Abstract
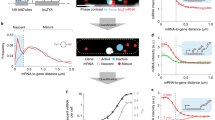
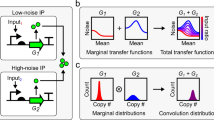
The presence of low-copy-number regulators and switch-like signal propagation in regulatory networks are expected to increase noise in cellular processes. We developed a noise amplifier that detects fluctuations in the level of low-abundance mRNAs in yeast. The observed fluctuations are not due to the low number of molecules expressed from a gene per se but originate in the random, rare events of gene activation. The frequency of these events and the correlation between stochastic expressions of genes in a single cell depend on the positioning of the genes along the chromosomes. Transcriptional regulators produced by such random expression propagate noise to their target genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Rao, C.V., Wolf, D.M. & Arkin, A.P. Control, exploitation and tolerance of intracellular noise. Nature 420, 231–237 (2002).
Becskei, A., Seraphin, B. & Serrano, L. Positive feedback in eukaryotic gene networks: cell differentiation by graded to binary response conversion. EMBO J. 20, 2528–2535 (2001).
Thattai, M. & van Oudenaarden, A. Stochastic gene expression in fluctuating environments. Genetics 167, 523–530 (2004).
Erdi, P. & Toth, J. Mathematical Models of Chemical Reactions: Theory and Applications of Deterministic and Stochastic Models (Manchester University Press, Manchester, UK, 1989).
Horsthemke, W., Doering, C.R., Ray, T.S. & Burschka, M.A. Fluctuations and correlations in a diffusion-reaction system - unified description of internal fluctuations and external noise. Phys. Rev. A. 45, 5492–5503 (1992).
Elowitz, M.B., Levine, A.J., Siggia, E.D. & Swain, P.S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004).
Pedraza, J.M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005).
Rosenfeld, N., Young, J.W., Alon, U., Swain, P.S. & Elowitz, M.B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Holland, M.J. Transcript abundance in yeast varies over six orders of magnitude. J. Biol. Chem. 277, 14363–14366 (2002).
Velculescu, V.E. et al. Characterization of the yeast transcriptome. Cell 88, 243–251 (1997).
Swain, P.S. Efficient attenuation of stochasticity in gene expression through post-transcriptional control. J. Mol. Biol. 344, 965–976 (2004).
Becskei, A., Boselli, M.G. & van Oudenaarden, A. Amplitude control of cell-cycle waves by nuclear import. Nat. Cell Biol. 6, 451–457 (2004).
Hasty, J., Dolnik, M., Rottschafer, V. & Collins, J.J. Synthetic gene network for entraining and amplifying cellular oscillations. Phys. Rev. Lett. 88, 148101 (2002).
Isaacs, F.J., Hasty, J., Cantor, C.R. & Collins, J.J. Prediction and measurement of an autoregulatory genetic module. Proc. Natl. Acad. Sci. USA 100, 7714–7719 (2003).
Basu, S., Mehreja, R., Thiberge, S., Chen, M.T. & Weiss, R. Spatiotemporal control of gene expression with pulse-generating networks. Proc. Natl. Acad. Sci. USA 101, 6355–6360 (2004).
Xiong, W. & Ferrell, J.E., Jr. A positive-feedback-based bistable 'memory module' that governs a cell fate decision. Nature 426, 460–465 (2003).
Pramila, T., Miles, S., GuhaThakurta, D., Jemiolo, D. & Breeden, L.L. Conserved homeodomain proteins interact with MADS box protein Mcm1 to restrict ECB-dependent transcription to the M/G1 phase of the cell cycle. Genes Dev. 16, 3034–3045 (2002).
Acar, M., Becskei, A. & van Oudenaarden, A. Enhancement of cellular memory by reducing stochastic transitions. Nature 435, 228–232 (2005).
Shibata, T. & Fujimoto, K. Noisy signal amplification in ultrasensitive signal transduction. Proc. Natl. Acad. Sci. USA 102, 331–336 (2005).
Hooshangi, S., Thiberge, S. & Weiss, R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl. Acad. Sci. USA 102, 3581–3586 (2005).
Blake, W.J., Kaern, M., Cantor, C.R. & Collins, J.J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003).
Raser, J.M. & O'Shea, E.K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004).
Li, G. & Widom, J. Nucleosomes facilitate their own invasion. Nat. Struct. Mol. Biol. 11, 763–769 (2004).
Vashee, S., Melcher, K., Ding, W.V., Johnston, S.A. & Kodadek, T. Evidence for two modes of cooperative DNA binding in vivo that do not involve direct protein-protein interactions. Curr. Biol. 8, 452–458 (1998).
Melcher, K. & Xu, H.E. Gal80-Gal80 interaction on adjacent Gal4p binding sites is required for complete GAL gene repression. EMBO J. 20, 841–851 (2001).
Harbison, C.T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004).
