Abstract
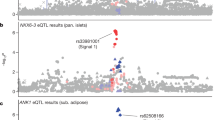
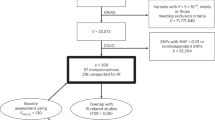
Genome-wide association studies have identified common variants that only partially explain the genetic risk for type 2 diabetes (T2D). Using genome-wide association data from 1,376 French individuals, we identified 16,360 SNPs nominally associated with T2D and studied these SNPs in an independent sample of 4,977 French individuals. We then selected the 28 best hits for replication in 7,698 Danish subjects and identified 4 SNPs showing strong association with T2D, one of which (rs2943641, P = 9.3 × 10−12, OR = 1.19) was located adjacent to the insulin receptor substrate 1 gene (IRS1). Unlike previously reported T2D risk loci, which predominantly associate with impaired beta cell function, the C allele of rs2943641 was associated with insulin resistance and hyperinsulinemia in 14,358 French, Danish and Finnish participants from population-based cohorts; this allele was also associated with reduced basal levels of IRS1 protein and decreased insulin induction of IRS1-associated phosphatidylinositol-3-OH kinase activity in human skeletal muscle biopsies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
13 September 2009
NOTE: In the version of this article initially published online, there were errors in the e-mail addresses of two of the corresponding authors. The correct e-mail address for Robert Sladek is robert.sladek@mcgill.ca; the correct e-mail address for Philippe Froguel is philippe.froguel@good.ibl.fr. These errors have been corrected for the print, PDF and HTML versions of this article.
References
Stumvoll, M., Goldstein, B.J. & van Haeften, T.W. Type 2 diabetes: principles of pathogenesis and therapy. Lancet 365, 1333–1346 (2005).
Bouatia-Naji, N. I A variant near MTNR1B is associated with increased fasting plasma glucose levels and type 2 diabetes risk. Nat. Genet. 41, 89–94 (2009).
Saxena, R. et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316, 1331–1336 (2007).
Scott, L.J. et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316, 1341–1345 (2007).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Steinthorsdottir, V. et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat. Genet. 39, 770–775 (2007).
Unoki, H. et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 40, 1098–1102 (2008).
Yasuda, K. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat. Genet. 40, 1092–1097 (2008).
Zeggini, E. et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 40, 638–645 (2008).
Zeggini, E. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316, 1336–1341 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Sakamoto, H. et al. Genetic variation in PSCA is associated with susceptibility to diffuse-type gastric cancer. Nat. Genet. 40, 730–740 (2008).
Thomas, G. et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 40, 310–315 (2008).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Samani, N.J. et al. Genomewide association analysis of coronary artery disease. N. Engl. J. Med. 357, 443–453 (2007).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn?s disease. Nat. Genet. 40, 955–962 (2008).
Gudmundsson, J. et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat. Genet. 39, 977–983 (2007).
Balkau, B. An epidemiologic survey from a network of French Health Examination Centres, (D.E.S.I.R.): epidemiologic data on the insulin resistance syndrome. Rev. Epidemiol. Sante Publique 44, 373–375 (1996).
Järvelin, M.R. et al. Ecological and individual predictors of birthweight in a northern Finland birth cohort 1986. Paediatr. Perinat. Epidemiol. 11, 298–312 (1997).
Jørgensen, T. et al. A randomized non-pharmacological intervention study for prevention of ischaemic heart disease: baseline results Inter99. Eur. J. Cardiovasc. Prev. Rehabil. 10, 377–386 (2003).
Alberti, K.G. & Zimmet, P.Z. Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet. Med. 15, 539–553 (1998).
Poulsen, P. et al. Impact of genetic versus environmental factors on the control of muscle glycogen synthase activation in twins. Diabetes 54, 1289–1296 (2005).
Aston-Mourney, K., Proietto, J., Morahan, G. & Andrikopoulos, S. Too much of a good thing: why it is bad to stimulate the beta cell to secrete insulin. Diabetologia 51, 540–545 (2008).
Nandi, A., Kitamura, Y., Kahn, C.R. & Accili, D. Mouse models of insulin resistance. Physiol. Rev. 84, 623–647 (2004).
Tamemoto, H. et al. Insulin resistance and growth retardation in mice lacking insulin receptor substrate-1. Nature 372, 182–186 (1994).
