Abstract
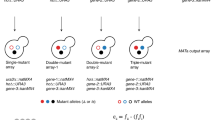
Physical and functional interactions define the molecular organization of the cell. Genetic interactions, or epistasis, tend to occur between gene products involved in parallel pathways or interlinked biological processes. High-throughput experimental systems to examine genetic interactions on a genome-wide scale have been devised for Saccharomyces cerevisiae, Schizosaccharomyces pombe, Caenorhabditis elegans and Drosophila melanogaster, but have not been reported previously for prokaryotes. Here we describe the development of a quantitative screening procedure for monitoring bacterial genetic interactions based on conjugation of Escherichia coli deletion or hypomorphic strains to create double mutants on a genome-wide scale. The patterns of synthetic sickness and synthetic lethality (aggravating genetic interactions) we observed for certain double mutant combinations provided information about functional relationships and redundancy between pathways and enabled us to group bacterial gene products into functional modules.
NOTE: In the version of this article initially published online two author names (Gabriel Moreno-Hagelseib and Constantine Christopolous) were spelled incorrectly. The correct author names are Gabriel Moreno-Hagelsieb and Constantine Christopoulos. The error has been corrected for the print, PDF and HTML versions of this article.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
17 August 2008
NOTE: In the version of this article initially published online two author names (Gabriel Moreno-Hagelseib and Constantine Christopolous) were spelled incorrectly. The correct author names are Gabriel Moreno-Hagelsieb and Constantine Christopoulos. The error has been corrected for the print, PDF and HTML versions of this article.
References
Boone, C., Bussey, H. & Andrews, B.J. Exploring genetic interactions and networks with yeast. Nat. Rev. Genet. 8, 437–449 (2007).
Costanzo, M., Giaever, G., Nislow, C. & Andrews, B. Experimental approaches to identify genetic networks. Curr. Opin. Biotechnol. 17, 472–480 (2006).
Beyer, A., Bandyopadhyay, S. & Ideker, T. Integrating physical and genetic maps: from genomes to interaction networks. Nat. Rev. Genet. 8, 699–710 (2007).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Collins, S.R., Schuldiner, M., Krogan, N.J. & Weissman, J.S. A strategy for extracting and analyzing large-scale quantitative epistatic interaction data. Genome Biol. 7, R63 (2006).
Collins, S.R. et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446, 806–810 (2007).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2 2006.0008 (2006).
Kogoma, T. Stable DNA replication: interplay between DNA replication, homologous recombination and transcription. Microbiol. Mol. Biol. Rev. 61, 212–238 (1997).
Ippen-Ihler, K.A. & Minkley, E.G. Jr. The conjugation system of F, the fertility factor of Escherichia coli. Annu. Rev. Genet. 20, 593–624 (1986).
Kowalczykowski, S.C., Dixon, D.A., Eggleston, A.K., Lauder, S.D. & Rehrauer, W.M. Biochemistry of homologous recombination in Escherichia coli. Microbiol. Mol. Biol. Rev. 58, 401–465 (1994).
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 5978–5983 (2000).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005).
Susman, M. General bacterial genetics. Annu. Rev. Genet. 4, 135–176 (1970).
Bachmann, B.J. Pedigrees of some mutant strains of Escherichia coli K-12. Bacteriol. Rev. 36, 525–557 (1972).
Fontecave, M., Ollagnier de Choudens, S. & Barras, B. Py, F. Mechanisms of iron-sulfur cluster assembly: the SUF machinery. J. Biol. Inorg. Chem. 10, 713–721 (2005).
Tokumoto, U., Kitamura, S., Fukuyama, K. & Takahashi, Y. Interchangeability and distinct properties of bacterial Fe-S cluster assembly systems: functional replacement of the isc and suf operons in Escherichia coli with the nifSU-like operon from Helicobacter pylori. J. Biochem. 136, 199–209 (2004).
Tokumoto, U. & Takahashi, Y. Genetic analysis of the isc operon in Escherichia coli involved in the biogenesis of cellular iron-sulfur proteins. J. Biochem. 130, 63–71 (2001).
Outten, F.W., Djaman, O. & Storz, G. A suf operon requirement for Fe–S cluster assembly during iron starvation in Escherichia coli. Mol. Microbiol. 52, 861–872 (2004).
