Abstract
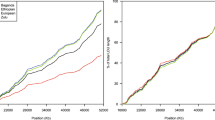
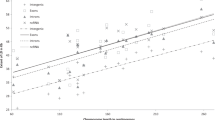
The genome-wide distribution of linkage disequilibrium (LD) determines the strategy for selecting markers for association studies, but it varies between populations. We assayed LD in large samples (200 individuals) from each of 11 well-described population isolates and an outbred European-derived sample, using SNP markers spaced across chromosome 22. Most isolates show substantially higher levels of LD than the outbred sample and many fewer regions of very low LD (termed 'holes'). Young isolates known to have had relatively few founders show particularly extensive LD with very few holes; these populations offer substantial advantages for genome-wide association mapping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Thomas, D.C., Haile, R.W. & Duggan, D. Recent developments in genomewide association scans: A workshop summary and review. Am. J. Hum. Genet. 77, 337–345 (2005).
Stumpf, M.P.H. & Goldstein, D.B. Demography, recombination hotspot intensity, and the block structure of linkage disequilibrium. Curr. Biol. 13, 1–8 (2003).
Peltonen, L., Palotie, A. & Lange, K. Use of population isolates for mapping complex traits. Nat. Rev. Genet. 1, 182–190 (2000).
Wright, A.F., Carothers, A.D. & Piratsu, M. Population choice in mapping genes for complex disease. Nat. Genet. 23, 397–404 (1999).
Pritchard, J.K. & Przeworski, M. Linkage disequilibrium in humans: models and data. Am. J. Hum. Genet. 69, 1–14 (2001).
Teare, M.D., Dunning, A.M., Durocher, F., Rennart, G. & Easton, D.F. Sampling distribution of summary linkage disequilibrium measures. Ann. Hum. Genet. 66, 223–233 (2002).
Tenesa, A. et al. Extent of linkage disequilbirum in a Sardinian sub-isolate: sampling and methodological considerations. Hum. Mol. Genet. 13, 25–33 (2004).
Groenewald, J.Z., Liebenberg, J., Groenewald, I.M. & Warnich, L. Linkage disequilibrium analysis in a recently founded population: evaluation of the variegate porphyria founder in South African Afrikaners. Am. J. Hum. Genet. 62, 1254–1258 (1998).
Gordon, D., Simonic, I. & Ott, J. Significant evidence for linkage disequilibrium over a 5-cM region among Afrikaners. Genomics 66, 87–92 (2000).
Service, S.K., Ophoff, R. & Freimer, N.B. The genomewide distribution of background linkage disequilbrium in a population isolate. Hum. Mol. Genet. 10, 545–551 (2001).
Hall, D., Wijsman, E.M., Roos, J.L., Gogos, J.A. & Karayiorgou, M. Extended intermarker linkage disequilibrium in the Afrikaners. Genome Res. 12, 956–961 (2002).
Varilo, T. et al. The interval of linkage disequilibrium (LD) detected with microsatellite and SNP markers in chromosomes of Finnish populations with different histories. Hum. Mol. Genet. 12, 51–59 (2003).
Aulchenko, Y.S. et al. Linkage disequilibrium in young genetically isolated Dutch population. Eur. J. Hum. Genet. 12, 527–534 (2004).
Maraganore, D.M. et al. High-resolution whole-genome association study of Parkinson disease. Am. J. Hum. Genet. 77, 685–693 (2005).
Nei, M. Genetic distance between populations. Am. Nat. 106, 283–292 (1972).
Dawson, E. et al. A first-generation linkage disequilibrium map of human chromosome 22. Nature 418, 544–548 (2002).
Maniatis, N. et al. The first linkage disequilibrium (LD) maps: Delineation of hot and cold blocks by diplotype analysis. Proc. Natl. Acad. Sci. USA 99, 2228–2233 (2002).
Ke, X. et al. The impact of SNP density on fine-scale patterns of linkage disequilibrium. Hum. Mol. Genet. 13, 577–588 (2004).
Tapper, W.J., Maniatis, N., Morton, N.E. & Collins, A. A metric linkage disequilibrium map of a human chromosome. Ann. Hum. Genet. 67, 487–494 (2003).
De La Vega, F.M. et al. The linkage disequilibrium maps of three human chromosomes across four populations reflect their demographic history and a common underlying recombination pattern. Genome Res. 15, 454–462 (2005).