Araki, E. et al. Alternative pathway of insulin signalling in mice with targeted disruption of the IRS-1 gene. Nature 372, 186–190 (1994).
Terauchi, Y. et al. Development of non-insulin-dependent diabetes mellitus in the double knockout mice with disruption of insulin receptor substrate-1 and beta cell glucokinase genes. Genetic reconstitution of diabetes as a polygenic disease. J. Clin. Invest. 99, 861–866 (1997).
Withers, D.J. et al. Disruption of IRS-2 causes type 2 diabetes in mice. Nature 391, 900–904 (1998).
Almind, K. et al. Amino acid polymorphisms of insulin receptor substrate-1 in non-insulin-dependent diabetes mellitus. Lancet 342, 828–832 (1993).
Almind, K., Inoue, G., Pedersen, O. & Kahn, C.R. A common amino acid polymorphism in insulin receptor substrate-1 causes impaired insulin signaling. Evidence from transfection studies. J. Clin. Invest. 97, 2569–2575 (1996).
Clausen, J.O. et al. Insulin resistance: interactions between obesity and a common variant of insulin receptor substrate-1. Lancet 346, 397–402 (1995).
Sesti, G. et al. Defects of the insulin receptor substrate (IRS) system in human metabolic disorders. FASEB J. 15, 2099–2111 (2001).
Florez, J.C. et al. Association testing of common variants in the insulin receptor substrate-1 gene (IRS1) with type 2 diabetes. Diabetologia 50, 1209–1217 (2007).
Florez, J.C. et al. Association testing in 9,000 people fails to confirm the association of the insulin receptor substrate-1 G972R polymorphism with type 2 diabetes. Diabetes 53, 3313–3318 (2004).
Jellema, A., Zeegers, M.P., Feskens, E.J., Dagnelie, P.C. & Mensink, R.P. Gly972Arg variant in the insulin receptor substrate-1 gene and association with type 2 diabetes: a meta-analysis of 27 studies. Diabetologia 46, 990–995 (2003).
Zeggini, E. et al. Association studies of insulin receptor substrate 1 gene (IRS1) variants in type 2 diabetes samples enriched for family history and early age of onset. Diabetes 53, 3319–3322 (2004).
Skol, A.D., Scott, L.J., Abecasis, G.R. & Boehnke, M. Optimal designs for two-stage genome-wide association studies. Genet. Epidemiol. 31, 776–788 (2007).
Lauritzen, T. et al. The ADDITION study: proposed trial of the cost-effectiveness of an intensive multifactorial intervention on morbidity and mortality among people with type 2 diabetes detected by screening. Int. J. Obes.Relat. Metab. Disord. 24 (suppl. 3), S6–S11 (2000).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Poulsen, P. et al. Heritability of insulin secretion, peripheral and hepatic insulin action, and intracellular glucose partitioning in young and old Danish twins. Diabetes 54, 275–283 (2005).
Højlund, K. et al. Increased phosphorylation of skeletal muscle glycogen synthase at NH2-terminal sites during physiological hyperinsulinemia in type 2 diabetes. Diabetes 52, 1393–1402 (2003).
