Abstract
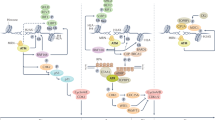
The MRN (Mre11-Rad50-Nbs1)-ATM (ataxia-telangiectasia mutated) pathway is essential for sensing and signaling from DNA double-strand breaks. The MRN complex acts as a DNA damage sensor, maintains genome stability during DNA replication, promotes homology-dependent DNA repair and activates ATM. MRN is essential for cell viability, which has limited functional studies of the complex. Small-molecule inhibitors of MRN could circumvent this experimental limitation and could also be used as cellular radio- and chemosensitization compounds. Using cell-free systems that recapitulate faithfully the MRN-ATM signaling pathway, we designed a forward chemical genetic screen to identify inhibitors of the pathway, and we isolated Z-5-(4-hydroxybenzylidene)-2-imino-1,3-thiazolidin-4-one (mirin, 1) as an inhibitor of MRN. Mirin prevents MRN-dependent activation of ATM without affecting ATM protein kinase activity, and it inhibits Mre11-associated exonuclease activity. Consistent with its ability to target the MRN complex, mirin abolishes the G2/M checkpoint and homology-dependent repair in mammalian cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
13 February 2009
In the version of this article initially published, the reported chemical structure and systematic name for mirin (compound 1), identified from a high-throughput screen, were incorrectly assigned in the commercial library. Investigations by Gartner, Pletnev and Eastman (Correspondence: Nat. Chem. Biol. 5, 129-130, 2009), which have been confirmed by us (Response: Nat. Chem. Biol. 5, 130, 2009), have identified the correct structure of mirin. The systematic name and chemical structure for mirin have been corrected in the HTML and PDF versions of the article, and in the graphical abstract and the PubChem database.
References
Zhou, B.B. & Elledge, S.J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439 (2000).
Kastan, M.B. & Bartek, J. Cell-cycle checkpoints and cancer. Nature 432, 316–323 (2004).
Lavin, M.F. et al. ATM signaling and genomic stability in response to DNA damage. Mutat. Res. 569, 123–132 (2005).
Bakkenist, C.J. & Kastan, M.B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421, 499–506 (2003).
You, Z., Chahwan, C., Bailis, J., Hunter, T. & Russell, P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol. Cell. Biol. 25, 5363–5379 (2005).
Lee, J.H. & Paull, T.T. Direct activation of the ATM protein kinase by the Mre11/Rad50/Nbs1 complex. Science 304, 93–96 (2004).
Lee, J.H. & Paull, T.T. ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science 308, 551–554 (2005).
Carson, C.T. et al. The Mre11 complex is required for ATM activation and the G2/M checkpoint. EMBO J. 22, 6610–6620 (2003).
Difilippantonio, S. et al. Role of Nbs1 in the activation of the Atm kinase revealed in humanized mouse models. Nat. Cell Biol. 7, 675–685 (2005).
Dupre, A., Boyer-Chatenet, L. & Gautier, J. Two-step activation of ATM by DNA and the Mre11-Rad50-Nbs1 complex. Nat. Struct. Mol. Biol. 13, 451–457 (2006).
Kozlov, S.V. et al. Involvement of novel autophosphorylation sites in ATM activation. EMBO J. 25, 3504–3514 (2006).
Gatei, M. et al. ATM-dependent phosphorylation of nibrin in response to radiation exposure. Nat. Genet. 25, 115–119 (2000).
Paull, T.T. & Lee, J.H. The Mre11/Rad50/Nbs1 complex and its role as a DNA double-strand break sensor for ATM. Cell Cycle 4, 737–740 (2005).
Falck, J., Coates, J. & Jackson, S.P. Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature 434, 605–611 (2005).
Uziel, T. et al. Requirement of the MRN complex for ATM activation by DNA damage. EMBO J. 22, 5612–5621 (2003).
Shiloh, Y. ATM and related protein kinases: safeguarding genome integrity. Nat. Rev. Cancer 3, 155–168 (2003).
D'Amours, D. & Jackson, S.P. The Mre11 complex: at the crossroads of dna repair and checkpoint signalling. Nat. Rev. Mol. Cell Biol. 3, 317–327 (2002).
Paull, T.T. & Gellert, M. The 3′ to 5′ exonuclease activity of Mre 11 facilitates repair of DNA double-strand breaks. Mol. Cell 1, 969–979 (1998).
Haber, J.E. The many interfaces of Mre11. Cell 95, 583–586 (1998).
Zhou, B.B. & Bartek, J. Targeting the checkpoint kinases: chemosensitization versus chemoprotection. Nat. Rev. Cancer 4, 216–225 (2004).
Costanzo, V. et al. Reconstitution of an ATM-dependent checkpoint that inhibits chromosomal DNA replication following DNA damage. Mol. Cell 6, 649–659 (2000).
