Abstract
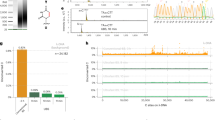
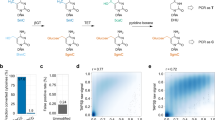
A complete understanding of the potential function of 5-hydroxymethylcytosine (5-hmC), a DNA cytosine modification in mammalian cells, requires an accurate single-base resolution sequencing method. Here we describe a modified bisulfite-sequencing method, Tet-assisted bisulfite sequencing (TAB-seq), which can identify 5-hmC at single-base resolution, as well as determine its abundance at each modification site. This protocol involves β-glucosyltransferase (β-GT)-mediated protection of 5-hmC (glucosylation) and recombinant mouse Tet1(mTet1)-mediated oxidation of 5-methylcytosine (5-mC) to 5-carboxylcytosine (5-caC). After the subsequent bisulfite treatment and PCR amplification, both cytosine and 5-caC (derived from 5-mC) are converted to thymine (T), whereas 5-hmC reads as C. The treated genomic DNA is suitable for both whole-genome and locus-specific sequencing. The entire procedure (which does not include data analysis) can be completed in 14 d for whole-genome sequencing or 7 d for locus-specific sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Penn, N.W., Suwalski, R., O'Riley, C., Bojanowski, K. & Yura, R. The presence of 5-hydroxymethylcytosine in animal deoxyribonucleic acid. Biochem. J. 126, 781–790 (1972).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Valinluck, V. et al. Oxidative damage to methyl-CpG sequences inhibits the binding of the methyl-CpG binding domain (MBD) of methyl-CpG binding protein 2 (MeCP2). Nucleic Acids Res. 32, 4100–4108 (2004).
Valinluck, V. & Sowers, L.C. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 67, 946–950 (2007).
Hashimoto, H. et al. Recognition and potential mechanisms for replication and erasure of cytosine hydroxymethylation. Nucleic Acids Res. 40, 4841–4849 (2012).
Jin, S.G., Kadam, S. & Pfeifer, G.P. Examination of the specificity of DNA methylation profiling techniques towards 5-methylcytosine and 5-hydroxymethylcytosine. Nucleic Acids Res. 38, e125 (2010).
Globisch, D. et al. Tissue distribution of 5-hydroxymethylcytosine and search for active demethylation intermediates. PloS ONE 5, e15367 (2010).
Munzel, M. et al. Quantification of the sixth DNA base hydroxymethylcytosine in the brain. Angew. Chem. Int. Ed. Engl. 49, 5375–5377 (2010).
Szwagierczak, A., Bultmann, S., Schmidt, C.S., Spada, F. & Leonhardt, H. Sensitive enzymatic quantification of 5-hydroxymethylcytosine in genomic DNA. Nucleic Acids Res. 38, e181 (2010).
Song, C.X. et al. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat. Biotechnol. 29, 68–72 (2011).
Wu, H. et al. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev. 25, 679–684 (2011).
Ito, S. et al. Role of Tet proteins in 5-mC to 5-hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Dawlaty, M.M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011).
Gu, T.P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Iqbal, K., Jin, S.G., Pfeifer, G.P. & Szabo, P.E. Reprogramming of the paternal genome upon fertilization involves genome-wide oxidation of 5-methylcytosine. Proc. Natl Acad. Sci. USA 108, 3642–3647 (2011).
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Koh, K.P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Szulwach, K.E. et al. Integrating 5-hydroxymethylcytosine into the epigenomic landscape of human embryonic stem cells. PLoS Genet. 7, e1002154 (2011).
Szulwach, K.E. et al. 5-hmC-mediated epigenetic dynamics during postnatal neurodevelopment and aging. Nat. Neurosci. 14, 1607–1616 (2011).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. Int. Ed. Engl. 50, 7008–7012 (2011).
Maiti, A. & Drohat, A.C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338 (2011).
Zhang, L. et al. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat. Chem. Biol. 8, 328–330 (2012).
Yu, M. et al. Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome. Cell 149, 1368–1380 (2012).
Huang, Y. et al. The behaviour of 5-hydroxymethylcytosine in bisulfite sequencing. PloS ONE 5, e8888 (2010).
Josse, J. & Kornberg, A. Glucosylation of deoxyribonucleic acid. III. α- and β-glucosyl transferases from T4-infected Escherichia coli. J. Biol. Chem. 237, 1968–1976 (1962).
Lariviere, L. & Morera, S. Structural evidence of a passive base-flipping mechanism for β-glucosyltransferase. J. Biol. Chem. 279, 34715–34720 (2004).
Song, C.X. et al. Detection of 5-hydroxymethylcytosine in DNA by transferring a keto-glucose by using T4 phage β-glucosyltransferase. Chembiochem 12, 1682–1685 (2011).
Pastor, W.A. et al. Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473, 394–397 (2011).
Robertson, A.B. et al. A novel method for the efficient and selective identification of 5-hydroxymethylcytosine in genomic DNA. Nucleic Acids Res. 39, e55 (2011).
Robertson, A.B., Dahl, J.A., Ougland, R. & Klungland, A. Pull-down of 5-hydroxymethylcytosine DNA using JBP1-coated magnetic beads. Nat. Protoc. 7, 340–350 (2012).
Booth, M.J. et al. Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 336, 934–937 (2012).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Xu, Y. et al. Genome-wide regulation of 5-hmC, 5-mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol. Cell. 42, 451–464 (2011).
Flusberg, B.A. et al. Direct detection of DNA methylation during single-molecule, real-time sequencing. Nat. Methods 7, 461–465 (2010).
Song, C.X. et al. Sensitive and specific single-molecule sequencing of 5-hydroxymethylcytosine. Nat. Methods 9, 75–77 (2012).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Acknowledgements
This study was supported by the US National Institutes of Health (GM071440 and HG006827 to C.H., U01 ES017166 to B.R., NS079625 and HD073162 to P.J.), a Catalyst Award (to C.H.) from the Chicago Biomedical Consortium with support from the Searle Funds at The Chicago Community Trust, the Ludwig Institute for Cancer Research (to B.R.) and the Emory Genetics Discovery Fund (to P.J.).
Author information
Authors and Affiliations
Contributions
M.Y., C.-X.S. and C.H. conceived the original idea. M.Y., C.-X.S. and C.H. designed the experiment with the help from B.R. and P.J.; M.Y. performed treatment of genomic DNA; M.Y., G.C.H. and K.E.S. performed locus-specific sequencing; and G.C.H. and K.E.S. performed genome-wide sequencing. M.Y. and C.H. drafted the manuscript, and all the authors participated in writing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Note 1
Sequence of 5hmC spike-in control (PDF 213 kb)
Supplementary Note 2
Insertion Sequence of mTet1. The mouse TET1 (1367-2039) gene with one flag tag was cloned into the insect cell expression plasmid pFastBac Dual (Invitrogen, cat. 10712-024). The restriction enzyme cutting sites are BssHII and NotI. (PDF 199 kb)
Rights and permissions
About this article
Cite this article
Yu, M., Hon, G., Szulwach, K. et al. Tet-assisted bisulfite sequencing of 5-hydroxymethylcytosine. Nat Protoc 7, 2159–2170 (2012). https://doi.org/10.1038/nprot.2012.137
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.137
This article is cited by
-
METTL3 acetylation impedes cancer metastasis via fine-tuning its nuclear and cytosolic functions
Nature Communications (2022)
-
Dual detection of chromatin accessibility and DNA methylation using ATAC-Me
Nature Protocols (2021)
-
B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.