Abstract
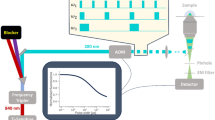
Visualizing conformational dynamics in proteins has been difficult, and the atomic-scale motions responsible for the behavior of most allosteric proteins are unknown. Here we report that fluorescence resonance energy transfer (FRET) between a small fluorescent dye and a nickel ion bound to a dihistidine motif can be used to monitor small structural rearrangements in proteins. This method provides several key advantages over classical FRET, including the ability to measure the dynamics of close-range interactions, the use of small probes with short linkers, a low orientation dependence, and the ability to add and remove unique tunable acceptors. We used this 'transition metal ion FRET' approach along with X-ray crystallography to determine the structural changes of the gating ring of the mouse hyperpolarization-activated cyclic nucleotide–regulated ion channel HCN2. Our results suggest a general model for the conformational switch in the cyclic nucleotide–binding site of cyclic nucleotide–regulated ion channels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Zagotta, W.N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003).
Selvin, P.R. Fluorescence resonance energy transfer. Methods Enzymol. 246, 300–334 (1995).
Posson, D.J. & Selvin, P.R. Extent of voltage sensor movement during gating of shaker K+ channels. Neuron 59, 98–109 (2008).
Best, R.B. et al. Effect of flexibility and cis residues in single-molecule FRET studies of polyproline. Proc. Natl. Acad. Sci. USA 104, 18964–18969 (2007).
Selvin, P.R. Principles and biophysical applications of lanthanide-based probes. Annu. Rev. Biophys. Biomol. Struct. 31, 275–302 (2002).
Richmond, T.A., Takahashi, T.T., Shimkhada, R. & Bernsdorf, J. Engineered metal binding sites on green fluorescence protein. Biochem. Biophys. Res. Commun. 268, 462–465 (2000).
Sandtner, W., Bezanilla, F. & Correa, A.M. In vivo measurement of intramolecular distances using genetically encoded reporters. Biophys. J. 93, L45–L47 (2007).
Vogel, S.S., Thaler, C. & Koushik, S.V. Fanciful FRET. Sci. STKE 2006, re2 (2006).
Lakowicz, J.R. Principles of Fluorescence Spectroscopy (Kluwer Academic/Plenum, New York, 1999).
Arnold, F.H. & Haymore, B.L. Engineered metal-binding proteins: purification to protein folding. Science 252, 1796–1797 (1991).
Suh, S.S., Haymore, B.L. & Arnold, F.H. Characterization of His-X3-His sites in alpha-helices of synthetic metal-binding bovine somatotropin. Protein Eng. 4, 301–305 (1991).
Craven, K.B. & Zagotta, W.N. CNG and HCN channels: two peas, one pod. Annu. Rev. Physiol. 68, 375–401 (2006).
Dale, R.E., Eisinger, J. & Blumberg, W.E. The orientational freedom of molecular probes. The orientation factor in intramolecular energy transfer. Biophys. J. 26, 161–193 (1979).
Figgis, B.N. & Hitchman, M.A. Ligand Field Theory and Its Applications (Wiley-VCH, New York, 2000).
Forster, T. Experimentelle und theoretische untersuchung des zwischenmolekularen ubergangs von elektronenanregungsenergie. Zeitschrift Fur Naturforschung Section A4, 321-3271949 A 4, 321–327 (1949).
Wolber, P.K. & Hudson, B.S. An analytic solution to the Forster energy transfer problem in two dimensions. Biophys. J. 28, 197–210 (1979).
Zimet, D.B., Thevenin, B.J., Verkman, A.S., Shohet, S.B. & Abney, J.R. Calculation of resonance energy transfer in crowded biological membranes. Biophys. J. 68, 1592–1603 (1995).
Miranda, P. et al. FRET with multiply labeled HERG K(+) channels as a reporter of the in vivo coarse architecture of the cytoplasmic domains. Biochim. Biophys. Acta 1783, 1681–1699 (2008).
Taraska, J.W. & Zagotta, W.N. Structural dynamics in the gating ring of cyclic nucleotide-gated ion channels. Nat. Struct. Mol. Biol. 14, 854–860 (2007).
Kellis, J.T. Jr., Todd, R.J. & Arnold, F.H. Protein stabilization by engineered metal chelation. Bio/Technology 9, 994–995 (1991).
