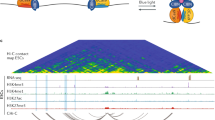
Abstract
Mammalian genomes contain numerous regulatory DNA sites with unknown target genes. We used mice with an extra β-globin locus control region (LCR) to investigate how a regulator searches the genome for target genes. We find that the LCR samples a restricted nuclear subvolume, wherein it preferentially contacts genes controlled by shared transcription factors. No contacted gene is detectably upregulated except for endogenous β-globin genes located on another chromosome. This demonstrates genetically that mammalian trans activation is possible, but suggests that it will be rare. Trans activation occurs not pan-cellularly, but in ‘jackpot’ cells enriched for the interchromosomal interaction. Therefore, cell-specific long-range DNA contacts can cause variegated expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Amano, T. et al. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev. Cell 16, 47–57 (2009).
Kleinjan, D. J. & Coutinho, P. Cis-ruption mechanisms: disruption of cis-regulatory control as a cause of human genetic disease. Brief Funct. Genomic Proteom. 8, 317–332 (2009).
Epner, E. et al. The β-globin LCR is not necessary for an open chromatin structure or developmentally regulated transcription of the native mouse β-globin locus. Mol. Cell 2, 447–455 (1998).
Grosveld, F., van Assendelft, G. B., Greaves, D. R. & Kollias, G. Position-independent, high-level expression of the human β-globin gene in transgenic mice. Cell 51, 975–985 (1987).
Wasylyk, B., Wasylyk, C., Augereau, P. & Chambon, P. The SV40 72 bp repeat preferentially potentiates transcription starting from proximal natural or substitute promoter elements. Cell 32, 503–514 (1983).
Wijgerde, M., Grosveld, F. & Fraser, P. Transcription complex stability and chromatin dynamics in vivo . Nature 377, 209–213 (1995).
Carter, D., Chakalova, L., Osborne, C. S., Dai, Y. F. & Fraser, P. Long-range chromatin regulatory interactions in vivo . Nat. Genet. 32, 623–626 (2002).
Tolhuis, B., Palstra, R. J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
de Laat, W. & Grosveld, F. Spatial organization of gene expression: the active chromatin hub. Chromosome Res. 11, 447–459 (2003).
Spilianakis, C. G. & Flavell, R. A. Long-range intrachromosomal interactions in the T helper type 2 cytokine locus. Nat. Immunol. 5, 1017–1027 (2004).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J. A., Wood, W. G. & Higgs, D. R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Brown, K. E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997).
Chambeyron, S., Da Silva, N. R., Lawson, K. A. & Bickmore, W. A. Nuclear re-organisation of the Hoxb complex during mouse embryonic development. Development 132, 2215–2223 (2005).
Skok, J. A. et al. Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nat. Immunol. 2, 848–854 (2001).
Hewitt, S. L. et al. Association between the Igk and Igh immunoglobulin loci mediated by the 3’ Igk enhancer induces ‘decontraction’ of the Igh locus in pre-B cells. Nat. Immunol. 9, 396–404 (2008).
Lundgren, M. et al. Transcription factor dosage affects changes in higher order chromatin structure associated with activation of a heterochromatic gene. Cell 103, 733–743 (2000).
Noordermeer, D. et al. Transcription and chromatin organization of a housekeeping gene cluster containing an integrated β-globin locus control region. PLoS Genet. 4, e1000016 (2008).
Ragoczy, T., Bender, M. A., Telling, A., Byron, R. & Groudine, M. The locus control region is required for association of the murine β-globin locus with engaged transcription factories during erythroid maturation. Genes Dev. 20, 1447–1457 (2006).
Schoenfelder, S. et al. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42, 53–61 (2010).
Apostolou, E. & Thanos, D. Virus infection induces NF-κB-dependent interchromosomal associations mediating monoallelic IFN-β gene expression. Cell 134, 85–96 (2008).
Ling, J. Q. et al. CTCF mediates interchromosomal colocalization between Igf2/H19 and Wsb1/Nf1. Science 312, 269–272 (2006).
Lomvardas, S. et al. Interchromosomal interactions and olfactory receptor choice. Cell 126, 403–413 (2006).
Sandhu, K. S. et al. Nonallelic transvection of multiple imprinted loci is organized by the H19 imprinting control region during germline development. Genes Dev. 23, 2598–2603 (2009).
Spilianakis, C. G., Lalioti, M. D., Town, T., Lee, G. R. & Flavell, R. A. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005).
Lewis, E. B. The theory and application of a new method of detectingchromosomal rearrangements in Drosophila melanogaster . Am. Naturalist 88, 225–239 (1954).
Pirrotta, V. Transvection and chromosomal trans-interaction effects. Biochim. Biophys. Acta 1424, M1-8 (1999).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, 826–842 (2005).
Bacher, C. P. et al. Transient colocalization of X-inactivation centres accompanies the initiation of X inactivation. Nat. Cell Biol. 8, 293–299 (2006).
Hewitt, S. L. et al. RAG-1 and ATM coordinate monoallelic recombinationand nuclear positioning of immunoglobulin loci. Nat. Immunol. 10, 655–664 (2009).
