Abstract
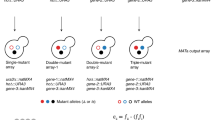
We describe a new synthetic lethality analysis by microarray (SLAM) technique that uses ∼4,600 Saccharomyces cerevisiae haploid deletion mutants with molecular 'bar codes' (TAGs). We used SGS1 and SRS2, two 3′→5′ DNA helicase genes, as 'queries' to identify their redundant and unique biological functions. We introduced these 'query mutations' into a haploid deletion pool by integrative transformation to disrupt the query gene in every cell, generating a double mutant pool. Optimization of integrative transformation efficiency was essential to the success of SLAM. Synthetic interactions defined a DNA helicase genetic network and predicted a role for SRS2 in processing damaged replication forks but, unlike SGS1, not in rDNA replication, DNA topology or lagging strand synthesis. SGS1 and SRS2 have synthetic defects with MRC1 but not RAD9, suggesting that SGS1 and SRS2 function in a parallel pathway with MRC1 to transduce the DNA replication stress signal to the general DNA damage checkpoint pathway. Both helicase genes have rad51-reversible synthetic defects with 5′→3′ DNA helicase RRM3, suggesting that RRM3 helps prevent formation of toxic recombination intermediates. SLAM detects synthetic lethality efficiently and ranks candidate genetic interactions, making it an especially useful method.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Dobzhansky, T. Genetics of natural populations. XIII. Recombination and variability in populations of Drosophila pseudoobscura. Genetics 31, 269–290 (1946).
Novick, P., Osmond, B.C. & Botstein, D. Suppressors of yeast actin mutations. Genetics 121, 659–674 (1989).
Bender, A. & Pringle, J.R. Use of a screen for synthetic lethal and multicopy suppressee mutants to identify two new genes involved in morphogenesis in Saccharomyces cerevisiae. Mol. Cell. Biol. 11, 1295–1305 (1991).
Hartman, J., Garvik, B. & Hartwell, L. Principles for the buffering of genetic variation. Science 291, 1001–1004 (2001).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Ooi, S.L., Shoemaker, D.D. & Boeke, J.D. A DNA microarray-based genetic screen for nonhomologous end-joining mutants in Saccharomyces cerevisiae. Science 294, 2552–2556 (2001).
Rong, L. & Klein, H.L. Purification and characterization of the SRS2 DNA helicase of the yeast Saccharomyces cerevisiae. J. Biol. Chem. 268, 1252–1259 (1993).
Lu, J. et al. Human homologues of yeast helicase. Nature 383, 678–679 (1996).
Bennett, R.J., Sharp, J.A. & Wang, J.C. Purification and characterization of the Sgs1 DNA helicase activity of Saccharomyces cerevisiae. J. Biol. Chem. 273, 9644–9650 (1998).
Hickson, I.D. RecQ helicases: caretakers of the genome. Nat. Rev. Cancer 3, 169–178 (2003).
Aboussekhra, A. et al. RADH, a gene of Saccharomyces cerevisiae encoding a putative DNA helicase involved in DNA repair. Characteristics of radH mutants and sequence of the gene. Nucleic Acids Res. 17, 7211–7219 (1989).
Lee, S.K., Johnson, R.E., Yu, S.L., Prakash, L. & Prakash, S. Requirement of yeast SGS1 and SRS2 genes for replication and transcription. Science 286, 2339–2342 (1999).
Aguilera, A. & Klein, H.L. Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics 119, 779–790 (1988).
Gangloff, S., Soustelle, C. & Fabre, F. Homologous recombination is responsible for cell death in the absence of the Sgs1 and Srs2 helicases. Nat. Genet. 25, 192–194 (2000).
Yamagata, K. et al. Bloom's and Werner's syndrome genes suppress hyperrecombination in yeast sgs1 mutant: implication for genomic instability in human diseases. Proc. Natl. Acad. Sci. USA 95, 8733–8738 (1998).
Aylon, Y., Liefshitz, B., Bitan-Banin, G. & Kupiec, M. Molecular dissection of mitotic recombination in the yeast Saccharomyces cerevisiae. Mol. Cell. Biol. 23, 1403–1417 (2003).
Vaze, M.B. et al. Recovery from checkpoint-mediated arrest after repair of a double-strand break requires Srs2 helicase. Mol. Cell 10, 373–385 (2002).
Frei, C. & Gasser, S.M. The yeast Sgs1p helicase acts upstream of Rad53p in the DNA replication checkpoint and colocalizes with Rad53p in S-phase-specific foci. Genes Dev. 14, 81–96 (2000).
Bennett, C.B. et al. Genes required for ionizing radiation resistance in yeast. Nat. Genet. 29, 426–434 (2001).
