Abstract
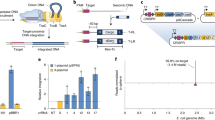
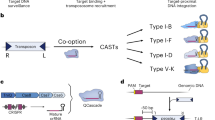
We previously constructed a series of mini-Tn7 chromosome integration vectors that, when provided only with the site-specific transposition machinery, generally transpose to a naturally evolved, neutral attTn7 site that is located 25-bp downstream of the glmS gene. Here we provide a protocol for application of the mini-Tn7 system in Proteus mirabilis as an example of a bacterium with a secondary attTn7 site that is not linked to glmS but, in this case, located in the carAB operon. The procedure involves, first, cloning of the genes of interest into an appropriate mini-Tn7 vector; second, co-transfer of the recombinant mini-Tn7 vector and a helper plasmid encoding the Tn7 site-specific transposition pathway into P. mirabilis by transformation, followed by selection of insertion-containing strains; third, PCR verification of mini-Tn7 insertions; and last, optional Flp-mediated excision of the antibiotic-resistance selection marker present on the chromosomally integrated mini-Tn7 element. When transposon-containing cells are selected on rich medium, insertions occur at both attTn7 sites with equal efficiency and frequency. Because carA mutants are arginine and pyrimidine auxotrophs, single-site insertions at the glmS attTn7 sites can be obtained by selection on minimal medium. From start to verification of the insertion events, the whole procedure takes 5 d. This chromosome integration system in P. mirabilis provides an important tool for animal and biofilm studies based on this bacterium. Vectors are available for gene complementation and expression, gene fusion analyses and tagging with a green fluorescent protein (GFP)-encoding reporter gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Warren, J.W. The catheter and urinary tract infection. Med. Clin. North. Am. 75, 481–493 (1991).
Warren, J.W., Muncie, H.L., Jr. & Hall-Craggs, M. Acute pyelonephritis associated with bacteriuria during long-term catheterization: a prospective clinicopathological study. J. Infect. Dis. 158, 1341–1346 (1988).
Belas, R. Proteus mirabilis swarmer cell differentiation and urinary tract infection. In Urinary Tract Infections: Molecular Pathogenesis and Clinical Management (eds. Mobley, H.L.T. & Warren, J.W.) 271–298 (ASM Press, Washington DC, 1996).
Fairley, K.F. et al. Site of infection in acute urinary-tract infection in general practice. Lancet 2, 615–618 (1971).
Jansen, A.M., Lockatell, C.V., Johnson, D.E. & Mobley, H.L. Visualization of Proteus mirabilis morphotypes in the urinary tract: the elongated swarmer cell is rarely observed in ascending urinary tract infection. Infect. Immun. 71, 3607–3613 (2003).
Burall, L.S. et al. Proteus mirabilis genes that contribute to pathogenesis of urinary tract infection: identification of 25 signature-tagged mutants attenuated at least 100-fold. Infect. Immun. 72, 2922–2938 (2004).
Choi, K.-H. et al. A Tn7-based broad-range bacterial cloning and expression system. Nat. Methods 2, 443–448 (2005).
Choi, K.-H. & Schweizer, H.P. mini-Tn7 insertion in bacteria with single attTn7 sites: example Pseudomonas aeruginosa. Nat. Protocols 1, 153–161 (2006).
Craig, N.L. Transposon Tn7. Curr. Top. Microbiol. Immunol. 204, 27–48 (1996).
Peters, J.E. & Craig, N.L. Tn7: smarter than we thought. Nat. Rev. Mol. Cell Biol. 2, 806–814 (2001).
Yanisch-Perron, C., Vieira, J. & Messing, J. Improved M13 cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33, 103–119 (1985).
Sambrook, J. & Russell, D.W. Molecular Cloning (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Hoang, T.T., Karkhoff-Schweizer, R.R., Kutchma, A.J. & Schweizer, H.P. A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212, 77–86 (1998).
Acknowledgements
This work was supported by a Public Health Service grant (AI058141) from the US National Institute of Allergy and Infectious Diseases (NIAID) to H.P.S.
Author information
Authors and Affiliations
Contributions
K.H.C. performed all experiments and assisted in writing the manuscript; H.P.S. supervised the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Choi, KH., Schweizer, H. mini-Tn7 insertion in bacteria with secondary, non-glmS-linked attTn7 sites: example Proteus mirabilis HI4320. Nat Protoc 1, 170–178 (2006). https://doi.org/10.1038/nprot.2006.26
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.26
This article is cited by
-
Conditional quorum-sensing induction of a cyanide-insensitive terminal oxidase stabilizes cooperating populations of Pseudomonas aeruginosa
Nature Communications (2019)
-
In vitro and in vivo efficacy, toxicity, bio-distribution and resistance selection of a novel antibacterial drug candidate
Scientific Reports (2016)
-
Design and characterization of a polyamine derivative inhibiting the expression of type III secretion system in Pseudomonas aeruginosa
Scientific Reports (2016)
-
Mini-Tn7 vectors for stable expression of diguanylate cyclase PleD* in Gram-negative bacteria
BMC Microbiology (2015)
-
Tn5/7-lux: a versatile tool for the identification and capture of promoters in Gram-negative bacteria
BMC Microbiology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.