Abstract
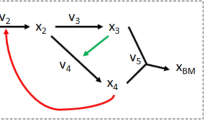
The manner in which microorganisms utilize their metabolic processes can be predicted using constraint-based analysis of genome-scale metabolic networks. Herein, we present the constraint-based reconstruction and analysis toolbox, a software package running in the Matlab environment, which allows for quantitative prediction of cellular behavior using a constraint-based approach. Specifically, this software allows predictive computations of both steady-state and dynamic optimal growth behavior, the effects of gene deletions, comprehensive robustness analyses, sampling the range of possible cellular metabolic states and the determination of network modules. Functions enabling these calculations are included in the toolbox, allowing a user to input a genome-scale metabolic model distributed in Systems Biology Markup Language format and perform these calculations with just a few lines of code. The results are predictions of cellular behavior that have been verified as accurate in a growing body of research. After software installation, calculation time is minimal, allowing the user to focus on the interpretation of the computational results.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Bork, P. Is there biological research beyond Systems Biology? A comparative analysis of terms. Mol. Syst. Biol. 1 Epub 2005 May 25 (2005).
Papin, J.A., Hunter, T., Palsson, B.O. & Subramaniam, S. Reconstruction of cellular signalling networks and analysis of their properties. Nat. Rev. Mol. Cell. Biol. 6, 99–111 (2005).
Covert, M.W., Knight, E.M., Reed, J.L., Herrgard, M.J. & Palsson, B.O. Integrating high-throughput and computational data elucidates bacterial networks. Nature 429, 92–96 (2004).
Brynildsen, M.P., Wong, W.W. & Liao, J.C. Transcriptional regulation and metabolism. Biochem. Soc. Trans. 33, 1423–1426 (2005).
Reed, J.L., Famili, I., Thiele, I. & Palsson, B.O. Towards multidimensional genome annotation. Nat. Rev. Genet. 7, 130–141 (2006).
Papin, J.A. et al. Comparison of network-based pathway analysis methods. Trends Biotechnol. 22, 400–405 (2004).
Papin, J.A. & Palsson, B.O. The JAK–STAT signaling network in the human B-cell: an extreme signaling pathway analysis. Biophys. J. 87, 37–46 (2004).
Reed, J.L., Vo, T.D., Schilling, C.H. & Palsson, B.O. An expanded genome-scale model of Escherichia coli K-12 (iJR904 GSM/GPR). Genome Biol. 4, R54 (2003).
Price, N.D., Reed, J.L. & Palsson, B.O. Genome-scale models of microbial cells: evaluating the consequences of constraints. Nat. Rev. Microbiol. 2, 886–897 (2004).
Fong, S.S. & Palsson, B.O. Metabolic gene-deletion strains of Escherichia coli evolve to computationally predicted growth phenotypes. Nat. Genet. 36, 1056–1058 (2004).
Hong, S.H. et al. The genome sequence of the capnophilic rumen bacterium Mannheimia succiniciproducens . Nat. Biotechnol. 22, 1275–1281 (2004).
David, H., Akesson, M. & Nielsen, J. Reconstruction of the central carbon metabolism of Aspergillus niger . Eur. J. Biochem./FEBS 270, 4243–4253 (2003).
Sheikh, K., Forster, J. & Nielsen, L.K. Modeling hybridoma cell metabolism using a generic genome-scale metabolic model of Mus musculus . Biotechnol. Prog. 21, 112–121 (2005).
Kuepfer, L., Sauer, U. & Blank, L.M. Metabolic functions of duplicate genes in Saccharomyces cerevisiae . Genome Res. 15, 1421–1430 (2005).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Duarte, N.C., Herrgard, M.J. & Palsson, B.O. Reconstruction and validation of Saccharomyces cerevisiae iND750, a fully compartmentalized genome-scale metabolic model. Genome Res. 14, 1298–1309 (2004).
Fong, S.S. et al. In silico design and adaptive evolution of Escherichia coli for production of lactic acid. Biotechnol. Bioeng. 91, 643–648 (2005).
Wang, Q., Chen, X., Yang, Y. & Zhao, X. Genome-scale in silico aided metabolic analysis and flux comparisons of Escherichia coli to improve succinate production. Appl. Microbiol. Biotechnol. (2006).
Alper, H., Jin, Y.S., Moxley, J.F. & Stephanopoulos, G. Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli . Metab. Eng. 7, 155–164 (2005).
Klamt, S., Stelling, J., Ginkel, M. & Gilles, E.D. FluxAnalyzer: exploring structure, pathways, and flux distributions in metabolic networks on interactive flux maps. Bioinformatics 19, 261–269 (2003).
Zamboni, N., Fischer, E. & Sauer, U. FiatFlux—a software for metabolic flux analysis from 13C-glucose experiments. BMC Bioinformatics 6, 209 (2005).
Hucka, M. et al. The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics 19, 524–531 (2003).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Varma, A. & Palsson, B.O. Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Appl. Environ. Microbiol. 60, 3724–3731 (1994).
Segre, D., Vitkup, D. & Church, G.M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. USA 99, 15112–15117 (2002).
Mahadevan, R. & Schilling, C.H. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 5, 264–276 (2003).
Reed, J.L. & Palsson, B.O. Genome-scale in silico models of E. coli have multiple equivalent phenotypic states: assessment of correlated reaction subsets that comprise network states. Genome Res. 14, 1797–1805 (2004).
Acknowledgements
We acknowledge Nathan Price, Vasiliy Portnoy, Jan Schellenberger and Christian Barrett for help with the COBRA Toolbox development and testing. Support for this work was provided by the National Institutes of Health (RO1 GM071808, 2R01 GM062791-04A2) and National Science Foundation (BES-0331342).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
These studies were supported by grants from the NIH/NSF (RO1 GM071808, 2R01 GM062791-04A2, BES-0331342). The investigator(s) have a financial interest in Genomatica, Inc. Although this grant has been identified for conflict of interest management based on the overall scope of the project and its potential to benefit Genomatica, Inc, the research findings included in this publication may not necessarily directly relate to the interests of Genomatica, Inc.
Supplementary information
Supplementary Manual
Quantitative Prediction of Cellular Metabolism with Constraint-based Models: The COBRA Toolbox (DOC 55 kb)
Rights and permissions
About this article
Cite this article
Becker, S., Feist, A., Mo, M. et al. Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox. Nat Protoc 2, 727–738 (2007). https://doi.org/10.1038/nprot.2007.99
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.99
This article is cited by
-
Engineering the synthetic β-alanine pathway in Komagataella phaffii for conversion of methanol into 3-hydroxypropionic acid
Microbial Cell Factories (2023)
-
Multivariate modular metabolic engineering for enhanced l-methionine biosynthesis in Escherichia coli
Biotechnology for Biofuels and Bioproducts (2023)
-
Co-utilization of Maltose and Sodium Acetate via Engineered Corynebacterium glutamicum for Improved Itaconic Acid Production
Biotechnology and Bioprocess Engineering (2023)
-
Monitoring and modelling the glutamine metabolic pathway: a review and future perspectives
Metabolomics (2023)
-
Substrate type and CO2 addition significantly influence succinic acid production of Basfia succiniciproducens
Biotechnology Letters (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.