Abstract
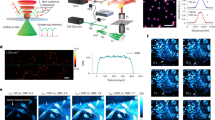
Protease activity is tightly regulated in both normal and disease conditions. However, it is often difficult to monitor the dynamic nature of this regulation in the context of a live cell or whole organism. To address this limitation, we developed a series of quenched activity-based probes (qABPs) that become fluorescent upon activity-dependent covalent modification of a protease target. These reagents freely penetrate cells and allow direct imaging of protease activity in living cells. Targeted proteases are directly identified and monitored biochemically by virtue of the resulting covalent tag, thereby allowing unambiguous assignment of protease activities observed in imaging studies. We report here the design and synthesis of a selective, cell-permeable qABP for the study of papain-family cysteine proteases. This probe is used to monitor real-time protease activity in live human cells with fluorescence microscopy techniques as well as standard biochemical methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Puente, X.S., Sánchez, L.M., Overall, C.M. & López-Otín, C. Human and mouse proteases: a comparative genomic approach. Nat. Rev. Genet. 4, 544–558 (2003).
Baruch, A., Jeffery, D.A. & Bogyo, M. Enzyme activity–it's all about image. Trends Cell Biol. 14, 29–35 (2004).
Berger, A.B., Vitorino, P.M. & Bogyo, M. Activity-based protein profiling: applications to biomarker discovery, in vivo imaging and drug discovery. Am. J. Pharmacogenomics 4, 371–381 (2004).
Speers, A.E. & Cravatt, B.F. Chemical strategies for activity-based proteomics. ChemBioChem 5, 41–47 (2004).
Jeffery, D.A. & Bogyo, M. Chemical proteomics and its application to drug discovery. Curr. Opin. Biotechnol. 14, 87–95 (2003).
Adam, G.C., Sorensen, E.J. & Cravatt, B.F. Chemical strategies for functional proteomics. Mol. Cell. Proteomics 1, 781–790 (2002).
Jessani, N. & Cravatt, B.F. The development and application of methods for activity-based protein profiling. Curr. Opin. Chem. Biol. 8, 54–59 (2004).
Liu, Y., Patricelli, M.P. & Cravatt, B.F. Activity-based protein profiling: the serine hydrolases. Proc. Natl. Acad. Sci. USA 96, 14694–14699 (1999).
Kato, D. et al. Activity-based probes that target diverse cysteine protease families. Nat. Chem. Biol. 1, 33–38 (2005).
Bogyo, M., Shin, S., McMaster, J.S. & Ploegh, H.L. Substrate binding and sequence preference of the proteasome revealed by active-site-directed affinity probes. Chem. Biol. 5, 307–320 (1998).
Bogyo, M., Verhelst, S., Bellingard-Dubouchaud, V., Toba, S. & Greenbaum, D. Selective targeting of lysosomal cysteine proteases with radiolabeled electrophilic substrate analogs. Chem. Biol. 7, 27–38 (2000).
Saghatelian, A., Jessani, N., Joseph, A., Humphrey, M. & Cravatt, B.F. Activity-based probes for the proteomic profiling of metalloproteases. Proc. Natl. Acad. Sci. USA 101, 10000–10005 (2004).
Chan, E.W., Chattopadhaya, S., Panicker, R.C., Huang, X. & Yao, S.Q. Developing photoactive affinity probes for proteomic profiling: hydroxamate-based probes for metalloproteases. J. Am. Chem. Soc. 126, 14435–14446 (2004).
Greenbaum, D., Medzihradszky, K.F., Burlingame, A. & Bogyo, M. Epoxide electrophiles as activity-dependent cysteine protease profiling and discovery tools. Chem. Biol. 7, 569–581 (2000).
Greenbaum, D. et al. Chemical approaches for functionally probing the proteome. Mol. Cell. Proteomics 1, 60–68 (2002).
Falgueyret, J.P. et al. An activity-based probe for the determination of cysteine cathepsin protease activities in whole cells. Anal. Biochem. 335, 218–227 (2004).
Hemelaar, J. et al. Specific and covalent targeting of conjugating and deconjugating enzymes of ubiquitin-like proteins. Mol. Cell. Biol. 24, 84–95 (2004).
Borodovsky, A. et al. A novel active site-directed probe specific for deubiquitylating enzymes reveals proteasome association of USP14. EMBO J. 20, 5187–5196 (2001).
Borodovsky, A. et al. Small-molecule inhibitors and probes for ubiquitin- and ubiquitin-like-specific proteases. ChemBioChem 6, 287–291 (2005).
Greenbaum, D.C. et al. A role for the protease falcipain 1 in host cell invasion by the human malaria parasite. Science 298, 2002–2006 (2002).
Yasothornsrikul, S. et al. Cathepsin L in secretory vesicles functions as a prohormone-processing enzyme for production of the enkephalin peptide neurotransmitter. Proc. Natl. Acad. Sci. USA 100, 9590–9595 (2003).
Goulet, B. et al. A cathepsin L isoform that is devoid of a signal peptide localizes to the nucleus in S phase and processes the CDP/Cux transcription factor. Mol. Cell 14, 207–219 (2004).
Baruch, A. et al. Defining a link between gap junction communication, proteolysis, and cataract formation. J. Biol. Chem. 276, 28999–29006 (2001).
