Key Points
-
Germline polymorphisms are known to influence the pharmacokinetics and pharmacological effects of a growing number of anticancer agents.
-
Polymorphisms in the human thiopurine methyltransferase (TPMT) gene lead to loss of the functional protein and predispose with high penetrance to severe haematopoietic toxicity in TPMT-deficient patients, unless their dose of mercaptopurine is reduced by 90–95%.
-
Candidate-gene strategies have shown that germline polymorphisms in additional genes — such as the glutathione S-transferase genes, the uridine-5′-diphosphate-glucuronosyl-transferase genes and the thymidylate synthetase gene — are associated with the efficacy or toxicity of chemotherapy for acute lymphoblastic leukaemia (ALL).
-
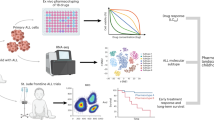
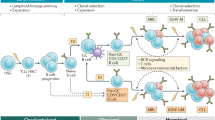
Recently, distinct gene-expression profiles of primary ALL cells have been linked to the sensitivity of leukaemia cells to several antileukaemic agents in vitro, and these expression signatures also predicted treatment outcome.
-
These findings provide momentum for future genome-wide studies to identify additional genomic determinants of ALL- treatment responses. These will allow the development of polygenic models that can be used to optimize the treatment of ALL and other human cancers.
Abstract
The use of combination chemotherapy to cure acute lymphoblastic leukaemia (ALL) in children emerged in the 1980s as a paradigm for curing any disseminated cancer, and many of the therapeutic principles were subsequently applied to the treatment of other disseminated human cancers. Similarly, elucidation of the pharmacogenomics of ALL and its translation into new chemotherapeutic approaches might serve as a model for optimizing the treatment of other human cancers. Germline polymorphisms and gene-expression patterns in ALL cells have been linked to the toxicity and efficacy of chemotherapy for ALL and are beginning to emerge as useful clinical diagnostics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Simone, J. V. Childhood leukemia as a model for cancer research: the Richard and Hinda Rosenthal Foundation Award Lecture. Cancer Res. 39, 4301–4307 (1979).
Simone, J. V. Childhood leukemia — successes and challenges for survivors. N. Engl. J. Med. 349, 627–628 (2003).
Pui, C. H. & Evans, W. E. Treatment of acute lymphoblastic leukemia. N. Engl. J. Med. 354, 166–178 (2006).
Pui, C. H., Relling, M. V. & Downing, J. R. Acute lymphoblastic leukemia. N. Engl. J. Med. 350, 1535–1548 (2004).
Pui, C. H. & Evans, W. E. Acute lymphoblastic leukemia. N. Engl. J. Med. 339, 605–615 (1998).
Pui, C. H., Campana, D. & Evans, W. E. Childhood acute lymphoblastic leukaemia — current status and future perspectives. Lancet Oncol. 2, 597–607 (2001).
Evans, W. E. & Relling, M. V. Pharmacogenomics: translating functional genomics into rational therapeutics. Science 286, 487–491 (1999).
Relling, M. V. & Dervieux, T. Pharmacogenetics and cancer therapy. Nature Rev. Cancer 1, 99–108 (2001).
Evans, W. E. & Relling, M. V. Moving towards individualized medicine with pharmacogenomics. Nature 429, 464–468 (2004).
Weinshilboum, R. Inheritance and drug response. N. Engl. J. Med. 348, 529–537 (2003).
Ulrich, C. M., Robien, K. & McLeod, H. L. Cancer pharmacogenetics: polymorphisms, pathways and beyond. Nature Rev. Cancer 3, 912–920 (2003).
Undevia, S. D., Gomez-Abuin, G. & Ratain, M. J. Pharmacokinetic variability of anticancer agents. Nature Rev. Cancer 5, 447–458 (2005).
Krynetski, E. Y. & Evans, W. E. Pharmacogenetics of cancer therapy: getting personal. Am. J. Hum. Genet. 63, 11–16 (1998).
McLeod, H. L., Krynetski, E. Y., Relling, M. V. & Evans, W. E. Genetic polymorphism of thiopurine methyltransferase and its clinical relevance for childhood acute lymphoblastic leukemia. Leukemia 14, 567–572 (2000).
