Abstract
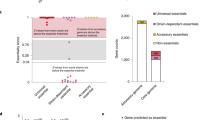
Biological pathways are structured in complex networks of interacting genes. Solving the architecture of such networks may provide valuable information, such as how microorganisms cause disease. Here we present a method (Tn-seq) for accurately determining quantitative genetic interactions on a genome-wide scale in microorganisms. Tn-seq is based on the assembly of a saturated Mariner transposon insertion library. After library selection, changes in frequency of each insertion mutant are determined by sequencing the flanking regions en masse. These changes are used to calculate each mutant's fitness. Using this approach, we determined fitness for each gene of Streptococcus pneumoniae, a causative agent of pneumonia and meningitis. A genome-wide screen for genetic interactions of five query genes identified both alleviating and aggravating interactions that could be divided into seven distinct categories. Owing to the wide activity of the Mariner transposon, Tn-seq has the potential to contribute to the exploration of complex pathways across many different species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Pan, X. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006).
Fiedler, D. et al. Functional organization of the S. cerevisiae phosphorylation network. Cell 136, 952–963 (2009).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Pan, X. et al. A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16, 487–496 (2004).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Roguev, A. et al. Conservation and rewiring of functional modules revealed by an epistasis map in fission yeast. Science 322, 405–410 (2008).
Typas, A. et al. High-throughput, quantitative analyses of genetic interactions in E. coli. Nat. Methods 5, 781–787 (2008).
Butland, G. et al. eSGA: E. coli synthetic genetic array analysis. Nat. Methods 5, 789–795 (2008).
Joshi, S.M. et al. Characterization of mycobacterial virulence genes through genetic interaction mapping. Proc. Natl. Acad. Sci. USA 103, 11760–11765 (2006).
Girgis, H. et al. A comprehensive genetic characterization of bacterial motility. PLoS Genet. 3, 1644–1660 (2007).
Lampe, D.J., Churchill, M. & Robertson, H. A purified mariner transposase is sufficient to mediate transposition in vitro. EMBO J. 15, 5470–5479 (1996).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Jacobs, M.A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 100, 14339–14344 (2003).
Fujita, Y. Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci. Biotechnol. Biochem. 73, 245–259 (2009).
Giammarinaro, P. & Paton, J. Role of RegM, a homologue of the catabolite repressor protein CcpA, in the virulence of Streptococcus pneumoniae. Infect. Immun. 70, 5454–5461 (2002).
Iyer, R., Baliga, N.S. & Camilli, A. Catabolite control protein A (CcpA) contributes to virulence and regulation of sugar metabolism in Streptococcus pneumoniae. J. Bacteriol. 187, 8340–8349 (2005).
Chapuy-Regaud, S. et al. RegR, a global LacI/GalR family regulator, modulates virulence and competence in Streptococcus pneumoniae. Infect. Immun. 71, 2615–2625 (2003).
Nieto, C., Espinosa, M. & Puyet, A. The maltose/maltodextrin regulon of Streptococcus pneumoniae differential promoter regulation by the transcriptional repressor MalR. J. Biol. Chem. 272, 30860–30865 (1997).
von Mering, C. et al. STRING 7–recent developments in the integration and prediction of protein interactions. Nucleic Acids Res. 35, D358–D362 (2007).
Mascher, T. et al. The Streptococcus pneumoniae cia regulon: CiaR target sites and transcription profile analysis. J. Bacteriol. 185, 60–70 (2003).
Throup, J.P. et al. A genomic analysis of two-component signal transduction in Streptococcus pneumoniae. Mol. Microbiol. 35, 566–576 (2000).
Sebert, M.E. et al. Microarray-based identification of htrA, a Streptococcus pneumoniae gene that is regulated by the CiaRH two-component system and contributes to nasopharyngeal colonization. Infect. Immun. 70, 4059–4067 (2002).
Drees, B.L. et al. Derivation of genetic interaction networks from quantitative phenotype data. Genome Biol. 6, R38 (2005).