Mankouri, H.W., Craig, T.J. & Morgan, A. SGS1 is a multicopy suppressor of srs2: functional overlap between DNA helicases. Nucleic Acids Res. 30, 1103–1113 (2002).
Mullen, J.R., Kaliraman, V., Ibrahim, S.S. & Brill, S.J. Requirement for three novel protein complexes in the absence of the Sgs1 DNA helicase in Saccharomyces cerevisiae. Genetics 157, 103–118 (2001).
Debrauwere, H., Loeillet, S., Lin, W., Lopes, J. & Nicolas, A. Links between replication and recombination in Saccharomyces cerevisiae: a hypersensitive requirement for homologous recombination in the absence of Rad27 activity. Proc. Natl. Acad. Sci. USA 98, 8263–8269 (2001).
Ogawa, T. et al. RecA-like recombination proteins in eukaryotes: functions and structures of RAD51 genes. Cold Spring Harb. Symp. Quant. Biol. 58, 567–576 (1993).
Sung, P., Trujillo, K.M. & Van Komen, S. Recombination factors of Saccharomyces cerevisiae. Mutat. Res. 451, 257–275 (2000).
Fabre, F., Chan, A., Heyer, W.D. & Gangloff, S. Alternate pathways involving Sgs1/Top3, Mus81/ Mms4, and Srs2 prevent formation of toxic recombination intermediates from single-stranded gaps created by DNA replication. Proc. Natl. Acad. Sci. USA 99, 16887–16892 (2002).
Morrow, D.M., Connelly, C. & Hieter, P. “Break copy” duplication: a model for chromosome fragment formation in Saccharomyces cerevisiae. Genetics 147, 371–382 (1997).
Alcasabas, A.A. et al. Mrc1 transduces signals of DNA replication stress to activate Rad53. Nat. Cell Biol. 3, 958–965 (2001).
Qiu, J., Qian, Y., Frank, P., Wintersberger, U. & Shen, B. Saccharomyces cerevisiae RNase H(35) functions in RNA primer removal during lagging-strand DNA synthesis, most efficiently in cooperation with Rad27 nuclease. Mol. Cell. Biol. 19, 8361–8371 (1999).
Uetz, P. et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Schmidt, K.H., Derry, K.L. & Kolodner, R.D. Saccharomyces cerevisiae RRM3, a 5′ to 3′ DNA helicase, physically interacts with proliferating cell nuclear antigen. J. Biol. Chem. 277, 45331–45337 (2002).
Ivessa, A.S., Zhou, J.Q. & Zakian, V.A. The Saccharomyces Pif1p DNA helicase and the highly related Rrm3p have opposite effects on replication fork progression in ribosomal DNA. Cell 100, 479–489 (2000).
Ivessa, A.S., Zhou, J.Q., Schulz, V.P., Monson, E.K. & Zakian, V.A. Saccharomyces Rrm3p, a 5′ to 3′ DNA helicase that promotes replication fork progression through telomeric and subtelomeric DNA. Genes Dev. 16, 1383–1396 (2002).
Hughes, T.R. et al. Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat. Biotechnol. 19, 342–347 (2001).
Breitkreutz, B.J., Stark, C. & Tyers, M. Osprey: a network visualization system. Genome Biol. 4, R22 (2003).
Acknowledgements
We thank R. Crouch and H. Jeong for sharing unpublished results; M. Lee and S. Sookhai-Mahadeo for optimizing the long range PCR; S. Sookhai-Mahadeo, A. IJpma and M. Lee for help with deletion pool construction; C. Boone and A. Tong for strains and discussion; G. Ira and R. Rothstein for discussions and reading the manuscripts; R. Irizarry and J. Bader for discussions on statistical aspects and network analysis; and E. Bolton, M. Lee, X. Pan, F. Spencer and D. Yuan for discussions. This work was supported by a grant from the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Ooi, S., Shoemaker, D. & Boeke, J. DNA helicase gene interaction network defined using synthetic lethality analyzed by microarray. Nat Genet 35, 277–286 (2003). https://doi.org/10.1038/ng1258
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1258
This article is cited by
-
Transposon insertion sequencing: a new tool for systems-level analysis of microorganisms
Nature Reviews Microbiology (2013)
-
An N-terminal acidic region of Sgs1 interacts with Rpa70 and recruits Rad53 kinase to stalled forks
The EMBO Journal (2012)
-
Extracting quantitative genetic interaction phenotypes from matrix combinatorial RNAi
BMC Bioinformatics (2011)
-
Chemical-genetic profile analysis of five inhibitory compounds in yeast
BMC Chemical Biology (2010)
-
Improving the iMM904 S. cerevisiae metabolic model using essentiality and synthetic lethality data
BMC Systems Biology (2010)