Abstract
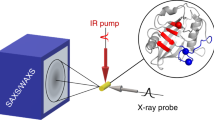
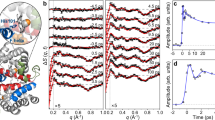
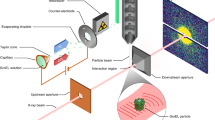
We demonstrate tracking of protein structural changes with time-resolved wide-angle X-ray scattering (TR-WAXS) with nanosecond time resolution. We investigated the tertiary and quaternary conformational changes of human hemoglobin under nearly physiological conditions triggered by laser-induced ligand photolysis. We also report data on optically induced tertiary relaxations of myoglobin and refolding of cytochrome c to illustrate the wide applicability of the technique. By providing insights into the structural dynamics of proteins functioning in their natural environment, TR-WAXS complements and extends results obtained with time-resolved optical spectroscopy and X-ray crystallography.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
29 September 2008
NOTE: In the version of this article initially published, the time scale reported in the Figure 2d legend is incorrect. The correct time scale should be 3 μs. Additionally, the time delay of 320 ms reported in Figure 5b is incorrect. The correct time delay is 200 ms. These errors have been corrected in the HTML and PDF versions of the article.
References
Moffat, K. Ultrafast time-resolved crystallography. Nat. Struct. Biol. 5, 641–643 (1998).
Schotte, F. et al. Watching a protein as it functions with 150-ps time-resolved X-ray crystallography. Science 300, 1944–1947 (2003).
Ihee, H. et al. Visualizing reaction pathways in photoactive yellow protein from nanoseconds to seconds. Proc. Natl. Acad. Sci. USA 102, 7145–7150 (2005).
Grishaev, A., Wu, J., Trewhella, J. & Bax, A. Refinement of multidomain protein structures by combination of solution small-angle X-ray scattering and NMR data. J. Am. Chem. Soc. 127, 16621–16628 (2005).
Xu, X. et al. Dynamics in a pure encounter complex of two proteins studied by solution scattering and paramagnetic NMR spectroscopy. J. Am. Chem. Soc. 130, 6395–6403 (2008).
Zuo, X. et al. Global molecular structure and interfaces: refining an RNA:RNA complex structure using solution X-ray scattering data. J. Am. Chem. Soc. 130, 3292–3293 (2008).
Hirai, M., Iwase, H., Hayakawa, T., Miura, K. & Inoue, K. Structural hierarchy of several proteins observed by wide-angle solution scattering. J. Synchrotron Radiat. 9, 202–205 (2002).
Menk, R.H. et al. Novel detector systems for time resolved SAXS experiments. J. Appl. Crystallogr. 33, 778–781 (2000).
Akiyama, S. et al. Conformational landscape of cytochrome c folding studied by microsecond-resolved small-angle X-ray scattering. Proc. Natl. Acad. Sci. USA 99, 1329–1334 (2002).
Kainosho, M. et al. Optimal isotope labelling for NMR protein structure determinations. Nature 440, 52–57 (2006).
Hofrichter, J., Sommer, J.H., Henry, E.R. & Eaton, W.A. Nanosecond absorption spectroscopy of hemoglobin: elementary processes in kinetic cooperativity. Proc. Natl. Acad. Sci. USA 80, 2235–2239 (1983).
Balakrishnan, G. et al. Time-resolved absorption and UV resonance Raman spectra reveal stepwise formation of T quaternary contacts in the allosteric pathway of hemoglobin. J. Mol. Biol. 340, 843–856 (2004).
Svergun, D.I. et al. Shape determination from solution scattering of biopolymers. J. Appl. Crystallogr. 30, 798–802 (1997).
Makowski, L. et al. Molecular crowding inhibits intramolecular breathing motions in proteins. J. Mol. Biol. 375, 529–546 (2008).
Bellelli, A., Brunori, M., Miele, A.E., Panetta, G. & Vallone, B. The allosteric properties of hemoglobin: insights from natural and site directed mutants. Curr. Protein Pept. Sci. 7, 17–45 (2006).
Eaton, W.A. et al. Evolution of allosteric models for hemoglobin. IUBMB Life 59, 586–599 (2007).