Lu, J., Yang, J., Tan, G. & Ding, H. Complementary roles of SufA and IscA in the biogenesis of iron-sulfur clusters in Escherichia coli. J. Biochem. 409, 535–543 (2008).
Mani, R., St. Onge, R.P., Hartman, J.L., Giaever, G. & Roth, F.P. Defining genetic interaction. Proc. Natl. Acad. Sci. USA 105, 3461–3466 (2008).
Fox, M.S. Some features of genetic recombination in prokaryotes. Annu. Rev. Genet. 12, 47–68 (1978).
St. Onge, R.P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nat. Genet. 39, 199–206 (2007).
Von Mering, C. et al. STRING 7-recent developments in the integration and prediction of protein interactions. Nucleic Acids Res. 35, D358–D362 (2007).
Lauhon, C.T. Requirement for IscS in biosynthesis of all thionucleosides in Escherichia coli. J. Bacteriol. 184, 6820–6829 (2002).
Davierwala, A.P. et al. The synthetic genetic interaction spectrum of essential genes. Nat. Genet. 37, 1147–1152 (2005).
Ding, H., Harrison, K. & Lu, J. Thioredoxin reducatse system mediates iron binding in iscA and iron delivery for the iron-sulfur cluster assembly in IscU. J. Biol. Chem. 280, 30432–30437 (2005).
Fernandes, A.P. et al. A novel monothiol glutaredoxin (Grx4) from Escherichia coli can serve as a substrate for thioredoxin reductase. J. Biol. Chem. 280, 24544–24552 (2005).
Mizoguchi, H., Mori, H. & Fujio, T. Escherichia coli minimum genome factory. Biotechnol. Appl. Biochem. 46, 157–167 (2007).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
We thank M. Costanzo for comments on the manuscript. This work was supported by a Canadian Institute of Health Research (CIHR) grant (82852) to J.F.G. and A.E., by a US National Institutes of Health grant to B.L.W. (GM62662), by Grant-in-Aid for Scientific Research on Priority Areas from the Ministry of Education, Sports, Science and Technology of Japan Core Research for Evolutional Science and Technology, and Japan Science and Technology grants to H.M., by a CIHR grant (GSP-41567) to B.J.A. and C.B., by a partial postdoctoral fellowship from the Mexican Science and Technology Research Council to J.J.D.-M., and by a Laboratory Directed Research and Development grant to G.B. The phage-λ Red system was kindly provided by D.L. Court (National Cancer Institute).
Author information
Authors and Affiliations
Contributions
M.B. and G.B. coordinated experimental design and data analysis. M.B., G.B., H.L., J.W., P.V., C.C., L.L., M.A., K.A.D., N.Y., H.M. and B.L.W. performed optimization studies and constructed recipient arrays. J.L., A.S.-N., W.Y., A.G.G., O.P., G.B. and M.B. generated query deletion knockouts. M.B., F.B., N.S., S.C., J.-Y.C. and A.N.-A. performed genome-wide screens. M.B., S.P., A.D., S.J., B.S., H.D., B.J.A. and C.B. designed and performed quantitative image analysis. J.J.D.-M., M.B., S.P. and G.M.-H. performed informatics analyses. G.B., B.G. and W.Y. performed validation experiments. G.B., M.B., J.F.G. and A.E. drafted the manuscript. J.F.G. and A.E. conceived, designed and directed the project.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Methods, Supplementary Results, Supplementary Discussion (PDF 1941 kb)
Supplementary Table 1
List of recipient F- KEIO deletion mutant strains and SPA-tagged essential genes used in this study. (XLS 701 kb)
Supplementary Table 2
List of query deletion mutant strains used in the 39 genome-wide screens. (XLS 32 kb)
Supplementary Table 3
High-confidence genetic interaction filtered after accounting for linkage (30 kbp-window) using a stringent cut-off of |Z score| > = 4 (P < 0.0001). (XLS 284 kb)
Supplementary Table 4
Raw colony sizes (see Sheet 1), Normalized median colony sizes (see Sheet 2), |Z scores| (see Sheet 3) and interaction (S) scores (see Sheet 4) of each mutant gene pair from 39 genome-wide screens without any filtering parameters. (XLS 35759 kb)
Rights and permissions
About this article
Cite this article
Butland, G., Babu, M., Díaz-Mejía, J. et al. eSGA: E. coli synthetic genetic array analysis. Nat Methods 5, 789–795 (2008). https://doi.org/10.1038/nmeth.1239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1239