Mahrus, S. & Craik, C.S. Selective chemical functional probes of granzymes A and B reveal granzyme B is a major effector of natural killer cell-mediated lysis of target cells. Chem. Biol. 12, 567–577 (2005).
Jessani, N., Liu, Y., Humphrey, M. & Cravatt, B.F. Enzyme activity profiles of the secreted and membrane proteome that depict cancer cell invasiveness. Proc. Natl. Acad. Sci. USA 99, 10335–10340 (2002).
Jessani, N. et al. Carcinoma and stromal enzyme activity profiles associated with breast tumor growth in vivo. Proc. Natl. Acad. Sci. USA 101, 13756–13761 (2004).
Joyce, J.A. et al. Cathepsin cysteine proteases are effectors of invasive growth and angiogenesis during multistage tumorigenesis. Cancer Cell 5, 443–453 (2004).
Okerberg, E.S. et al. High-resolution functional proteomics by active-site peptide profiling. Proc. Natl. Acad. Sci. USA 102, 4996–5001 (2005).
Powers, J.C., Asgian, J.L., Ekici, O.D. & James, K.E. Irreversible inhibitors of serine, cysteine, and threonine proteases. Chem. Rev. 102, 4639–4750 (2002).
Sando, S. & Kool, E.T. Quencher as leaving group: efficient detection of DNA-joining reactions. J. Am. Chem. Soc. 124, 2096–2097 (2002).
Sando, S. & Kool, E.T. Imaging of RNA in bacteria with self-ligating quenched probes. J. Am. Chem. Soc. 124, 9686–9687 (2002).
Turk, D., Guncar, G., Podobnik, M. & Turk, B. Revised definition of substrate binding sites of papain-like cysteine proteases. Biol. Chem. 379, 137–147 (1998).
Musil, D. et al. The refined 2.15 A X-ray crystal structure of human liver cathepsin B: the structural basis for its specificity. EMBO J. 10, 2321–2330 (1991).
O'Brien, L.E., Zegers, M.M. & Mostov, K.E. Opinion: Building epithelial architecture: insights from three-dimensional culture models. Nat. Rev. Mol. Cell Biol. 3, 531–537 (2002).
Debnath, J., Muthuswamy, S.K. & Brugge, J.S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Bieth, J.G. Theoretical and practical aspects of proteinase inhibition kinetics. Methods Enzymol. 248, 59–84 (1995).
Barlic-Maganja, D., Dolinar, M. & Turk, V. The influence of Ala205 on the specificity of cathepsin L produced by dextran sulfate assisted activation of the recombinant proenzyme. Biol. Chem. 379, 1449–1452 (1998).
Deval, C., Bechet, D., Obled, A. & Ferrara, M. Purification and properties of different isoforms of bovine cathepsin B. Biochem. Cell Biol. 68, 822–826 (1990).
Moin, K. et al. Human tumour cathepsin B. Comparison with normal liver cathepsin B. Biochem. J. 285, 427–434 (1992).
Acknowledgements
We thank B. Goulet and A. Nepveu (McGill University) for cathepsin L–deficient MEF cells and V. Turk and B. Turk (J. Stefan Institute) for recombinant human cathepsin L. The authors thank C. Gilon for helpful advice on peptide synthesis, G. von Degenfeld for technical assistance throughout the project, K. Boatright and S. Verhelst for helpful discussion of the manuscript. This work was supported by a Turman Fellowship at Stanford University (to M.B.), a US National Institutes of Health National Technology Center for Networks and Pathways grant U54 RR020843 (to M.B.), and a Department of Defense Breast Cancer Center of Excellence grant DAMD-17-02-0693 (to B.F.S.; M.B. subcontract).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Departments of Microbiology and Immunology, Stanford University School of Medicine, 300 Pasteur Dr., Stanford, 94305, California, USA
Supplementary information
Supplementary Fig. 1
Determination of quenching efficiency of the qABP GB117 relative to the unquenched control ABP GB111. (PDF 530 kb)
Supplementary Fig. 2
GB111 and GB117 where docked into cathepsin B (1SP4) and cathepsin L (1MHW) using the MMF94 force field using MOE (Chemical Computing Group). (PDF 3430 kb)
Supplementary Table 1
Inhibition rate constants of various probes for human cathepsin L and bovine cathepsin B. (PDF 35 kb)
Rights and permissions
About this article
Cite this article
Blum, G., Mullins, S., Keren, K. et al. Dynamic imaging of protease activity with fluorescently quenched activity-based probes. Nat Chem Biol 1, 203–209 (2005). https://doi.org/10.1038/nchembio728
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio728
This article is cited by
-
N-Acryloylindole-alkyne (NAIA) enables imaging and profiling new ligandable cysteines and oxidized thiols by chemoproteomics
Nature Communications (2023)
-
AND-gate contrast agents for enhanced fluorescence-guided surgery
Nature Biomedical Engineering (2020)
-
Amphiphilic nanocarrier-induced modulation of PLK1 and miR-34a leads to improved therapeutic response in pancreatic cancer
Nature Communications (2018)
-
Labeling of active proteases in fresh-frozen tissues by topical application of quenched activity-based probes
Nature Protocols (2016)
-
Detecting cathepsin activity in human osteoarthritis via activity-based probes
Arthritis Research & Therapy (2015)