Pazmino, P. A., Sladek, S. L. & Weinshilboum, R. M. Thiol S-methylation in uremia: erythrocyte enzyme activities and plasma inhibitors. Clin. Pharmacol. Ther. 28, 356–367 (1980).
Weinshilboum, R. M. & Sladek, S. L. Mercaptopurine pharmacogenetics: monogenic inheritance of erythrocyte thiopurine methyltransferase activity. Am. J. Hum. Genet. 32, 651–662 (1980).
Lennard, L., Lilleyman, J. S., Van, L. J. & Weinshilboum, R. M. Genetic variation in response to 6-mercaptopurine for childhood acute lymphoblastic leukaemia. Lancet 336, 225–229 (1990). The first study to show a relation between a patient's TPMT activity and the active metabolite of mercaptopurine (thioguanine nucleotides) in erythrocytes.
Krynetski, E. Y., Krynetskaia, N. F., Yanishevski, Y. & Evans, W. E. Methylation of mercaptopurine, thioguanine, and their nucleotide metabolites by heterologously expressed human thiopurine S-methyltransferase. Mol. Pharmacol. 47, 1141–1147 (1995).
Loennechen, T. et al. Isolation of a human thiopurine S-methyltransferase (TPMT) complementary DNA with a single nucleotide transition A719G (TPMT*3C) and its association with loss of TPMT protein and catalytic activity in humans. Clin. Pharmacol. Ther. 64, 46–51 (1998).
Tai, H. L. et al. Thiopurine S-methyltransferase deficiency: two nucleotide transitions define the most prevalent mutant allele associated with loss of catalytic activity in Caucasians. Am. J. Hum. Genet. 58, 694–702 (1996).
Otterness, D. et al. Human thiopurine methyltransferase pharmacogenetics: gene sequence polymorphisms. Clin. Pharmacol. Ther. 62, 60–73 (1997).
Yates, C. R. et al. Molecular diagnosis of thiopurine S-methyltransferase deficiency: genetic basis for azathioprine and mercaptopurine intolerance. Ann. Intern. Med. 126, 608–614 (1997). The first study to show high concordance between TPMT genotype and TPMT catalytic activity in patients.
Krynetski, E. Y. et al. A single point mutation leading to loss of catalytic activity in human thiopurine S-methyltransferase. Proc. Natl Acad. Sci. USA 92, 949–953 (1995). The first human TPMT variant allele to be cloned and characterized, followed by the identification of other variant alleles and the development of genotyping methods to diagnose TPMT deficiency.
Schaeffeler, E. et al. Comprehensive analysis of thiopurine S-methyltransferase phenotype-genotype correlation in a large population of German-Caucasians and identification of novel TPMT variants. Pharmacogenetics 14, 407–417 (2004). A study in a large cohort of patients that establishes the sensitivity and specificity of diagnosing TPMT deficiency on the basis of TPMT genotype.
Hon, Y. Y. et al. Polymorphism of the thiopurine S-methyltransferase gene in African-Americans. Hum. Mol. Genet. 8, 371–376 (1999).
Cheng, Q. et al. Karyotypic abnormalities create discordance of germline genotype and cancer cell phenotypes. Nature Genet. 37, 878–882 (2005). Initial study to show that acquisition of an additional chromosome in leukaemia cells alters the concordance of genotype and phenotype depending on whether the acquired chromosome contains a wild-type or variant allele for the gene of interest.
McLeod, H. L., Relling, M. V., Liu, Q., Pui, C. H. & Evans, W. E. Polymorphic thiopurine methyltransferase in erythrocytes is indicative of activity in leukemic blasts from children with acute lymphoblastic leukemia. Blood 85, 1897–1902 (1995).
Stanulla, M. et al. Thiopurine methyltransferase (TPMT) genotype and early treatment response to mercaptopurine in childhood acute lymphoblastic leukemia. JAMA 293, 1485–1489 (2005). This report demonstrated that TPMT genotypes were associated with the early response to treatment of acute lymphoblastic leukaemia.
Evans, W. E., Horner, M., Chu, Y. Q., Kalwinsky, D. & Roberts, W. M. Altered mercaptopurine metabolism, toxic effects, and dosage requirement in a thiopurine methyltransferase-deficient child with acute lymphocytic leukemia. J. Pediatr. 119, 985–989 (1991).
