Abstract
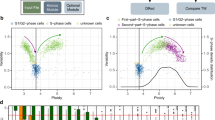
Replication timing profiles are cell type–specific and reflect genome organization changes during differentiation. In this protocol, we describe how to analyze genome-wide replication timing (RT) in mammalian cells. Asynchronously cycling cells are pulse labeled with the nucleotide analog 5-bromo-2-deoxyuridine (BrdU) and sorted into S-phase fractions on the basis of DNA content using flow cytometry. BrdU-labeled DNA from each fraction is immunoprecipitated, amplified, differentially labeled and co-hybridized to a whole-genome comparative genomic hybridization microarray, which is currently more cost effective than high-throughput sequencing and equally capable of resolving features at the biologically relevant level of tens to hundreds of kilobases. We also present a guide to analyzing the resulting data sets based on methods we use routinely. Subjects include normalization, scaling and data quality measures, LOESS (local polynomial) smoothing of RT values, segmentation of data into domains and assignment of timing values to gene promoters. Finally, we cover clustering methods and means to relate changes in the replication program to gene expression and other genetic and epigenetic data sets. Some experience with R or similar programming languages is assumed. All together, the protocol takes ∼3 weeks per batch of samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Pope, B.D., Hiratani, I. & Gilbert, D.M. Domain-wide regulation of DNA replication timing during mammalian development. Chromosome Res. 18, 127–136 (2010).
Yaffe, E. et al. Comparative analysis of DNA replication timing reveals conserved large-scale chromosomal architecture. PLoS Genet. 6, e1001011 (2010).
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010).
Gilbert, D.M. et al. Space and time in the nucleus: developmental control of replication timing and chromosome architecture. Cold Spring Harb. Symp. Quant. Biol. doi:10.1101/sqb.2010.75.011 (2010).
Hiratani, I. et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 20, 155–169 (2010).
Hiratani, I. et al. Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol. 6, e245 (2008).
Schwaiger, M. et al. Heterochromatin protein 1 (HP1) modulates replication timing of the Drosophila genome. Genome Res. 20, 771–780 (2010).
Schwaiger, M. et al. Chromatin state marks cell-type- and gender-specific replication of the Drosophila genome. Genes Dev. 23, 589–601 (2009).
Schübeler, D. et al. Genome-wide DNA replication profile for Drosophila melanogaster: a link between transcription and replication timing. Nat. Genet. 32, 438–442 (2002).
Lee, T.-J. et al. Arabidopsis thaliana chromosome 4 replicates in two phases that correlate with chromatin state. PLoS Genet. 6, e1000982 (2010).
Koren, A., Soifer, I. & Barkai, N. MRC1-dependent scaling of the budding yeast DNA replication timing program. Genome Res. 20, 781–790 (2010).
Raghuraman, M.K. & Brewer, B.J. Molecular analysis of the replication program in unicellular model organisms. Chromosome Res. 18, 19–34 (2010).
Hayashi, M. et al. Genome-wide localization of pre-RC sites and identification of replication origins in fission yeast. EMBO J. 26, 1327–1339 (2007).
Karnani, N., Taylor, C.M. & Dutta, A. Microarray analysis of DNA replication timing. Methods Mol. Biol. 556, 191–203 (2009).
Farkash-Amar, S. & Simon, I. Genome-wide analysis of the replication program in mammals. Chromosome Res. 18, 115–125 (2009).
Sasaki, T. et al. The Chinese hamster dihydrofolate reductase replication origin decision point follows activation of transcription and suppresses initiation of replication within transcription units. Mol. Cel. Biol. 26, 1051–1062 (2006).
Gilbert, D.M. Replication origin plasticity, taylor-made: inhibition vs recruitment of origins under conditions of replication stress. Chromosoma 116, 341–347 (2007).
Anglana, M. et al. Dynamics of DNA replication in mammalian somatic cells: nucleotide pool modulates origin choice and interorigin spacing. Cell 114, 385–394 (2003).
Gilbert, D.M. Evaluating genome-scale approaches to eukaryotic DNA replication. Nat. Rev. Genet. 11, 673–684 (2010).
Gilbert, D.M. Temporal order of replication of Xenopus laevis 5S ribosomal RNA genes in somatic cells. Proc. Natl. Acad. Sci. USA 83, 2924–2928 (1986).
Gilbert, D.M. & Cohen, S.N. Bovine papilloma virus plasmids replicate randomly in mouse fibroblasts throughout S phase of the cell cycle. Cell 50, 59–68 (1987).
Hansen, R.S. et al. Association of fragile X syndrome with delayed replication of the FMR1 gene. Cell 73, 1403–1409 (1993).
Yokochi, T. et al. G9a selectively represses a class of late-replicating genes at the nuclear periphery. Proc. Natl. Acad. Sci. USA 106, 19363–19368 (2009).
Lu, J. et al. G2 phase chromatin lacks determinants of replication timing. J Cell Biol. 189, 967–980 (2010).
