Abstract
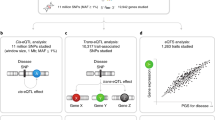
Quantitative differences in gene expression are thought to contribute to phenotypic differences between individuals. We generated genome-wide transcriptional profiles of lymphocyte samples from 1,240 participants in the San Antonio Family Heart Study. The expression levels of 85% of the 19,648 detected autosomal transcripts were significantly heritable. Linkage analysis uncovered >1,000 cis-regulated transcripts at a false discovery rate of 5% and showed that the expression quantitative trait loci with the most significant linkage evidence are often located at the structural locus of a given transcript. To highlight the usefulness of this much-enlarged map of cis-regulated transcripts for the discovery of genes that influence complex traits in humans, as an example we selected high-density lipoprotein cholesterol concentration as a phenotype of clinical importance, and identified the cis-regulated vanin 1 (VNN1) gene as harboring sequence variants that influence high-density lipoprotein cholesterol concentrations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Jansen, R.C. & Nap, J.P. Genetical genomics: the added value from segregation. Trends Genet. 17, 388–391 (2001).
Brem, R.B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002).
Yvert, G. et al. Trans-acting regulatory variation in Saccharomyces cerevisiae and the role of transcription factors. Nat. Genet. 35, 57–64 (2003).
Storey, J.D., Akey, J.M. & Kruglyak, L. Multiple locus linkage analysis of genomewide expression in yeast. PLoS Biol. 3, e267 (2005).
Brem, R.B. & Kruglyak, L. The landscape of genetic complexity across 5,700 gene expression traits in yeast. Proc. Natl. Acad. Sci. USA 102, 1572–1577 (2005).
Schadt, E.E. et al. Genetics of gene expression surveyed in maize, mouse and man. Nature 422, 297–302 (2003).
Kirst, M. et al. Coordinated genetic regulation of growth and lignin revealed by quantitative trait locus analysis of cDNA microarray data in an interspecific backcross of eucalyptus. Plant Physiol. 135, 2368–2378 (2004).
Wayne, M.L. & McIntyre, L.M. Combining mapping and arraying: An approach to candidate gene identification. Proc. Natl. Acad. Sci. USA 99, 14903–14906 (2002).
Bystrykh, L. et al. Uncovering regulatory pathways that affect hematopoietic stem cell function using 'genetical genomics'. Nat. Genet. 37, 225–232 (2005).
Chesler, E.J. et al. Complex trait analysis of gene expression uncovers polygenic and pleiotropic networks that modulate nervous system function. Nat. Genet. 37, 233–242 (2005).
Hubner, N. et al. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nat. Genet. 37, 243–253 (2005).
Dausset, J. et al. Centre d'etude du polymorphisme humain (CEPH): collaborative genetic mapping of the human genome. Genomics 6, 575–577 (1990).
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004).
Monks, S.A. et al. Genetic inheritance of gene expression in human cell lines. Am. J. Hum. Genet. 75, 1094–1105 (2004).
Cheung, V.G. et al. Mapping determinants of human gene expression by regional and genome-wide association. Nature 437, 1365–1369 (2005).
Stranger, B.E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005).
Deutsch, S. et al. Gene expression variation and expression quantitative trait mapping of human chromosome 21 genes. Hum. Mol. Genet. 14, 3741–3749 (2005).
Pastinen, T., Ge, B. & Hudson, T.J. Influence of human genome polymorphism on gene expression. Hum. Mol. Genet. 15, R9–R16 (2006).
de Koning, D.J. & Haley, C.S. Genetical genomics in humans and model organisms. Trends Genet. 21, 377–381 (2005).
Pant, P.V. et al. Analysis of allelic differential expression in human white blood cells. Genome Res. 16, 331–339 (2006).
Whitney, A.R. et al. Individuality and variation in gene expression patterns in human blood. Proc. Natl. Acad. Sci. USA 100, 1896–1901 (2003).