Kong, X. et al. A combined linkage-physical map of the human genome. Am. J. Hum. Genet. 75, 1143–1148 (2004).
Maniatis, N. et al. Positional cloning by linkage disequilibrium. Am. J. Hum. Genet. 74, 846–855 (2004).
Carvajal-Carmona, L.G. et al. Genetic demography of Antioquia (Colombia) and the central valley of Costa Rica. Hum. Genet. 112, 534–541 (2003).
The International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
Fan, J.-B. et al. Highly parallel SNP genotyping. Cold Spring Harb. Symp. Quant. Biol. 68, 69–78 (2004).
Abecasis, G.R. & Cookson, W.O. GOLD–graphical overview of linkage disequilibrium. Bioinformatics 16, 182–183 (2000).
Collins, A. & Morton, N.E. Mapping a disease locus by allelic association. Proc. Natl. Acad. Sci. USA 95, 1741–1745 (1998).
Morton, N.E. et al. The optimal measure of allelic association. Proc. Natl. Acad. Sci. USA 98, 5217–5221 (2001).
Efron, B. & Tibshirani, R.J. An Introduction to the Bootstrap (Chapman and Hall, New York, 1993).
McVean, G.A. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584 (2004).
Acknowledgements
We thank all of the study participants; the Center of Medical Systems Biology (CMSB); D. Ruano (Life and Health Sciences Research Institute (ICVS), University of Minho, Braga, Portugal); M.J. Soares, J. Valente and M.H. Azevedo (Instituto de Psicologia Médica, Faculdade de Medicina, Coimbra, Portugal); C. Pato and M.T. Pato (Center for Psychiatric and Molecular Genetics, and Department of Psychiatry, State University of New York, Syracuse, New York, and the Veterans Administration Medical Center, Washington, D.C.); A. Gabbas (Haematology Division and Bone Marrow Transplantation Unit, San Francesco Hospital, Nuoro, Italy); R. van Wyk, C. Botha and G. Valencia (Universidad de Antioquia) and P. Snijders for recruitment of subjects; M. Almonte and E. Slaten for genotyping assistance; R. Ophoff for comments on the manuscript and M. Levinson for assistance with graphics. We acknowledge funding from the Biotechnology and Biological Sciences Research Council (A.C.), the US National Institutes of Health (N.F., M.K., A.R.-L.), Newfound Genomics (P.R.), Colciencias (A.R.-L.), the Universidad de Antioquia (A.R.-L.), the National Alliance for Research in Schizophrenia and Depression (A.R.-L.), the Wellcome Trust (A.R.-L.), the Netherlands Organisation for Scientific Research (C. van D.), the Netherlands Diabetes Fund (C. van D.), the Netherlands Kidney Fund (C. van D.), the Netherlands Heart Foundation (C. van D.), the International Alzheimer Organisation (C. van D.), the Netherlands Brain Fund (C. van D.), the Center of Excellence of the Academy of Finland (L.P.), Nordic Center of Excellence in Disease Genetics (L.P.) and Biocentrum Helsinki, Finland (L.P.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
S.M. and L.G. are employed by Illumina, Inc.
Supplementary information
Supplementary Table 1
Demographic history of population isolates. (PDF 20 kb)
Rights and permissions
About this article
Cite this article
Service, S., DeYoung, J., Karayiorgou, M. et al. Magnitude and distribution of linkage disequilibrium in population isolates and implications for genome-wide association studies. Nat Genet 38, 556–560 (2006). https://doi.org/10.1038/ng1770
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1770
This article is cited by
-
Identification of eQTLs using different sets of single nucleotide polymorphisms associated with carcass and body composition traits in pigs
BMC Genomics (2024)
-
The Newfoundland and Labrador mosaic founder population descends from an Irish and British diaspora from 300 years ago
Communications Biology (2023)
-
Genetic history of Calabrian Greeks reveals ancient events and long term isolation in the Aspromonte area of Southern Italy
Scientific Reports (2021)
-
Linkage disequilibrium maps for European and African populations constructed from whole genome sequence data
Scientific Data (2019)
-
Heterogeneity in the extent of linkage disequilibrium among exonic, intronic, non-coding RNA and intergenic chromosome regions
European Journal of Human Genetics (2019)