Pollack, J.R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat. Genet. 23, 41–46 (1999).
Acevedo, L.G. et al. Genome-scale ChIP-chip analysis using 10,000 human cells. BioTechniques 43, 791–797 (2007).
Smyth, G.K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 3 (2004).
Yang, Y.H. et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 30, e15 (2002).
Bolstad, B.M. et al. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Core, R.D. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2008).
Gentleman, R.C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Ihaka, R. & Gentleman, R. R: A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Spector, P. Data Manipulation with R (Springer Publishing Company, 2008).
Chambers, J.M. Software for Data Analysis: Programming with R (Springer Publishing Company, 2008).
Crawley, M.J. The R Book (Wiley, 2007).
Wettenhall, J.M. & Smyth, G.K. limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics 20, 3705–3706 (2004).
Hansen, R.S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl. Acad. Sci. USA 107, 139–144 (2010).
Pombo, A. & Gilbert, D.M. Nucleus and gene expression: the structure and function conundrum. Curr. Opin. Cell Biol. 22, 269–270 (2010).
O'Geen, H. et al. Comparison of sample preparation methods for ChIP-chip assays. BioTechniques 41, 577–580 (2006).
Peng, S. et al. Normalization and experimental design for ChIP-chip data. BMC Bioinformatics 8, 219 (2007).
Reimers, M. & Weinstein, J.N. Quality assessment of microarrays: visualization of spatial artifacts and quantitation of regional biases. BMC Bioinformatics 6, 166 (2005).
Venkatraman, E.S. & Olshen, A.B. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007).
Dellinger, A.E. et al. Comparative analyses of seven algorithms for copy number variant identification from single nucleotide polymorphism arrays. Nucleic Acids Res. 38, e105 (2010).
Willenbrock, H. & Fridlyand, J. A comparison study: applying segmentation to array CGH data for downstream analyses. Bioinformatics 21, 4084–4091 (2005).
Lai, W.R. et al. Comparative analysis of algorithms for identifying amplifications and deletions in array CGH data. Bioinformatics 21, 3763–3770 (2005).
Olshen, A.B. et al. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Eisen, M.B. et al. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
ENCODE Project Consortium. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640 (2004).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Rosenbloom, K.R. et al. ENCODE whole-genome data in the UCSC genome browser. Nucleic Acids Res. 38, D620–D625 (2010).
Edgar, R., Domrachev, M. & Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Barrett, T. et al. NCBI GEO: archive for high-throughput functional genomic data. Nucleic Acids Res. 37, D885–D890 (2009).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Salcedo-Amaya, A.M. et al. Dynamic histone H3 epigenome marking during the intraerythrocytic cycle of Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 106, 9655–9660 (2009).
Guenther, M.G. et al. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–88 (2007).
Hon, G., Wang, W. & Ren, B. Discovery and annotation of functional chromatin signatures in the human genome. PLoS Comput. Biol. 5, e1000566 (2009).
Aladjem, M.I. et al. Replication initiation patterns in the beta-globin loci of totipotent and differentiated murine cells: evidence for multiple initiation regions. Mol. Cel. Biol. 22, 442–452 (2002).
Woodfine, K. et al. Replication timing of the human genome. Hum. Mol. Genet. 13, 191–202 (2004).
Acknowledgements
We thank J.C. Rivera Mulia and A. Rycyk for helpful comments on the manuscript. Research in the Gilbert lab is funded by NIH Grants GM083337 and GM085354.
Author information
Authors and Affiliations
Contributions
D.M.G. and I.H. conceived the study and designed the experiments. T.R. and I.H. devised the computational methods. T.R., D.B., B.D.P. and D.M.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data 1
Example data file L1210LymphoblastR1_532.pair (TXT 38582 kb)
Supplementary Data 2
Example data file L1210LymphoblastR1_635.pair (TXT 38532 kb)
Supplementary Data 3
Example data file L1210LymphoblastR2_532.pair (TXT 39297 kb)
Supplementary Data 4
Example data file L1210LymphoblastR2_635.pair (TXT 39203 kb)
Supplementary Data 5
Example targets file T.txt (TXT 0 kb)
Rights and permissions
About this article
Cite this article
Ryba, T., Battaglia, D., Pope, B. et al. Genome-scale analysis of replication timing: from bench to bioinformatics. Nat Protoc 6, 870–895 (2011). https://doi.org/10.1038/nprot.2011.328
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.328
This article is cited by
-
Optimized Repli-seq: improved DNA replication timing analysis by next-generation sequencing
Chromosome Research (2022)
-
Progerin impairs 3D genome organization and induces fragile telomeres by limiting the dNTP pools
Scientific Reports (2021)
-
Single-cell DNA replication profiling identifies spatiotemporal developmental dynamics of chromosome organization
Nature Genetics (2019)
-
5-hydroxymethylcytosine Marks Mammalian Origins Acting as a Barrier to Replication
Scientific Reports (2019)
-
Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.