Stranger, B.E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Storey, J.D. et al. Gene-expression variation within and among human populations. Am. J. Hum. Genet. 80, 502–509 (2007).
Mitchell, B.D. et al. Genetic and environmental contributions to cardiovascular risk factors in Mexican Americans. The San Antonio Family Heart Study. Circulation 94, 2159–2170 (1996).
Pruitt, K.D., Tatusova, T. & Maglott, D.R. NCBI Reference Sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 33, D501–D504 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and poerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Jin, W. et al. The contributions of sex, genotype and age to transcriptional variance in Drosophila melanogaster. Nat. Genet. 29, 389–395 (2001).
Heath, S.C. Markov chain Monte Carlo segregation and linkage analysis for oligogenic models. Am. J. Hum. Genet. 61, 748–760 (1997).
Almasy, L. & Blangero, J. Multipoint quantitative-trait linkage analysis in general pedigrees. Am. J. Hum. Genet. 62, 1198–1211 (1998).
Kong, A. et al. A high-resolution recombination map of the human genome. Nat. Genet. 31, 241–247 (2002).
Göring, H.H., Terwilliger, J.D. & Blangero, J. Large upward bias in estimation of locus-specific effects from genomewide scans. Am. J. Hum. Genet. 69, 1357–1369 (2001).
Williams, J.T. & Blangero, J. Power of variance component linkage analysis to detect quantitative trait loci. Ann. Hum. Genet. 63, 545–563 (1999).
Young, C.E., Karas, R.H. & Kuvin, J.T. High-density lipoprotein cholesterol and coronary heart disease. Cardiol. Rev. 12, 107–119 (2004).
Yamazaki, K., Kuromitsu, J. & Tanaka, I. Microarray analysis of gene expression changes in mouse liver induced by peroxisome proliferator- activated receptor alpha agonists. Biochem. Biophys. Res. Commun. 290, 1114–1122 (2002).
Blangero, J. et al. Quantitative trait nucleotide analysis using Bayesian model selection. Hum. Biol. 77, 541–559 (2005).
Kent, J.W. Jr., Dyer, T.D., Göring, H.H.H. & Blangero, J. Type I error rates in association versus joint linkage/association tests in related individuals. Genet. Epidemiol. 31, 173–177 (2007).
Chekmenev, D.S., Haid, C. & Kel, A.E. P-Match: transcription factor binding site search by combining patterns and weight matrices. Nucleic Acids Res. 33, W432–W437 (2005).
Wingender, E. et al. TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res. 28, 316–319 (2000).
Kadonaga, J.T., Carner, K.R., Masiarz, F.R. & Tjian, R. Isolation of cDNA encoding transcription factor Sp1 and functional analysis of the DNA binding domain. Cell 51, 1079–1090 (1987).
Turner, J. & Crossley, M. Mammalian Kruppel-like transcription factors: more than just a pretty finger. Trends Biochem. Sci. 24, 236–240 (1999).
Kaczynski, J., Cook, T. & Urrutia, R. Sp1- and Kruppel-like transcription factors. Genome Biol. 4, 206 (2003).
Perez-Enciso, M. In silico study of transcriptome genetic variation in outbred populations. Genetics 166, 547–554 (2004).
Schadt, E.E. et al. An integrative genomics approach to infer causal associations between gene expression and disease. Nat. Genet. 37, 710–717 (2005).
McPeek, M.S. & Sun, L. Statistical tests for detection of misspecified relationships by use of genome-screen data. Am. J. Hum. Genet. 66, 1076–1094 (2000).
Sobel, E. & Lange, K. Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am. J. Hum. Genet. 58, 1323–1337 (1996).
Sobel, E., Papp, J.C. & Lange, K. Detection and integration of genotyping errors in statistical genetics. Am. J. Hum. Genet. 70, 496–508 (2002).
