Abstract
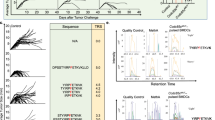
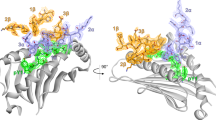
Structural studies of complexes of T cell receptor (TCR) and peptide–major histocompatibility complex (MHC) have focused on TCRs specific for foreign antigens or native self. An unexplored category of TCRs includes those specific for self determinants bearing alterations resulting from disease, notably cancer. We determined here the structure of a human melanoma–specific TCR (E8) bound to the MHC molecule HLA-DR1 and an epitope from mutant triosephosphate isomerase. The structure had features intermediate between 'anti-foreign' and autoimmune TCR–peptide–MHC class II complexes that may reflect the hybrid nature of altered self. E8 manifested very low affinity for mutant triosephosphate isomerase–HLA-DR1 despite the highly tumor-reactive properties of E8 cells. A second TCR (G4) had even lower affinity but underwent peptide-specific formation of dimers, suggesting this as a mechanism for enhancing low-affinity TCR-peptide-MHC interactions for T cell activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Rosenberg, S.A. Progress in human tumour immunology and immunotherapy. Nature 411, 380–384 (2001).
Houghton, A.N. & Guevara-Patino, J.A. Immune recognition of self in immunity against cancer. J. Clin. Invest. 114, 468–471 (2004).
Wucherpfennig, K.W. & Strominger, J.L. Molecular mimicry in T-cell mediated autoimmunity: viral peptides activate human T cell clones specific for myelin basic protein. Cell 80, 695–705 (1995).
Engelhorn, M.E. et al. Autoimmunity and tumor immunity induced by immune responses to mutations in self. Nat. Med. 12, 198–206 (2006).
Rudolph, M.G., Stanfield, R.L. & Wilson, I.A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Nicholson, M.J., Hahn, M. & Wucherpfennig, K.W. Unusual features of self-peptide/MHC binding by autoimmune T cell receptors. Immunity 23, 351–360 (2005).
Maynard, J. et al. Structure of an autoimmune T cell receptor complexed with class II peptide-MHC: insights into MHC bias and antigen specificity. Immunity 22, 81–92 (2005).
Hahn, M., Nicholson, M.J., Pyrdol, J. & Wucherpfennig, K.W. Unconventional topology of self peptide-major histocompatibility complex binding by a human autoimmune T cell receptor. Nat. Immunol. 6, 490–496 (2005).
Li, Y. et al. Structure of a human autoimmune TCR bound to a myelin basic protein self-peptide and a multiple sclerosis-associated MHC class II molecule. EMBO J. 24, 2968–2979 (2005).
Pieper, R. et al. Biochemical identification of a mutated human melanoma antigen recognized by CD4+ T cells. J. Exp. Med. 189, 757–766 (1999).
Sundberg, E.J. et al. Minor structural changes in a mutated human melanoma antigen correspond to dramatically enhanced stimulation of a CD4+ tumor-infiltrating lymphocyte line. J. Mol. Biol. 319, 449–461 (2002).
Krogsgaard, M. et al. Agonist/endogenous peptide-MHC heterodimers drive T cell activation and sensitivity. Nature 434, 238–243 (2005).
Cebecauer, M. et al. CD8+ cytotoxic T lymphocyte activation by soluble major histocompatibility complex-peptide dimers. J. Biol. Chem. 280, 23820–23828 (2005).
van der Merwe, P.A. & Davis, S.J. Molecular interactions mediating T cell antigen recognition. Annu. Rev. Immunol. 21, 659–684 (2003).
Hennecke, J., Carfi, A. & Wiley, D.C. Structure of a covalently stabilized complex of a human αβ T-cell receptor, influenza HA peptide and MHC class II molecule, HLA-DR1. EMBO J. 19, 5611–5624 (2000).