Xu, N., Donohoe, M. E., Silva, S. S. & Lee, J. T. Evidence that homologous X-chromosome pairing requires transcription and Ctcf protein. Nat. Genet. 39, 1390–1396 (2007).
Branco, M. R. & Pombo, A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol. 4, 780–788 (2006).
Strouboulis, J., Dillon, N. & Grosveld, F. Developmental regulation of a complete 70-kb human β-globin locus in transgenic mice. Genes Dev. 6, 1857–1864 (1992).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
de Wit, E., Braunschweig, U., Greil, F., Bussemaker, H. J. & van Steensel, B. Global chromatin domain organization of the Drosophila genome. PLoS Genet. 4, e1000045 (2008).
Simonis, M., Kooren, J. & de Laat, W. An evaluation of 3C-based methods to capture DNA interactions. Nat. Methods 4, 895–901 (2007).
Drissen, R. et al. The erythroid phenotype of EKLF-null mice: defects in hemoglobin metabolism and membrane stability. Mol. Cell Biol. 25, 5205–5214 (2005).
Hodge, D. et al. A global role for EKLF in definitive and primitive erythropoiesis. Blood 107, 3359–3370 (2006).
Welch, J. J. et al. Global regulation of erythroid gene expression by transcription factor GATA-1. Blood 104, 3136–3147 (2004).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ . Science 280, 585–590 (1998).
Cajiao, I., Zhang, A., Yoo, E. J., Cooke, N. E. & Liebhaber, S. A. Bystander gene activation by a locus control region. EMBO J. 23, 3854–3863 (2004).
Lower, K. M. et al. Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc. Natl Acad. Sci. USA 106, 21771–21776 (2009).
Fuss, S. H., Omura, M. & Mombaerts, P. Local and cis effects of the H element on expression of odorant receptor genes in mouse. Cell 130, 373–384 (2007).
Nishizumi, H., Kumasaka, K., Inoue, N., Nakashima, A. & Sakano, H. Deletion of the core-H region in mice abolishes the expression of three proximal odorant receptor genes in cis. Proc. Natl. Acad. Sci. USA 104, 20067–20072 (2007).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Muller, H. J. Types of visible variations induced by X-rays in Drosophila . J. Genet. 22, 299–334 (1930).
Solovei, I. et al. Spatial preservation of nuclear chromatin architecture during three-dimensional fluorescence in situ hybridization (3D-FISH). Exp. Cell Res. 276, 10–23 (2002).
Levsky, J. M., Shenoy, S. M., Pezo, R. C. & Singer, R. H. Single-cell gene expression profiling. Science 297, 836–840 (2002).
Pau, G., Fuchs, F., Sklyar, O., Boutros, M. & Huber, W. EBImage–an R package for image processing with applications to cellular phenotypes. Bioinformatics 26, 979–981 (2010).
Kolountzakis, M. N. & Kutulakos, K. N. Fast computation of the Euclidian distance maps for binary images. Inform. Process. Lett. 43, 181–184 (1992).
Vincent, L. & Soille, P. Watersheds in digital spaces: an efficient algorithm based on immersion simulations. Pattern Analysis and Machine Intelligence, IEEE Trans. 13, 583–598 (2002).
Smyth, G. K. & Speed, T. Normalization of cDNA microarray data. Methods 31, 265–273 (2003).
Acknowledgements
We thank J. Marteijn and W. Vermeulen for providing Rad23a knockout material, Y. Oz for counting FISH slides, V. Buckle for providing BAC probes, J. van Rheenen and the Hubrecht Imaging Center for help with image analysis and E. Splinter and other members of the group for assistance. This work was financially supported by grants from the Dutch Scientific Organization (NWO) (91204082 and 935170621) and a European Research Council Starting Grant (209700, ‘4C’) to W.d.L.
Author information
Authors and Affiliations
Contributions
D.N. and W.d.L. designed the experiments, analysed the data and, with help from E.d.W., wrote the manuscript. D.N. and P.K. carried out experiments. E.d.W. analysed 4C data and developed the automated FISH image analysis. H.v.d.W. analysed 4C and microarray expression data. M.S. carried out 4C experiments. M.L-J. and R.H.S. designed and synthesized RNA-FISH probes. B.E. and A.d.K. helped with the FISH experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3545 kb)
Supplementary Information
Supplementary Table 1 (XLS 124 kb)
Rights and permissions
About this article
Cite this article
Noordermeer, D., de Wit, E., Klous, P. et al. Variegated gene expression caused by cell-specific long-range DNA interactions. Nat Cell Biol 13, 944–951 (2011). https://doi.org/10.1038/ncb2278
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2278
This article is cited by
-
Chromosomes come together to help mice distinguish odours
Nature (2019)
-
LHX2- and LDB1-mediated trans interactions regulate olfactory receptor choice
Nature (2019)
-
Transcription factors and 3D genome conformation in cell-fate decisions
Nature (2019)
-
Dynamic adaptation of mesenchymal stem cell physiology upon exposure to surface micropatterns
Scientific Reports (2019)
-
3C and 3C-based techniques: the powerful tools for spatial genome organization deciphering
Molecular Cytogenetics (2018)