Basehoar, A.D., Zanton, S.J. & Pugh, B.F. Identification and distinct regulation of yeast TATA box-containing genes. Cell 116, 699–709 (2004).
Maillet, L. et al. Evidence for silencing compartments within the yeast nucleus: a role for telomere proximity and Sir protein concentration in silencer-mediated repression. Genes Dev. 10, 1796–1811 (1996).
Carr, A.J. & Whitmore, D. Imaging of single light-responsive clock cells reveals fluctuating free-running periods. Nat. Cell Biol. 7, 319–321 (2005).
Lengronne, A. & Schwob, E. The yeast CDK inhibitor Sic1 prevents genomic instability by promoting replication origin licensing in late G(1). Mol. Cell 9, 1067–1078 (2002).
Thornton, B.R., Chen, K.C., Cross, F.R., Tyson, J.J. & Toczyski, D.P. Cycling without the cyclosome: modeling a yeast strain lacking the APC. Cell Cycle 3, 629–633 (2004).
Magee, J.A., Abdulkadir, S.A. & Milbrandt, J. Haploinsufficiency at the Nkx3.1 locus. A paradigm for stochastic, dosage-sensitive gene regulation during tumor initiation. Cancer Cell 3, 273–283 (2003).
Sveiczer, A., Csikasz-Nagy, A., Gyorffy, B., Tyson, J.J. & Novak, B. Modeling the fission yeast cell cycle: quantized cycle times in wee1–cdc25Delta mutant cells. Proc. Natl. Acad. Sci. USA 97, 7865–7870 (2000).
Pirone, J.R. & Elston, T.C. Fluctuations in transcription factor binding can explain the graded and binary responses observed in inducible gene expression. J. Theor. Biol. 226, 111–121 (2004).
Menon, B.B. et al. Reverse recruitment: The Nup84 nuclear pore subcomplex mediates Rap1/Gcr1/Gcr2 transcriptional activation. Proc. Natl. Acad. Sci. USA 102, 5749–5754 (2005).
Casolari, J.M. et al. Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117, 427–439 (2004).
Hurst, L.D., Pal, C. & Lercher, M.J. The evolutionary dynamics of eukaryotic gene order. Nat. Rev. Genet. 5, 299–310 (2004).
Osborne, C.S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Struhl, K. Fundamentally different logic of gene regulation in eukaryotes and prokaryotes. Cell 98, 1–4 (1999).
Martin, D.I. Transcriptional enhancers–on/off gene regulation as an adaptation to silencing in higher eukaryotic nuclei. Trends Genet. 17, 444–448 (2001).
Ahmad, K. & Henikoff, S. Modulation of a transcription factor counteracts heterochromatic gene silencing in Drosophila. Cell 104, 839–847 (2001).
Misteli, T. Concepts in nuclear architecture. Bioessays 27, 477–487 (2005).
Roix, J.J., McQueen, P.G., Munson, P.J., Parada, L.A. & Misteli, T. Spatial proximity of translocation-prone gene loci in human lymphomas. Nat. Genet. 34, 287–291 (2003).
Dernburg, A.F. et al. Perturbation of nuclear architecture by long-distance chromosome interactions. Cell 85, 745–759 (1996).
Nutt, S.L. et al. Independent regulation of the two Pax5 alleles during B-cell development. Nat. Genet. 21, 390–395 (1999).
Peccoud, J. & Ycard, B. Markovian modelling of gene product synthesis. Theor. Popul. Biol. 48, 222–234 (1995).
Iyer, V. & Struhl, K. Absolute mRNA levels and transcriptional initiation rates in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 93, 5208–5212 (1996).
Guptasarma, P. Does replication-induced transcription regulate synthesis of the myriad low copy number proteins of Escherichia coli?. Bioessays 17, 987–997 (1995).
Acknowledgements
We thank J. Pedraza, W. Tansey and M. Thattai for discussions. A.B. is a Long Term Fellow of the Human Frontier Science Program. This work was supported by a grant from the US National Institutes of Health and a US National Science Foundation CAREER grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Moments of distribution of YFP fluorescence. (PDF 91 kb)
Supplementary Fig. 2
Fitting of the input noise. (PDF 134 kb)
Supplementary Table 1
Yeast strains. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Becskei, A., Kaufmann, B. & van Oudenaarden, A. Contributions of low molecule number and chromosomal positioning to stochastic gene expression. Nat Genet 37, 937–944 (2005). https://doi.org/10.1038/ng1616
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1616
This article is cited by
-
Heritable transcriptional defects from aberrations of nuclear architecture
Nature (2023)
-
Pairing of segmentation clock genes drives robust pattern formation
Nature (2021)
-
Independent control of mean and noise by convolution of gene expression distributions
Nature Communications (2021)
-
Inter-embryo gene expression variability recapitulates the hourglass pattern of evo-devo
BMC Biology (2020)
-
Insights about collective decision-making at the genetic level
Biophysical Reviews (2020)