St. Onge, R.P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nat. Genet. 39, 199–206 (2007).
King, S.J., Hippe, K.R. & Weiser, J. Deglycosylation of human glycoconjugates by the sequential activities of exoglycosidases expressed by Streptococcus pneumoniae. Mol. Microbiol. 59, 961–974 (2006).
Lampe, D.J. et al. Hyperactive transposase mutants of the Himar1 mariner transposon. Proc. Natl. Acad. Sci. USA 96, 11428–11433 (1999).
Bae, T. et al. Generating a collection of insertion mutations in the Staphylococcus aureus genome using bursa aurealis. Methods Mol. Biol. 416, 103–116 (2008).
Akerley, B.J. et al. Systematic identification of essential genes by in vitro mariner mutagenesis. Proc. Natl. Acad. Sci. USA 95, 8927–8932 (1998).
Sassetti, C.M., Boyd, D.H. & Rubin, E.J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 48, 77–84 (2003).
Bricker, A.L. & Camilli, A. Transformation of a type 4 encapsulated strain of Streptococcus pneumoniae. FEMS Microbiol. Lett. 172, 131–135 (1999).
Hava, D.L. & Camilli, A. Large-scale identification of serotype 4 Streptococcus pneumoniae virulence factors. Mol. Microbiol. 45, 1389–1406 (2002).
Langmead, B. et al. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Lenski, R.E., Rose, M.R.S. & Tadler, S.C. Long-term experimental evolution in Escherichia coli. I. Adaptation and divergence during 2,000 generations. Am. Nat. 138, 1315–1341 (1991).
van Opijnen, T., Boerlijst, M.C. & Berkhout, B. Effects of random mutations in the human immunodeficiency virus type 1 transcriptional promoter on viral fitness in different host cell environments. J. Virol. 80, 6678–6685 (2006).
Hava, D.L. & Camilli, A. Large-scale identification of serotype 4 Streptococcus pneumoniae virulence factors. Mol. Microbiol. 45, 1389–1406 (2002).
Marcusson, L.L. et al. Interplay in the selection of fluoroquinolone resistance and bacterial fitness. PLoS Pathog. 5, e1000541 (2009).
Shannon, P. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We thank L. Sonenshein, R. Isberg, R. Iyer and N. Vastenhouw for both discussions and comments on the manuscript, members of the Tufts University core facility for Illumina sequencing and members of the Camilli lab, particularly D. Lazinski, and A. Boorsma for helpful discussions. This work was supported by a grant from the Netherlands Organization for Scientific Research (NWO-Rubicon) to T.v.O. and by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.v.O. and A.C. designed research, analyzed the data and wrote the paper. T.v.O. performed research and K.L.B. contributed to data analysis.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1,5–7 (PDF 193 kb)
Supplementary Table 2
Wild-type library fitness values (XLS 426 kb)
Supplementary Table 3
GSEA enrichment analysis (XLS 26 kb)
Supplementary Table 4
Genetic interactions and fitness values for five query genes (XLS 47 kb)
Rights and permissions
About this article
Cite this article
van Opijnen, T., Bodi, K. & Camilli, A. Tn-seq: high-throughput parallel sequencing for fitness and genetic interaction studies in microorganisms. Nat Methods 6, 767–772 (2009). https://doi.org/10.1038/nmeth.1377
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1377
This article is cited by
-
Bacterial genome engineering using CRISPR-associated transposases
Nature Protocols (2024)
-
Barcoding Populations of Pseudomonas fluorescens SBW25
Journal of Molecular Evolution (2023)
-
Vt35 antitoxin plays a central regulatory role in virulence of Pseudomonas savastanoi pv. glycinea on soybean
Journal of General Plant Pathology (2023)
-
A bacterial pan-genome makes gene essentiality strain-dependent and evolvable
Nature Microbiology (2022)
-
The genetic architecture underlying prey-dependent performance in a microbial predator
Nature Communications (2022)