Todd, R.J., Van Dam, M.E., Casimiro, D., Haymore, B.L. & Arnold, F.H. Cu(II)-binding properties of a cytochrome c with a synthetic metal-binding site: His-X3-His in an alpha-helix. Proteins 10, 156–161 (1991).
Rehmann, H., Wittinghofer, A. & Bos, J.L. Capturing cyclic nucleotides in action: snapshots from crystallographic studies. Nat. Rev. Mol. Cell Biol. 8, 63–73 (2007).
Altieri, S.L. et al. Structural and energetic analysis of activation by a cyclic nucleotide binding domain. J. Mol. Biol. 381, 655–669 (2008).
Kim, C., Xuong, N.H. & Taylor, S.S. Crystal structure of a complex between the catalytic and regulatory (RIalpha) subunits of PKA. Science 307, 690–696 (2005).
Rehmann, H., Das, J., Knipscheer, P., Wittinghofer, A. & Bos, J.L. Structure of the cyclic-AMP-responsive exchange factor Epac2 in its auto-inhibited state. Nature 439, 625–628 (2006).
Horrocks, W.D. Jr., Holmquist, B. & Vallee, B.L. Energy transfer between terbium (III) and cobalt (II) in thermolysin: a new class of metal–metal distance probes. Proc. Natl. Acad. Sci. USA 72, 4764–4768 (1975).
Flynn, G.E., Black, K.D., Islas, L.D., Sankaran, B. & Zagotta, W.N. Structure and rearrangements in the carboxy-terminal region of SpIH channels. Structure 15, 671–682 (2007).
McCoy, A.J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Navaza, J. AMoRe—an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron-density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Brunger, A.T. Assessment of phase accuracy by cross validation—the free R-value—methods and applications. Acta Crystallogr. D Biol. Crystallogr. 49, 24–36 (1993).
The C.C.P. 4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Schreiter, E.R. et al. Crystal structure of the nickel-responsive transcription factor NikR. Nat. Struct. Biol. 10, 794–799 (2003).
Telmer, P.G. & Shilton, B.H. Structural studies of an engineered zinc biosensor reveal an unanticipated mode of zinc binding. J. Mol. Biol. 354, 829–840 (2005).
Wray, J.W., Baase, W.A., Ostheimer, G.J., Zhang, X.J. & Matthews, B.W. Use of a non-rigid region in T4 lysozyme to design an adaptable metal-binding site. Protein Eng. 13, 313–321 (2000).
Acknowledgements
We thank K. Black, S. Cunnington, S. Simmons and G. Sheridan for technical assistance, the beamline staff at the Advanced Light Source and National Synchrotron Light Source for help with data collection, W. Almers, B. Hille and A. Merz for comments on the manuscript, and E. Gouaux and L. Islas for stimulating discussions. This work was supported by the Howard Hughes Medical Institute and National Institutes of Health (NIH) grants EY10329 to W.N.Z. and R01 NS038631 to E. Gouaux. M.C.P was supported by a Ruth Kirschstein National Research Service Award from the National Eye Institute. J.W.T was supported by a fellowship from the Jane Coffin Childs Foundation and a NIH Pathway to Independence Award (1K99NS064213).
Author information
Authors and Affiliations
Contributions
J.W.T designed the experiments and analysis. J.W.T and M.C.P. performed fluorescence experiments. N.B.O, G.E.F. and W.N.Z. performed the crystallography. J.W.T and W.N.Z. wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–2 (PDF 2096 kb)
Rights and permissions
About this article
Cite this article
Taraska, J., Puljung, M., Olivier, N. et al. Mapping the structure and conformational movements of proteins with transition metal ion FRET. Nat Methods 6, 532–537 (2009). https://doi.org/10.1038/nmeth.1341
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1341
This article is cited by
-
A high affinity switch for cAMP in the HCN pacemaker channels
Nature Communications (2024)
-
Conformational trajectory of allosteric gating of the human cone photoreceptor cyclic nucleotide-gated channel
Nature Communications (2023)
-
Direct regulation of the voltage sensor of HCN channels by membrane lipid compartmentalization
Nature Communications (2023)
-
Determining the gas-phase structures of α-helical peptides from shape, microsolvation, and intramolecular distance data
Nature Communications (2023)
-
Single-molecule imaging with cell-derived nanovesicles reveals early binding dynamics at a cyclic nucleotide-gated ion channel
Nature Communications (2021)