1) in prostate cancer cells. Oncogene 21, 1048–1061 (2002).
Frigola, J. et al. Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nature Genet. 38, 540–549 (2006).
Johnson, W. E. et al. Model-based analysis of tiling-arrays for ChIP-chip. Proc. Natl Acad. Sci. USA 103, 12457–12462 (2006).
Liu, P. et al. Sex-determining region Y Box 4 is a transforming oncogene in human prostate cancer cells. Cancer Res. 66, 4011–4019 (2006).
Clark, S. J., Harrison, J., Paul, C. L. & Frommer, M. High sensitivity mapping of methylated cytosines. Nucleic Acids Res. 22, 2990–2997 (1994).
Coolen, M. W., Statham, A. L., Gardiner-Garden, M. & Clark, S. J. Genomic profiling of CpG methylation and allelic specificity using quantitative high-throughput mass spectrometry: critical evaluation and improvements. Nucleic Acids Res. 35, e119 (2007).
Varambally, S. et al. Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 8, 393–406 (2005).
Khalili, A. et al. γ-Normal-γ mixture model for dtecting differentially methylated loci in three breast cancer cell lines. Cancer Informatics 3, 43–54 (2007).
Acknowledgements
We thank K. Patterson, P. Molloy and T. Hulf for reviewing the manuscript. We thank the Ramaciotti Centre, University of NSW (Sydney, Australia) for array hybridizations. This work is supported by Cancer Institute NSW (CINSW) program (S.J.C.), CINSW Fellowship (M.W.C.) and CINSW Student (A.L.S.) grants and National Health and Medical Research Council (NH&MRC) project (427614, 481347) and Fellowship grants (S.J.C. and T.S.) and NBCF program grant.
Author information
Authors and Affiliations
Contributions
S.J.C. initiated and supervised the study, and with M.W.C. and C.S., designed the experiments and wrote the paper; M.W.C., C.S., J.Z.S. and A.L.S. performed experiments and helped with data analysis; Z.K. helped with the experiments; M.D.R., T.P.S., P.L, M.C. and W.K. helped with the data analysis; C.S.M., A.N.Y and V.V. prepared the clinical samples.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 989 kb)
Supplementary Information
Additional File 1 (PDF 626 kb)
Supplementary Information
Additional File 2 (PDF 350 kb)
Supplementary Information
Additional File 3 (PDF 1012 kb)
Supplementary Information
Additional File 4 (PDF 2773 kb)
Supplementary Information
Additional Bioinformatics Data (PDF 1235 kb)
Rights and permissions
About this article
Cite this article
Coolen, M., Stirzaker, C., Song, J. et al. Consolidation of the cancer genome into domains of repressive chromatin by long-range epigenetic silencing (LRES) reduces transcriptional plasticity. Nat Cell Biol 12, 235–246 (2010). https://doi.org/10.1038/ncb2023
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2023
This article is cited by
-
Characterisation and reproducibility of the HumanMethylationEPIC v2.0 BeadChip for DNA methylation profiling
BMC Genomics (2024)
-
Extensive fragmentation and re-organization of transcription in Systemic Lupus Erythematosus
Scientific Reports (2020)
-
Constitutively bound CTCF sites maintain 3D chromatin architecture and long-range epigenetically regulated domains
Nature Communications (2020)
-
Epigenetic reprogramming at estrogen-receptor binding sites alters 3D chromatin landscape in endocrine-resistant breast cancer
Nature Communications (2020)
-
The DNA methylation landscape of advanced prostate cancer
Nature Genetics (2020)