Costanzo, V. et al. Mre11 protein complex prevents double-strand break accumulation during chromosomal DNA replication. Mol. Cell 8, 137–147 (2001).
Peterson, J.R., Lokey, R.S., Mitchison, T.J. & Kirschner, M.W. A chemical inhibitor of N-WASP reveals a new mechanism for targeting protein interactions. Proc. Natl. Acad. Sci. USA 98, 10624–10629 (2001).
Verma, R. et al. Ubistatins inhibit proteasome-dependent degradation by binding the ubiquitin chain. Science 306, 117–120 (2004).
Wignall, S.M. et al. Identification of a novel protein regulating microtubule stability through a chemical approach. Chem. Biol. 11, 135–146 (2004).
Costanzo, V., Paull, T., Gottesman, M. & Gautier, J. Mre11 assembles linear DNA fragments into DNA damage signaling complexes. PLoS Biol. 2, E110 (2004).
Komarov, P.G. et al. A chemical inhibitor of p53 that protects mice from the side effects of cancer therapy. Science 285, 1733–1737 (1999).
Mayer, T.U. Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science 286, 971–974 (1999); comment 286, 913–914 (1999).
Cimprich, K.A., Shin, T.B., Keith, C.T. & Schreiber, S.L. cDNA cloning and gene mapping of a candidate human cell cycle checkpoint protein. Proc. Natl. Acad. Sci. USA 93, 2850–2855 (1996).
Kitagawa, R., Bakkenist, C.J., McKinnon, P.J. & Kastan, M.B. Phosphorylation of SMC1 is a critical downstream event in the ATM-NBS1-BRCA1 pathway. Genes Dev. 18, 1423–1438 (2004).
Lee, J.H. & Paull, T.T. Purification and biochemical characterization of ataxia-telangiectasia mutated and Mre11/Rad50/Nbs1. Methods Enzymol. 408, 529–539 (2006).
Bhaskara, V. et al. Rad50 adenylate kinase activity regulates DNA tethering activities of Mre11/Rad50 complexes. Mol. Cell 25, 647–661 (2007).
Trenz, K., Smith, E., Smith, S. & Costanzo, V. ATM and ATR promote Mre11 dependent restart of collapsed replication forks and prevent accumulation of DNA breaks. EMBO J. 25, 1764–1774 (2006).
Hickson, I. et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 64, 9152–9159 (2004).
Pierce, A.J., Johnson, R.D., Thompson, L.H. & Jasin, M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 13, 2633–2638 (1999).
Esashi, F. et al. CDK-dependent phosphorylation of BRCA2 as a regulatory mechanism for recombinational repair. Nature 434, 598–604 (2005).
Arthur, L.M. et al. Structural and functional analysis of Mre11–3. Nucleic Acids Res. 32, 1886–1893 (2004).
Symington, L.S. Role of RAD52 epistasis group genes in homologous recombination and double-strand break repair. Microbiol. Mol. Biol. Rev. 66, 630–670 (2002).
Smythe, C. & Newport, J.W. Systems for the study of nuclear assembly, DNA replication, and nuclear breakdown in Xenopus laevis egg extracts. Methods Cell Biol. 35, 449–468 (1991).
Acknowledgements
We thank H. Lindsay for Chk2 antibody and B. Stockwell, O. Haccard and the members of the Gautier laboratory for helpful discussions. We thank G. Smith (KuDOS Pharmaceuticals) for the inhibitor KU-55933. We are indebted to J. Decatur and Y. Itagaki of the Chemistry Department for providing NMR and mass spectral analysis of mirin. This work was supported by US National Institutes of Health grants CA95866 and CA92245 (J. Gautier), CA97403 (R. Baer) and GM54668 (M. Jasin).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Methods (PDF 505 kb)
Rights and permissions
About this article
Cite this article
Dupré, A., Boyer-Chatenet, L., Sattler, R. et al. A forward chemical genetic screen reveals an inhibitor of the Mre11–Rad50–Nbs1 complex. Nat Chem Biol 4, 119–125 (2008). https://doi.org/10.1038/nchembio.63
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.63
This article is cited by
-
The DNA damage response pathway regulates the expression of the immune checkpoint CD47
Communications Biology (2023)
-
Farrerol directly activates the deubiqutinase UCHL3 to promote DNA repair and reprogramming when mediated by somatic cell nuclear transfer
Nature Communications (2023)
-
Replication fork uncoupling causes nascent strand degradation and fork reversal
Nature Structural & Molecular Biology (2023)
-
O-GlcNAc transferase couples MRE11 to transcriptionally active chromatin to suppress DNA damage
Journal of Biomedical Science (2022)
-
Genome-wide CRISPR screen identified Rad18 as a determinant of doxorubicin sensitivity in osteosarcoma
Journal of Experimental & Clinical Cancer Research (2022)