Acknowledgements
This work was funded by Genome Canada, Génome Québec and the Canada Foundation for Innovation. This work was also supported by the French Government (Agence Nationale de la Recherche), the French Region of Nord Pas De Calais (Contrat de Projets État-Région), and the following charities: Association Française des Diabétiques, Programme National de Recherche sur le Diabète and Association de Langue Française pour l'Etude du Diabète et des Maladies Métaboliques. The D.E.S.I.R. study has been supported by the Caisse Nationale d?Assurance Maladie des Travailleurs Salariés, Lilly, Novartis Pharma and Sanofi-Aventis, Institut National de la Santé et de la Recherche Médicale (INSERM) (Réseaux en Santé Publique, Interactions entre les déterminants de la santé), Association Diabète Risque Vasculaire, Fédération Française de Cardiologie, Fondation de France, Association de Langue Francaise pour l'Etude du Diabete et des Maladies Metaboliques, Office National Interprofessionnel des Vins, Ardix Medical, Bayer Diagnostics, Becton Dickinson, Cardionics, Merck Santé, Novo Nordisk, Pierre Fabre, Roche and Topcon. The D.E.S.I.R. Study Group: INSERM U780: B. Balkau, P. Ducimetière, E. Eschwège; INSERM U367: F. Alhenc-Gelas; Centre Hospitalier Universitaire D'Angers: Y. Gallois, A. Girault; Bichat Hospital: F. Fumeron, M. Marre; Medical Examination Services: Alençon, Angers, Caen, Chateauroux, Cholet, Le Mans and Tours; Research Institute for General Medicine: J. Cogneau; General practitioners of the region; Cross-Regional Institute for Health: C. Born, E. Caces, M. Cailleau, J.G. Moreau, F. Rakotozafy, J. Tichet, S. Vol. The NFBC 1986 study has been supported by the Oulu University Hospital, Finland, the Academy of Finland and the European Commission (Framework 5 award QLG1-CT-2000-01643). The Danish study was supported by grants from the Lundbeck Foundation Centre of Applied Medical Genomics for Personalized Disease Prediction, Prevention and Care (LUCAMP), the European Union (EUGENE2) grant LSHM-CT-2004-512013, EXGENESIS grant LSHM-CT-2004-005272, the Danish Diabetes Association and the Danish Agency for Science, Technology and Innovation, grant no. 271-06-0539. This study was supported in part by a grant from the European Union (Integrated Project EuroDia LSHM-CT-2006-518153 in the Framework Programme 6 [FP6] of the European Community). M.P. is the recipient of a Canadian Chair in diabetes and metabolism. R.S. is a Chercheur-boursier of the Fonds de la recherche en santé du Québec and the recipient of an MGH One Hundred and Seventy-Fifth Anniversary Award from Research Institute of the Montreal General Hospital Foundation. We thank L. Peltonen-Palotie for her contribution in DNA extraction and distribution; M. Deweider and F. Allegaert for the DNA bank management; R. Frechette, V. Catudal, D. Vincent, A. Forman, I.-L. Wantzin, T. Lorentzen, M. Stendal, M.H. Kristensen, T. Brunt and A.L. Nielsen for their technical or management assistance. Large-scale computations were performed using the Consortium Laval, Université du Québec, McGill and Eastern Quebec supercomputer facility. We are sincerely indebted to all subjects who participated in this study.
Author information
Authors and Affiliations
Contributions
B.B., G.C., P.E., S.H., M.-R.J., J.L., M.M., O.P., T.H., T.L., K.B.-J., C. Pisinger, A.P., A.R., A.S. and J.T. recruited and phenotyped patient cohorts. F.B., A.B., E.D., A. Montpetit and L.S. designed and performed genotyping studies. P.P., R.R.-M., A.V. and J.F.P.W. performed hyperinsulinemic and euglycemic clamp studies as well as studies of muscle biopsies. A.A., S.C., C.C.-P., A. Mazur, G.R., J.R., D.S. and L.S. analyzed genotyping data. A.A., S.C., C.D., P.F., D.M., O.P., C. Polychronakos, B.P., M.P., J.R., R.S. and M.V. wrote the first draft of the manuscript. R.S., P.F. and O.P. planned and coordinated the study. All authors contributed to the final version of the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–4, Supplementary Tables 1–10. (PDF 1405 kb)
Rights and permissions
About this article
Cite this article
Rung, J., Cauchi, S., Albrechtsen, A. et al. Genetic variant near IRS1 is associated with type 2 diabetes, insulin resistance and hyperinsulinemia. Nat Genet 41, 1110–1115 (2009). https://doi.org/10.1038/ng.443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.443
This article is cited by
-
Age effect on the shared etiology of glycemic traits and serum lipids: evidence from a Chinese twin study
Journal of Endocrinological Investigation (2023)
-
H19 inhibition increases HDAC6 and regulates IRS1 levels and insulin signaling in the skeletal muscle during diabetes
Molecular Medicine (2022)
-
Phase separation of insulin receptor substrate 1 drives the formation of insulin/IGF-1 signalosomes
Cell Discovery (2022)
-
Genome-wide DNA methylation profiling in anorexia nervosa discordant identical twins
Translational Psychiatry (2022)
-
Involvement of Cdkal1 in the etiology of type 2 diabetes mellitus and microvascular diabetic complications: a review
Journal of Diabetes & Metabolic Disorders (2022)