Reinherz, E.L. et al. The crystal structure of a T cell receptor in complex with peptide and MHC class II. Science 286, 1913–1921 (1999).
Lawrence, M.C. & Colman, P.M. Shape complementarity at protein-protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Garcia, K.C. et al. Structural basis of plasticity in T cell receptor recognition of a self peptide-MHC antigen. Science 279, 1166–1172 (1998).
Kjer-Nielsen, L. et al. A structural basis for the selection of dominant αβ T cell receptors in antiviral immunity. Immunity 18, 53–64 (2003).
Chen, J.-L. et al. Structural and kinetic basis for heightened immunogenicity of T cell vaccines. J. Exp. Med. 201, 1243–1255 (2005).
Dam, J. et al. Variable MHC class I engagement by Ly49 natural killer cell receptors demonstrated by the crystal structure of Ly49C bound to H-2Kb. Nat. Immunol. 4, 1213–1222 (2003).
Reich, Z. et al. Ligand-specific oligomerization of T-cell receptor molecules. Nature 387, 617–620 (1997).
Alam, S.M. et al. Qualitative and quantitative differences in T cell receptor binding of agonist and antagonist ligands. Immunity 10, 227–237 (1999).
Baker, B.M. & Wiley, D.C. αβ T cell receptor ligand-specific oligomerization revisited. Immunity 14, 681–692 (2001).
Stewart-Jones, G.B.E., McMichael, A.J., Bell, J.I., Stuart, D.I. & Jones, E.Y. A structural basis for immunodominant human T cell receptor recognition. Nat. Immunol. 4, 657–663 (2003).
Tynan, F.E. et al. T cell receptor recognition of a 'super-bulged' major histocompatibility complex class I–bound peptide. Nat. Immunol. 6, 1114–1122 (2005).
Clements, C.S., Dunstone, M.A., Macdonald, W.A., McCluskey, J. & Rossjohn, J. Specificity on a knife-edge: the αβ T cell receptor. Curr. Opin. Struct. Biol. 16, 1–9 (2006).
Kedl, R.M., Kappler, J.W. & Marrack, P. Epitope dominance, competition and affinity maturation. Curr. Opin. Immunol. 15, 120–127 (2003).
Hoare, H.L. et al. Structural basis for a major histocompatibility complex class Ib–restricted T cell response. Nat. Immunol. 7, 256–264 (2006).
Goodnow, C.C., Sprent, J., de St Groth, B.F. & Vinuesa, C.G. Cellular and genetic mechanisms of self tolerance and autoimmunity. Nature 435, 590–597 (2005).
Gronski, M.A. et al. TCR affinity and negative regulation limit autoimmunity. Nat. Med. 10, 1234–1239 (2004).
Zehn, D. & Bevan, M.J. Cells with low avidity for a tissue-restricted antigen routinely evade central and peripheral tolerance and cause autoimmunity. Immunity 25, 261–270 (2006).
Baker, B.M., Gagnon, S.J., Biddison, W.E. & Wiley, D.C. Conversion of a T cell antagonist into an agonist by repairing a defect in the TCR/peptide/MHC interface: implications for TCR signaling. Immunity 13, 475–484 (2000).
Degano, M. et al. A functional hot spot for antigen recognition in a superagonist TCR/MHC complex. Immunity 12, 251–261 (2000).
Call, M.E. & Wucherpfennig, K.W. The T cell receptor: critical role of the membrane environment in receptor assembly and function. Annu. Rev. Immunol. 23, 101–125 (2005).
Kuhns, M.S., Davis, M.M. & Garcia, K.C. Deconstructing the form and function of the TCR/CD3 complex. Immunity 24, 133–139 (2006).
van der Merwe, P.A., Davis, S.J., Shaw, A.S. & Dustin, M.L. Cytoskeletal polarization and redistribution of cell-surface molecules during T cell recognition. Semin. Immunol. 12, 5–21 (2000).
