Abstract
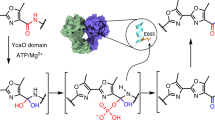
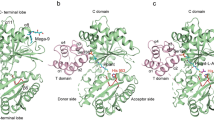
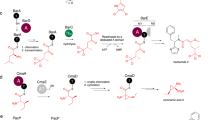
Thiazole/oxazole-modified microcins (TOMMs) encompass a recently defined class of ribosomally synthesized natural products with a diverse set of biological activities. Although TOMM biosynthesis has been investigated for over a decade, the mechanism of heterocycle formation by the synthetase enzymes remains poorly understood. Using substrate analogs and isotopic labeling, we demonstrate that ATP is used to directly phosphorylate the peptide amide backbone during TOMM heterocycle formation. Moreover, we present what is to our knowledge the first experimental evidence that the D-protein component of the heterocycle-forming synthetase (YcaO/domain of unknown function 181 family member), formerly annotated as a docking protein involved in complex formation and regulation, is able to perform the ATP-dependent cyclodehydration reaction in the absence of the other TOMM biosynthetic proteins. Together, these data reveal the role of ATP in the biosynthesis of azole and azoline heterocycles in ribosomal natural products and prompt a reclassification of the enzymes involved in their installation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Melby, J.O., Nard, N.J. & Mitchell, D.A. Thiazole/oxazole-modified microcins: complex natural products from ribosomal templates. Curr. Opin. Chem. Biol. 15, 369–378 (2011).
Li, Y.M., Milne, J.C., Madison, L.L., Kolter, R. & Walsh, C.T. From peptide precursors to oxazole and thiazole-containing peptide antibiotics: microcin B17 synthase. Science 274, 1188–1193 (1996).
Lee, S.W. et al. Discovery of a widely distributed toxin biosynthetic gene cluster. Proc. Natl. Acad. Sci. USA 105, 5879–5884 (2008).
McIntosh, J.A., Donia, M.S. & Schmidt, E.W. Insights into heterocyclization from two highly similar enzymes. J. Am. Chem. Soc. 132, 4089–4091 (2010).
McIntosh, J.A. & Schmidt, E.W. Marine molecular machines: heterocyclization in cyanobactin biosynthesis. ChemBioChem 11, 1413–1421 (2010).
Milne, J.C. et al. Cofactor requirements and reconstitution of microcin B17 synthetase: a multienzyme complex that catalyzes the formation of oxazoles and thiazoles in the antibiotic microcin B17. Biochemistry 38, 4768–4781 (1999).
Kelleher, N.L., Hendrickson, C.L. & Walsh, C.T. Posttranslational heterocyclization of cysteine and serine residues in the antibiotic microcin B17: distributivity and directionality. Biochemistry 38, 15623–15630 (1999).
Milne, J.C., Eliot, A.C., Kelleher, N.L. & Walsh, C.T. ATP/GTP hydrolysis is required for oxazole and thiazole biosynthesis in the peptide antibiotic microcin B17. Biochemistry 37, 13250–13261 (1998).
Schmidt, E.W. et al. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 102, 7315–7320 (2005).
Zamble, D.B., McClure, C.P., Penner-Hahn, J.E. & Walsh, C.T. The McbB component of microcin B17 synthetase is a zinc metalloprotein. Biochemistry 39, 16190–16199 (2000).
Melby, J.O., Dunbar, K.L. & Mitchell, D.A. Selectivity, directionality, and promiscuity in peptide processing from a Bacillus sp. Al Hakam cyclodehydratase. J. Am. Chem. Soc. 134, 5309–5316 (2012).
Yu, Y. et al. Nosiheptide biosynthesis featuring a unique indole side ring formation on the characteristic thiopeptide framework. ACS Chem. Biol. 4, 855–864 (2009).
Wieland Brown, L.C., Acker, M.G., Clardy, J., Walsh, C.T. & Fischbach, M.A. Thirteen posttranslational modifications convert a 14-residue peptide into the antibiotic thiocillin. Proc. Natl. Acad. Sci. USA 106, 2549–2553 (2009).
Morris, R.P. et al. Ribosomally synthesized thiopeptide antibiotics targeting elongation factor Tu. J. Am. Chem. Soc. 131, 5946–5955 (2009).
Liao, R. et al. Thiopeptide biosynthesis featuring ribosomally synthesized precursor peptides and conserved posttranslational modifications. Chem. Biol. 16, 141–147 (2009).
Kelly, W.L., Pan, L. & Li, C. Thiostrepton biosynthesis: prototype for a new family of bacteriocins. J. Am. Chem. Soc. 131, 4327–4334 (2009).
Myers, C.L., Hang, P.C., Ng, G., Yuen, J. & Honek, J.F. Semi-synthetic analogues of thiostrepton delimit the critical nature of tail region modifications in the control of protein biosynthesis and antibacterial activity. Bioorg. Med. Chem. 18, 4231–4237 (2010).
Bowers, A.A., Acker, M.G., Koglin, A. & Walsh, C.T. Manipulation of thiocillin variants by prepeptide gene replacement: structure, conformation, and activity of heterocycle substitution mutants. J. Am. Chem. Soc. 132, 7519–7527 (2010).
Molohon, K.J. et al. Structure determination and interception of biosynthetic intermediates for the plantazolicin class of highly discriminating antibiotics. ACS Chem. Biol. 6, 1307–1313 (2011).