Evans, W. E. et al. Preponderance of thiopurine S-methyltransferase deficiency and heterozygosity among patients intolerant to mercaptopurine or azathioprine. J. Clin. Oncol. 19, 2293–2301 (2001).
Relling, M. V. et al. Mercaptopurine therapy intolerance and heterozygosity at the thiopurine S-methyltransferase gene locus. J. Natl Cancer Inst. 91, 2001–2008 (1999). First study to show that TPMT heterozygotes are at higher risk of mercaptopurine dose-limiting haematopoietic toxicity.
Schutz, E., Gummert, J., Armstrong, V. W., Mohr, F. W. & Oellerich, M. Azathioprine pharmacogenetics: the relationship between 6-thioguanine nucleotides and thiopurine methyltransferase in patients after heart and kidney transplantation. Eur. J. Clin. Chem. Clin. Biochem. 34, 199–205 (1996).
Lennard, L., Gibson, B. E., Nicole, T. & Lilleyman, J. S. Congenital thiopurine methyltransferase deficiency and 6-mercaptopurine toxicity during treatment for acute lymphoblastic leukaemia. Arch. Dis. Child 69, 577–579 (1993).
Bo, J. et al. Possible carcinogenic effect of 6-mercaptopurine on bone marrow stem cells: relation to thiopurine metabolism. Cancer 86, 1080–1086 (1999).
Relling, M. V. et al. Etoposide and antimetabolite pharmacology in patients who develop secondary acute myeloid leukemia. Leukemia 12, 346–352 (1998).
Evans, W. E. Thiopurine S-methyltransferase: a genetic polymorphism that affects a small number of drugs in a big way. Pharmacogenetics 12, 421–423 (2002).
Relling, M. V. et al. High incidence of secondary brain tumours after radiotherapy and antimetabolites. Lancet 354, 34–39 (1999).
Marshall, E. Preventing toxicity with a gene test. Science 302, 588–590 (2003).
Abbott, A. With your genes? Take one of these, three times a day. Nature 425, 760–762 (2003).
Chen, C. L. et al. Higher frequency of glutathione S-transferase deletions in black children with acute lymphoblastic leukemia. Blood 89, 1701–1707 (1997).
Hayes, J. D., Flanagan, J. U. & Jowsey, I. R. Glutathione transferases. Annu. Rev. Pharmacol. Toxicol. 45, 51–88 (2005).
Woo, M. H. et al. Glutathione S-transferase genotypes in children who develop treatment-related acute myeloid malignancies. Leukemia 14, 232–237 (2000).
Allan, J. M. et al. Polymorphism in glutathione S-transferase P1 is associated with susceptibility to chemotherapy-induced leukemia. Proc. Natl Acad. Sci. USA 98, 11592–11597 (2001).
Anderer, G. et al. Polymorphisms within glutathione S-transferase genes and initial response to glucocorticoids in childhood acute lymphoblastic leukaemia. Pharmacogenetics 10, 715–726 (2000).
Davies, S. M. et al. Glutathione S-transferase polymorphisms and outcome of chemotherapy in childhood acute myeloid leukemia. J. Clin. Oncol. 19, 1279–1287 (2001).
Haase, D. et al. Increased risk for therapy-associated hematologic malignancies in patients with carcinoma of the breast and combined homozygous gene deletions of glutathione transferases M1 and T1. Leuk. Res. 26, 249–254 (2002).
Kishi, S. et al. Effects of prednisone and genetic polymorphisms on etoposide disposition in children with acute lymphoblastic leukemia. Blood 103, 67–72 (2004).
Stanulla, M., Schrappe, M., Brechlin, A. M., Zimmermann, M. & Welte, K. Polymorphisms within glutathione S-transferase genes (GSTM1, GSTT1, GSTP1) and risk of relapse in childhood B-cell precursor acute lymphoblastic leukemia: a case-control study. Blood 95, 1222–1228 (2000).
Takanashi, M. et al. Impact of glutathione S-transferase gene deletion on early relapse in childhood B-precursor acute lymphoblastic leukemia. Haematologica 88, 1238–1244 (2003).