Svergun, D.I., Barberato, C. & Koch, M.H.J. CRYSOL - a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Svergun, D.I. & Koch, M.H.J. Small-angle scattering studies of biological macromolecules in solution. Rep. Prog. Phys. 66, 1735–1782 (2003).
Sawicki, C.A. & Gibson, Q.H. Quaternary conformational changes in human hemoglobin studied by laser photolysis of carboxyhemoglobin. J. Biol. Chem. 251, 1533–1542 (1976).
Cammarata, M. et al. Impulsive solvent heating probed by picosecond X-ray diffraction. J. Chem. Phys. 124, 124504(1)–124504(9) (2006).
Fader, W.J. Density perturbations caused by weak absorption of a laser pulse. J. Appl. Phys. 47, 1975–1978 (1976).
Eaton, W.A., Henry, E.R. & Hofrichter, J. Application of linear free energy relations to protein conformational changes: the quaternary structural change of hemoglobin. Proc. Natl. Acad. Sci. USA 88, 4472–4475 (1991).
Fermi, G., Perutz, M.F., Shaanan, B. & Fourme, R. The crystal structure of human deoxyhaemoglobin at 1.74 Angstrom resolution. J. Mol. Biol. 175, 159–174 (1984).
Silva, M.M., Rogers, P.H. & Arnone, A. A third quaternary structure of human hemoglobin A at 1.7 Angstrom resolution. J. Biol. Chem. 267, 17248–17256 (1992).
Lukin, J.A. et al. Quaternary structure of hemoglobin in solution. Proc. Natl. Acad. Sci. USA 100, 517–520 (2003).
Srinivasan, R. & Rose, G.D. The T-to-R transformation in hemoglobin: a reevaluation. Proc. Natl. Acad. Sci. USA 91, 11113–11117 (1994).
Safo, M.K. & Abraham, D.J. The enigma of the liganded hemoglobin end state: a novel quaternary structure of human carbonmonoxy hemoglobin. Biochemistry 44, 8347–8359 (2005).
Guallar, V., Jarzecki, A.A., Friesner, R.A. & Spiro, T.G. Modeling of ligation-induced helix/loop displacements in myoglobin: toward an understanding of hemoglobin allostery. J. Am. Chem. Soc. 128, 5427–5435 (2006).
Kachalova, G.S., Popov, A.N. & Bartunik, H.D. A steric mechanism for inhibition of CO binding to heme proteins. Science 284, 473–476 (1999).
Jones, C.M. et al. Fast events in protein-folding initiated by nanosecond laser photolysis. Proc. Natl. Acad. Sci. USA 90, 11860–11864 (1993).
Arcovito, A., Gianni, S., Brunori, M., Travaglini-Allocatelli, C. & Bellelli, A. Fast coordination changes in cytochrome c do not necessarily imply folding. J. Biol. Chem. 276, 41073–41078 (2001).
Acknowledgements
We thank W.A. Eaton and E. Henry for helpful comments, H.-S. Cho and S. Ahn for helpful discussions about the data analysis, and Y.O. Jung, K.H. Kim and J.H. Lee for their assistance with sample preparation and experiments. This research was supported in part by the Intramural Research Program of the National Institutes of Health to P.A.A., by EU grant FLASH: FP6-503641 to M.W., and a grant from the Creative Research Initiatives (Center for Time-Resolved Diffraction) of the Ministry of Education, Science and Technology, Korea Science and Engineering Foundation to H.I.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–4, Supplementary Methods (PDF 975 kb)
Rights and permissions
About this article
Cite this article
Cammarata, M., Levantino, M., Schotte, F. et al. Tracking the structural dynamics of proteins in solution using time-resolved wide-angle X-ray scattering. Nat Methods 5, 881–886 (2008). https://doi.org/10.1038/nmeth.1255
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1255
This article is cited by
-
Small-angle X-ray and neutron scattering
Nature Reviews Methods Primers (2021)
-
Tracking Membrane Protein Dynamics in Real Time
The Journal of Membrane Biology (2021)
-
Short-lived metal-centered excited state initiates iron-methionine photodissociation in ferrous cytochrome c
Nature Communications (2021)
-
Molecular model of a sensor of two-component signaling system
Scientific Reports (2021)
-
Light-induced structural changes in a full-length cyanobacterial phytochrome probed by time-resolved X-ray scattering
Communications Biology (2019)