Alarcon, B., Gil, D., Delgado, P. & Schamel, W.W. Initiation of TCR signaling: regulation within CD3 dimers. Immunol. Rev. 191, 38–46 (2003).
Frank, S.J. Receptor dimerization in GH and erythropoietin action–it takes two to tango, but how? Endocrinology 143, 2–10 (2002).
Cochran, J.R., Cameron, T.O. & Stern, L.J. The relationship of MHC-peptide binding and T cell activation probed using chemically defined MHC class II oligomers. Immunity 12, 241–250 (2000).
Adams, E.J., Chien, Y.-H. & Garcia, K.C. Structure of a γδ T cell receptor in complex with the nonclassical MHC T22. Science 308, 227–231 (2005).
Nishimura, M.I. et al. T-cell receptor repertoire in tumor-infiltrating lymphocytes. Analysis of melanoma-specific long-term lines. J. Immunother. 16, 85–94 (1994).
Boulter, J.M. et al. Stable, soluble T-cell receptor molecules for crystallization and therapeutics. Protein Eng. 16, 707–711 (2003).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Storoni, L.C., McCoy, A.J. & Read, R.J. Likelihood-enhanced fast rotation functions. Acta Crystallogr. D 60, 432–438 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
McRee, D.E. XtalView/Xfit–A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: A program to check the stereo chemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Collaborative Computational Project. No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 240–255 (1994).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallog. 24, 946–950 (1991).
Bacon, D.J. & Anderson, W.F. A fast algorithm for rendering space-filling molecule pictures. J. Mol. Graph. 6, 219–220 (1988).
Schuck, P., Perugini, M.A., Gonzales, N.R., Howlett, G.J. & Schubert, D. Size-distribution analysis of proteins by analytical ultracentrifugation: strategies and application to model systems. Biophys. J. 82, 1096–1111 (2002).
Brown, P.H. & Schuck, P. Macromolecular size-and-shape distributions by sedimentation velocity analytical ultracentrifugation. Biophys. J. 90, 4651–4661 (2006).
Acknowledgements
We thank D.M. Pardoll for critical reading of the manuscript; E.J. Sundberg for initial studies; Y. Yin and S. Li for assistance with peptide synthesis; and T. McMiller for assistance with T cell assays. Supported by the National Institutes of Health (AI036900 to R.A.M.), the Intramural Research Program of the National Institutes of Health, National Cancer Institute, Center for Cancer Research (M.I.G. and S.L.T.) and the Cancer Research Institute (L.D.).
Author information
Authors and Affiliations
Contributions
L.D., X-ray crystallographic analyses; R.J.L., P.H.B., G.X., L.T. and Q.W., biochemical and biophysical experiments; M.I.G., G.G.C. and M.I.N., derivation and characterization of T cell clones; and S.L.T. and R.A.M., project direction.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Interactions between TCR and MHC molecules in the E8-mutTPI-DR1, HA1.7-HA-DR1 and 3A6-MBP-DR2 complexes. (PDF 84 kb)
Rights and permissions
About this article
Cite this article
Deng, L., Langley, R., Brown, P. et al. Structural basis for the recognition of mutant self by a tumor-specific, MHC class II–restricted T cell receptor. Nat Immunol 8, 398–408 (2007). https://doi.org/10.1038/ni1447
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1447
This article is cited by
-
Cooperative binding of T cell receptor and CD4 to peptide-MHC enhances antigen sensitivity
Nature Communications (2022)
-
Structural basis for oligoclonal T cell recognition of a shared p53 cancer neoantigen
Nature Communications (2020)
-
DynaDom: structure-based prediction of T cell receptor inter-domain and T cell receptor-peptide-MHC (class I) association angles
BMC Structural Biology (2018)
-
Recognition of host Clr-b by the inhibitory NKR-P1B receptor provides a basis for missing-self recognition
Nature Communications (2018)
-
Minimal conformational plasticity enables TCR cross-reactivity to different MHC class II heterodimers
Scientific Reports (2012)