Naoe, T. et al. Analysis of genetic polymorphism in NQO1, GST-M1, GST-T1, and CYP3A4 in 469 Japanese patients with therapy-related leukemia/myelodysplastic syndrome and de novo acute myeloid leukemia. Clin. Cancer Res. 6, 4091–4095 (2000).
Stanulla, M. et al. GSTP1 and MDR1 genotypes and central nervous system relapse in childhood acute lymphoblastic leukemia. Int. J. Hematol. 81, 39–44 (2005).
Meissner, B. et al. The GSTT1 deletion polymorphism is associated with initial response to glucocorticoids in childhood acute lymphoblastic leukemia. Leukemia 18, 1920–1923 (2004).
Davies, S. M. et al. Glutathione S-transferase genotypes, genetic susceptibility, and outcome of therapy in childhood acute lymphoblastic leukemia. Blood 100, 67–71 (2002).
Rocha, J. C. et al. Pharmacogenetics of outcome in children with acute lymphoblastic leukemia. Blood 105, 4752–4758 (2005).
Relling, M. V., Hancock, M. L., Boyett, J. M., Pui, C. H. & Evans, W. E. Prognostic importance of 6-mercaptopurine dose intensity in acute lymphoblastic leukemia. Blood 93, 2817–2823 (1999).
Pieters, R. et al. Hypoxanthine-guanine phosphoribosyl-transferase in childhood leukemia: relation with immunophenotype, in vitro drug resistance and clinical prognosis. Int. J. Cancer 51, 213–217 (1992).
Zaza, G. et al. Gene expression and thioguanine nucleotide disposition in acute lymphoblastic leukemia after in vivo mercaptopurine treatment. Blood 106, 1778–1785 (2005).
Scherf, U. et al. A gene expression database for the molecular pharmacology of cancer. Nature Genet. 24, 236–244 (2000). Description of a cancer-drug discovery resource that integrates large-scale drug sensitivities with baseline gene-expression profiles in 60 cancer cell lines.
Zaza, G. et al. Acute lymphoblastic leukemia with TEL–AML1 fusion has lower expression of genes involved in purine metabolism and lower de novo purine synthesis. Blood 104, 1435–1441 (2004).
Ross, M. E. et al. Classification of pediatric acute lymphoblastic leukemia by gene expression profiling. Blood 102, 2951–2959 (2003).
Coulthard, S. A. et al. The effect of thiopurine methyltransferase expression on sensitivity to thiopurine drugs. Mol. Pharmacol. 62, 102–109 (2002).
Dervieux, T. et al. Differing contribution of thiopurine methyltransferase to mercaptopurine versus thioguanine effects in human leukemic cells. Cancer Res. 61, 5810–5816 (2001).
Estlin, E. J., Lowis, S. P. & Hall, A. G. Optimizing antimetabolite-based chemotherapy for the treatment of childhood acute lymphoblastic leukaemia. Br. J. Haematol. 110, 29–40 (2000).
Jansen, G. et al. A structurally altered human reduced folate carrier with increased folic acid transport mediates a novel mechanism of antifolate resistance. J. Biol. Chem. 273, 30189–30198 (1998).
Wong, S. C. et al. Impaired membrane transport in methotrexate-resistant CCRF–CEM cells involves early translation termination and increased turnover of a mutant reduced folate carrier. J. Biol. Chem. 274, 10388–10394 (1999).
Chango, A. et al. A polymorphism (80G->A) in the reduced folate carrier gene and its associations with folate status and homocysteinemia. Mol. Genet. Metab. 70, 310–315 (2000).
Laverdiere, C., Chiasson, S., Costea, I., Moghrabi, A. & Krajinovic, M. Polymorphism G80A in the reduced folate carrier gene and its relationship to methotrexate plasma levels and outcome of childhood acute lymphoblastic leukemia. Blood 100, 3832–3834 (2002).
Whetstine, J. R. et al. Single nucleotide polymorphisms in the human reduced folate carrier: characterization of a high-frequency G/A variant at position 80 and transport properties of the His(27) and Arg(27) carriers. Clin. Cancer Res. 7, 3416–3422 (2001).