Webb, M.R. A continuous spectrophotometric assay for inorganic phosphate and for measuring phosphate release kinetics in biological systems. Proc. Natl. Acad. Sci. USA 89, 4884–4887 (1992).
Sinha Roy, R., Belshaw, P.J. & Walsh, C.T. Mutational analysis of posttranslational heterocycle biosynthesis in the gyrase inhibitor microcin B17: distance dependence from propeptide and tolerance for substitution in a GSCG cyclizable sequence. Biochemistry 37, 4125–4136 (1998).
Cohn, M. & Hu, A. Isotopic (18O) shift in 31P nuclear magnetic resonance applied to a study of enzyme-catalyzed phosphate-phosphate exchange and phosphate (oxygen)-water exchange reactions. Proc. Natl. Acad. Sci. USA 75, 200–203 (1978).
Roush, R.F., Nolan, E.M., Lohr, F. & Walsh, C.T. Maturation of an Escherichia coli ribosomal peptide antibiotic by ATP-consuming N-P bond formation in microcin C7. J. Am. Chem. Soc. 130, 3603–3609 (2008).
Schrimsher, J.L., Schendel, F.J., Stubbe, J. & Smith, J.M. Purification and characterization of aminoimidazole ribonucleotide synthetase from Escherichia coli. Biochemistry 25, 4366–4371 (1986).
Schendel, F.J., Mueller, E., Stubbe, J., Shiau, A. & Smith, J.M. Formylglycinamide ribonucleotide synthetase from Escherichia coli: cloning, sequencing, overproduction, isolation, and characterization. Biochemistry 28, 2459–2471 (1989).
Zhang, Y., Morar, M. & Ealick, S.E. Structural biology of the purine biosynthetic pathway. Cell. Mol. Life Sci. 65, 3699–3724 (2008).
Li, C., Kappock, T.J., Stubbe, J., Weaver, T.M. & Ealick, S.E. X-ray crystal structure of aminoimidazole ribonucleotide synthetase (PurM), from the Escherichia coli purine biosynthetic pathway at 2.5 A resolution. Structure 7, 1155–1166 (1999).
Morar, M., Anand, R., Hoskins, A.A., Stubbe, J. & Ealick, S.E. Complexed structures of formylglycinamide ribonucleotide amidotransferase from Thermotoga maritima describe a novel ATP binding protein superfamily. Biochemistry 45, 14880–14895 (2006).
Baldwin, J. Rules for ring closure. J. Chem. Soc. Chem. Comm. 734–736 (1976).
Chong, S. et al. Protein splicing involving the Saccharomyces cerevisiae VMA intein. The steps in the splicing pathway, side reactions leading to protein cleavage, and establishment of an in vitro splicing system. J. Biol. Chem. 271, 22159–22168 (1996).
Lee, J.J. et al. Autoproteolysis in hedgehog protein biogenesis. Science 266, 1528–1537 (1994).
Perler, F.B., Xu, M.Q. & Paulus, H. Protein splicing and autoproteolysis mechanisms. Curr. Opin. Chem. Biol. 1, 292–299 (1997).
Ghilarov, D., Serebryakova, M., Shkundina, I. & Severinov, K. A Major portion of DNA gyrase inhibitor microcin B17 undergoes an N,O-peptidyl shift during synthesis. J. Biol. Chem. 286, 26308–26318 (2011).
Ludwig, C., Schwarzer, D. & Mootz, H.D. Interaction studies and alanine scanning analysis of a semi-synthetic split intein reveal thiazoline ring formation from an intermediate of the protein splicing reaction. J. Biol. Chem. 283, 25264–25272 (2008).
Strader, M.B. et al. A proteomic and transcriptomic approach reveals new insight into beta-methylthiolation of Escherichia coli ribosomal protein S12. Mol. Cell. Proteomics 10, M110 005199 (2011).
Acknowledgements
We are grateful to W. van der Donk for technical advice and for suggestions in the preparation of the manuscript. We thank members of the Mitchell lab and M. Marletta for critical review of this manuscript. This work was supported in part by the institutional funds provided by the University of Illinois and the US National Institutes of Health (NIH) (1R01 GM097142 to D.A.M). D.A.M. is also the recipient of the NIH Director's New Innovator Award (DP2 OD008463). K.L.D. was supported by the NIH Training Program in the Chemistry-Biology Interface (2T32 GM070421). J.O.M. was supported by the University of Illinois Department of Chemistry Chinoree T. Kimiyo Enta Fellowship.
Author information
Authors and Affiliations
Contributions
Experiments were designed by D.A.M., K.L.D. and J.O.M. and were performed by K.L.D. and J.O.M. The manuscript was written by D.A.M. and K.L.D. with critical editorial input from J.O.M. The study was conceived and overseen by D.A.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 11904 kb)
Rights and permissions
About this article
Cite this article
Dunbar, K., Melby, J. & Mitchell, D. YcaO domains use ATP to activate amide backbones during peptide cyclodehydrations. Nat Chem Biol 8, 569–575 (2012). https://doi.org/10.1038/nchembio.944
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.944
This article is cited by
-
Caught in the act
Nature Chemical Biology (2023)
-
Electron transfer in protein modifications: from detection to imaging
Science China Chemistry (2023)
-
Functional elucidation of TfuA in peptide backbone thioamidation
Nature Chemical Biology (2021)
-
Posttranslational chemical installation of azoles into translated peptides
Nature Communications (2021)
-
Klebsazolicin inhibits 70S ribosome by obstructing the peptide exit tunnel
Nature Chemical Biology (2017)