Boehnke, M. Allele frequency estimation from data on relatives. Am. J. Hum. Genet. 48, 22–25 (1991).
Boerwinkle, E., Chakraborty, R. & Sing, C.F. The use of measured genotype information in the analysis of quantitative phenotypes in man. I. Models and analytical methods. Ann. Hum. Genet. 50, 181–194 (1986).
Warnick, G.R., Benderson, J. & Albers, J.J. Dextran sulfate-Mg2+ precipitation procedure for quantitation of high-density-lipoprotein cholesterol. Clin. Chem. 28, 1379–1388 (1982).
Li, Y.C., Ross, J., Scheppler, J.A. & Franza, B.R. Jr. An in vitro transcription analysis of early responses of the human immunodeficiency virus type 1 long terminal repeat to different transcriptional activators. Mol. Cell. Biol. 11, 1883–1893 (1991).
Acknowledgements
We are grateful to the participants in the San Antonio Family Heart Study. Data collection was supported by a grant from the US National Institute for Heart, Lungs and Blood (HL045222). A donation from the Azar and Shepperd families paid for the transcriptional profiling. Additional funds for transcriptional profiling, sequencing, genotyping and statistical analysis were provided by ChemGenex Pharmaceuticals. The SOLAR statistical genetics computer package is supported by a grant from the US National Institute of Mental Health (MH059490). The supercomputing facilities used for this work at the AT&T Genetics Computing Center were supported in part by a gift from the SBC Foundation. The laboratory work was carried out in facilities that were constructed with support from the US National Center for Research Resources (RR013556).
Author information
Authors and Affiliations
Contributions
J.B., E.K.M. and G.R.C. initiated the study. H.H.H.G. and J.B. performed or supervised all aspects of the statistical analysis and were aided by T.D.D. and J.C. E.K.M., J.E.C. and L.J.A. were responsible for all molecular analyses, including transcriptional profiles, resequencing, SNP typing and functional analysis, with aid from M.P.J. J.B., J.W.M., M.C.M., A.G.C., and L.A. were responsible for the Mexican American family samples. S.A.C. and J.B. were responsible for the 10-cM STR typing. D.L.R. was responsible for the HDL-C measurement. J.B.M.J. and J.C. performed bioinformatic analyses. A.H.K. and G.R.C. provided additional biological interpretation.
Corresponding author
Ethics declarations
Competing interests
Funds for resequencing, genotyping, transcriptional profiling, and statistical analyses were provided in part by ChemGenex Pharmaceuticals. G.R.C. has personal financial interests in ChemGenex. G.R.C. is also the Chief Executive Officer and Managing Director of ChemGenex. J.B. is a member of the Scientific Advisory Board of ChemGenex. J.B. also serves as Senior Director of Human Genomics.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3,5 and Supplementary Figure 1 (PDF 290 kb)
Supplementary Table 4
Heritability estimates, cis lod scores and maximum trans lod scores for all transcripts (XLS 1684 kb)
Rights and permissions
About this article
Cite this article
Göring, H., Curran, J., Johnson, M. et al. Discovery of expression QTLs using large-scale transcriptional profiling in human lymphocytes. Nat Genet 39, 1208–1216 (2007). https://doi.org/10.1038/ng2119
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2119
This article is cited by
-
Tissue-specific enhancer–gene maps from multimodal single-cell data identify causal disease alleles
Nature Genetics (2024)
-
Robust identification of regulatory variants (eQTLs) using a differential expression framework developed for RNA-sequencing
Journal of Animal Science and Biotechnology (2023)
-
Expression quantitative trait loci in sheep liver and muscle contribute to variations in meat traits
Genetics Selection Evolution (2021)
-
A 16q22.1 variant confers susceptibility to colorectal cancer as a distal regulator of ZFP90
Oncogene (2020)
-
OSCA: a tool for omic-data-based complex trait analysis
Genome Biology (2019)