Levy, A. S. et al. Reduced folate carrier and dihydrofolate reductase expression in acute lymphocytic leukemia may predict outcome: a Children's Cancer Group Study. J. Pediatr. Hematol. Oncol. 25, 688–695 (2003).
Belkov, V. M. et al. Reduced folate carrier expression in acute lymphoblastic leukemia: a mechanism for ploidy but not lineage differences in methotrexate accumulation. Blood 93, 1643–1650 (1999).
Kager, L. et al. Folate pathway gene expression differs in subtypes of acute lymphoblastic leukemia and influences methotrexate pharmacodynamics. J. Clin. Invest. 115, 110–117 (2005). A candidate-gene approach that linked levels of active metabolites of methotrexate in leukaemic cells to the expression of folate-pathway genes in leukaemia cells.
Cheng, Q. et al. A substrate specific functional polymorphism of human γ-glutamyl hydrolase alters catalytic activity and methotrexate polyglutamate accumulation in acute lymphoblastic leukaemia cells. Pharmacogenetics 14, 557–567 (2004).
Krajinovic, M., Costea, I. & Chiasson, S. Polymorphism of the thymidylate synthase gene and outcome of acute lymphoblastic leukaemia. Lancet 359, 1033–1034 (2002).
Lauten, M., Asgedom, G., Welte, K., Schrappe, M. & Stanulla, M. Thymidylate synthase gene polymorphism and its association with relapse in childhood B-cell precursor acute lymphoblastic leukemia. Haematologica 88, 353–354 (2003).
Frosst, P. et al. A candidate genetic risk factor for vascular disease: a common mutation in methylenetetrahydrofolate reductase. Nature Genet. 10, 111–113 (1995).
Molloy, A. M. et al. Thermolabile variant of 5,10-methylenetetrahydrofolate reductase associated with low red-cell folates: implications for folate intake recommendations. Lancet 349, 1591–1593 (1997).
Ulrich, C. M. et al. Pharmacogenetics of methotrexate: toxicity among marrow transplantation patients varies with the methylenetetrahydrofolate reductase C677T polymorphism. Blood 98, 231–234 (2001).
Skibola, C. F. et al. Polymorphisms in the methylenetetrahydrofolate reductase gene are associated with susceptibility to acute leukemia in adults. Proc. Natl Acad. Sci. USA 96, 12810–12815 (1999).
Wiemels, J. L. et al. Methylenetetrahydrofolate reductase (MTHFR) polymorphisms and risk of molecularly defined subtypes of childhood acute leukemia. Proc. Natl Acad. Sci. USA 98, 4004–4009 (2001).
Krajinovic, M. et al. Role of polymorphisms in MTHFR and MTHFD1 genes in the outcome of childhood acute lymphoblastic leukemia. Pharmacogenomics J. 4, 66–72 (2004).
Synold, T. W. et al. Blast cell methotrexate-polyglutamate accumulation in vivo differs by lineage, ploidy, and methotrexate dose in acute lymphoblastic leukemia. J. Clin. Invest. 94, 1996–2001 (1994).
Galpin, A. J. et al. Differences in folylpolyglutamate synthetase and dihydrofolate reductase expression in human B-lineage versus T-lineage leukemic lymphoblasts: mechanisms for lineage differences in methotrexate polyglutamylation and cytotoxicity. Mol. Pharmacol. 52, 155–163 (1997).
Barredo, J. C. et al. Differences in constitutive and post-methotrexate folylpolyglutamate synthetase activity in B-lineage and T-lineage leukemia. Blood 84, 564–569 (1994).
Armstrong, S. A. et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nature Genet. 30, 41–47 (2002).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999). First study to show that gene-expression profiling can accurately discriminate ALL from AML.
Ramaswamy, S. & Golub, T. R. DNA microarrays in clinical oncology. J. Clin. Oncol. 20, 1932–1941 (2002).
Moos, P. J. et al. Identification of gene expression profiles that segregate patients with childhood leukemia. Clin. Cancer Res. 8, 3118–3130 (2002).
Ross, M. E. et al. Gene expression profiling of pediatric acute myelogenous leukemia. Blood 104, 3679–3687 (2004).
Yeoh, E.-J. et al. Classification, subtype discovery, and prediction of outcome in pediatric acute lymphoblastic leukemia by gene expression profiling. Cancer Cell 1, 133–143 (2002). First study to show distinct gene-expression patterns in ALL of different lineage and molecular subgroups.
Teuffel, O. et al. Gene expression profiles and risk stratification in childhood acute lymphoblastic leukemia. Haematologica 89, 801–808 (2004).
Willenbrock, H., Juncker, A. S., Schmiegelow, K., Knudsen, S. & Ryder, L. P. Prediction of immunophenotype, treatment response, and relapse in childhood acute lymphoblastic leukemia using DNA microarrays. Leukemia 18, 1270–1277 (2004).
Qian, Z., Fernald, A. A., Godley, L. A., Larson, R. A. & Le Beau, M. M. Expression profiling of CD34+ hematopoietic stem/progenitor cells reveals distinct subtypes of therapy-related acute myeloid leukemia. Proc. Natl Acad. Sci. USA 99, 14925–14930 (2002).
Brown, P. et al. FLT3 inhibition selectively kills childhood acute lymphoblastic leukemia cells with high levels of FLT3 expression. Blood 105, 812–820 (2005).
Edick, M. J. et al. Lymphoid gene expression as a predictor of risk of secondary brain tumors. Genes Chromosomes Cancer 42, 107–116 (2005).
Staunton, J. E. et al. Chemosensitivity prediction by transcriptional profiling. Proc. Natl Acad. Sci. USA 98, 10787–10792 (2001).
Holleman, A. et al. Gene-expression patterns in drug-resistant acute lymphoblastic leukemia cells and response to treatment. N. Engl. J. Med. 351, 533–542 (2004). First study to show that gene-expression patterns in drug-sensitive and drug-resistant ALL cells differ significantly and predict treatment outcome.
Lugthart, S. et al. Identification of genes associated with chemotherapy crossresistance and treatment response in childhood acute lymphoblastic leukemia. Cancer Cell 7, 375–386 (2005).
French, D. et al. Global gene expression as a function of germline genetic variation. Hum. Mol. Genet. 14, 1621–1629 (2005).
Cheok, M. H. et al. Treatment-specific changes in gene expression discriminate in vivo drug response in human leukemia cells. Nature Genet. 34, 85–90 (2003). First study to show that antileukaemic agents evoke treatment-specific changes in gene expression in ALL cells in vivo.
Evans, W. E. & Guy, R. K. Gene expression as a drug discovery tool. Nature Genet. 36, 214–215 (2004).
Stegmaier, K. et al. Gene expression-based high-throughput screening(GE-HTS) and application to leukemia differentiation. Nature Genet. 36, 257–263 (2004).
Hofmann, W. K. et al. Relation between resistance of Philadelphia-chromosome-positive acute lymphoblastic leukaemia to the tyrosine kinase inhibitor STI571 and gene-expression profiles: a gene-expression study. Lancet 359, 481–486 (2002).
Golub, T. R. Mining the genome for combination therapies. Nature Med. 9, 510–511 (2003).
Kennedy, G. C. et al. Large-scale genotyping of complex DNA. Nature Biotechnol. 21, 1233–1237 (2003).
Irving, J. A. et al. Loss of heterozygosity in childhood acute lymphoblastic leukemia detected by genome-wide microarray single nucleotide polymorphism analysis. Cancer Res. 65, 3053–3058 (2005).
Takeuchi, S. et al. Long-term study of the clinical significance of loss of heterozygosity in childhood acute lymphoblastic leukemia. Leukemia 17, 149–154 (2003).
Sherr, C. J. The INK4a–ARF network in tumour suppression. Nature Rev. Mol. Cell Biol. 2, 731–737 (2001).
Innocenti, F. et al. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J. Clin. Oncol. 22, 1382–1388 (2004).
van't Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Chen, C. D. et al. Molecular determinants of resistance to antiandrogen therapy. Nature Med. 10, 33–39 (2004).
Pomeroy, S. L. et al. Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature 415, 436–442 (2002).
Roepman, P. et al. An expression profile for diagnosis of lymph node metastases from primary head and neck squamous cell carcinomas. Nature Genet. 37, 182–186 (2005).
Evans, W. E. & McLeod, H. L. Pharmacogenomics — drug disposition, drug targets, and side effects. N. Engl. J. Med. 348, 538–549 (2003).
Matherly, L. H. et al. Increased frequency of expression of elevated dihydrofolate reductase in T-cell versus B-precursor acute lymphoblastic leukemia in children. Blood 90, 578–589 (1997).
Zaza, G. et al. Thiopurine Pathway [online], <http://www.pharmgkb.org/search/pathway/thiopurine/thiopurine.jsp> (2004).
Krynetski, E. Y. et al. Genetic polymorphism of thiopurine S-methyltransferase: clinical importance and molecular mechanisms. Pharmacogenetics 6, 279–290 (1996).
French, D. et al. Methotrexate Pathway [online], <http://www.pharmgkb.org/search/pathway/mtx/methotrexate.jsp> (2004).
Acknowledgements
This work was supported in part by National Institutes of Health grants and by the American Lebanese Syrian Associated Charities. We would also like to thank C.-H. Pui, M.V. Relling and M. Schwab for helpful discussions; and J.R. Davies for editorial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
National Cancer Institute
FURTHER INFORMATION
Japanese Single Nucleotide Polymorphisms database
NCBIs Single Nucleotide Polymorphism Database
Glossary
- Pharmacogenomics
-
Often used synonymously with pharmacogenetics, but also used to refer to polygenic strategies within pharmacogenetics.
- Germline polymorphism
-
The existence of two or more variants of a gene locus (alleles, sequence variants or chromosomal structural variants) at significant frequencies (usually >1%) in the population.
- Pharmacogenetics
-
Focuses on the inherited variability in drug targets, target pathways, drug absorption, distribution, metabolism, transport and elimination, or in genes that indirectly influence drug response, and how these genetic variations influence drug-response phenotypes.
- Minimal residual disease
-
Tumour cells remaining in patients after treatment (for example, surgical debulking, chemotherapy or radiotherapy), often at sub-microscopic levels.
- Dose escalation
-
The administration of successively higher doses of drug to a cohort of patients until a subset of these patients experience unacceptable side effects. The highest dose at which patients do not experience serious side effects is the maximum tolerated dose.
- Purine-salvage pathway
-
The synthesis of nucleotides from purine bases, which are recycled to produce the corresponding nucleotides after phosphoribosylation. A key enzyme in this pathway is hypoxanthine-guanine phosphoribosyltransferase.
- ABC transporters
-
Transmembrane proteins that transport a wide variety of substrates (for example, metabolites, lipids, sterols and drugs). Overexpression can lead to drug-resistant cancer. There are 48 known human ABC transporters in 7 distinct subfamilies.
- Penetrance
-
The proportion of individuals who develop a phenotype when they inherit a polymorphism that is associated with this phenotype. If a polymorphism has complete penetrance, the phenotype is present in all carriers, whereas reduced or incomplete penetrance exists if the phenotype is not always present.
- Loss of heterozygosity
-
Loss of one allele in a cell (for example, a tumour cell) of a gene (or chromosomal region) that is heterozygous in normal cells of the same individual.
- Laser-capture microdissection
-
Laser-aided removal of cells from a tissue sample, especially when it consists of different cell types (for example, normal and cancer cells). Individual cells are attached to a transfer film for further analysis.
Rights and permissions
About this article
Cite this article
Cheok, M., Evans, W. Acute lymphoblastic leukaemia: a model for the pharmacogenomics of cancer therapy. Nat Rev Cancer 6, 117–129 (2006). https://doi.org/10.1038/nrc1800
Issue Date:
DOI: https://doi.org/10.1038/nrc1800
This article is cited by
-
Mathematical models of leukaemia and its treatment: a review
SeMA Journal (2022)
-
Results of a phase II clinical trial of 6-mercaptopurine (6MP) and methotrexate in patients with BRCA-defective tumours
British Journal of Cancer (2020)
-
Targeting EIF4E signaling with ribavirin in infant acute lymphoblastic leukemia
Oncogene (2019)
-
Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9
Nature Biotechnology (2016)
-
Integrative computational in-depth analysis of dysregulated miRNA-mRNA interactions in drug-resistant pediatric acute lymphoblastic leukemia cells: an attempt to obtain new potential gene-miRNA pathways involved in response to treatment
Tumor Biology (2016)