Key Points
-
Gene expression in the adult cerebral cortex has been compared in humans, chimpanzees and other primates in several independent microarray studies.
-
These studies yield qualitatively similar results when analysed in similar ways, despite differences in the cortical regions examined and the non-human species that were compared with humans.
-
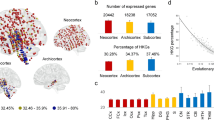
A large number of genes, representing at least 2% of expressed sequences, are differentially expressed in the cortex of humans compared with chimpanzees, the animals that are most closely related to humans.
-
There is a bias towards genes showing increased, rather than decreased, expression levels in human cortex (that is, upregulation). This bias persists even when artefacts introduced by sequence differences between species are considered.
-
Tissues other than the brain do not show such a bias, but show similar numbers of genes with increased and decreased expression levels.
-
Gene-expression changes in the adult brain that occurred during human evolution were more pronounced than those that occurred during chimpanzee evolution.
-
There were in fact, fewer gene-expression changes between humans and chimpanzees in the brain than in non-neural tissue, such as that of the liver and heart.
-
Gene-ontology analyses indicate that the human brain was modified to support higher levels of neural activity.
Abstract
Several recent microarray studies have compared gene-expression patterns n humans, chimpanzees and other non-human primates to identify evolutionary changes that contribute to the distinctive cognitive and behavioural characteristics of humans. These studies support the surprising conclusion that the evolution of the human brain involved an upregulation of gene expression relative to non-human primates, a finding that could be relevant to understanding human cerebral physiology and function. These results show how genetic and genomic methods can shed light on the basis of human neural and cognitive specializations, and have important implications for neuroscience, anthropology and medicine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Carroll, S. B. Genetics and the making of Homo sapiens. Nature 422, 849–857 (2003).
Hacia, J. G. Genome of the apes. Trends Genet. 17, 637–645 (2001).
Gagneux, P. & Varki, A. Genetic differences between humans and great apes. Mol. Phylogenet. Evol. 18, 2–13 (2001).
Gearing, M., Tigges, J., Mori, H. & Mirra, S. S. β-Amyloid (Aβ) deposition in the brains of aged orangutans. Neurobiol. Aging 18, 139–146 (1997).
Gearing, M., Rebeck, G. W., Hyman, B. T., Tigges, J. & Mirra, S. S. Neuropathology and apolipoprotein E profile of aged chimpanzees: implications for Alzheimer disease. Proc. Natl Acad. Sci. USA 91, 9382–9386 (1994).
Walker, L. C. & Cork, L. C. in Alzheimer's Disease 2nd edn (eds Terry, R. D., Katzman, R., Bick, K. L. & Sisodia, S. S.) 233–243 (Lippincott Williams & Wilkins, Philadelphia, 1999).
Rutjens, E. et al. Lentivirus infections and mechanisms of disease resistance in chimpanzees. Front Biosci. 8, d1134–1145 (2003).
McClure, H. M. Tumors in nonhuman primates: observations during a six-year period in the Yerkes primate center colony. Am. J. Phys. Anthropol. 38, 425–429 (1973).
Blurton Jones, N. G., Hawkes, K. & O'Connell, J. F. Antiquity of postreproductive life: are there modern impacts on hunter-gatherer postreproductive life spans? Am. J. Human Biol. 14, 184–205 (2002).
Hawkes, K., O'Connell, J. F., Jones, N. G., Alvarez, H. & Charnov, E. L. Grandmothering, menopause, and the evolution of human life histories. Proc. Natl Acad. Sci. USA 95, 1336–1339 (1998).
McConkey, E. H. & Varki, A. A primate genome project deserves high priority. Science 289, 1295–1296 (2000).
Olson, M. V. & Varki, A. Sequencing the chimpanzee genome: insights into human evolution and disease. Nature Rev. Genet. 4, 20–28 (2003).
Varki, A. A chimpanzee genome project is a biomedical imperative. Genome Res. 10, 1065–1070 (2000).
Watanabe, H. et al. DNA sequence and comparative analysis of chimpanzee chromosome 22. Nature 429, 382–388 (2004). The first comparison of the whole sequence of a human chromosome and its chimpanzee homologue, identifying numerous insertions and deletions, and changes in most of the encoded proteins.
Chou, H. H. et al. A mutation in human CMP-sialic acid hydroxylase occurred after the Homo-Pan divergence. Proc. Natl Acad. Sci. USA 95, 11751–11756 (1998).
Stedman, H. H. et al. Myosin gene mutation correlates with anatomical changes in the human lineage. Nature 428, 415–418 (2004).
Gilad, Y., Man, O., Paabo, S. & Lancet, D. Human specific loss of olfactory receptor genes. Proc. Natl Acad. Sci. USA 100, 3324–3327 (2003).
Zhang, J. Evolution of the human ASPM gene, a major determinant of brain size. Genetics 165, 2063–2070 (2003).
Evans, P. D. et al. Adaptive evolution of ASPM, a major determinant of cerebral cortical size in humans. Hum. Mol. Genet. 13, 489–494 (2004).
Kouprina, N. et al. Accelerated evolution of the ASPM gene controlling brain size begins prior to human brain expansion. PLoS Biol 2, E126 (2004).
Evans, P. D., Anderson, J. R., Vallender, E. J., Choi, S. S. & Lahn, B. T. Reconstructing the evolutionary history of microcephalin, a gene controlling human brain size. Hum. Mol. Genet. 13, 1139–1145 (2004).
Enard, W. et al. Molecular evolution of FOXP2, a gene involved in speech and language. Nature 418, 869–872 (2002). A study of the molecular evolution of the first gene related to a human speech and language disorder; the encoded protein contains two human-specific amino acids in positions that are conserved in all other species.
Lai, C. S., Fisher, S. E., Hurst, J. A., Vargha-Khadem, F. & Monaco, A. P. A forkhead-domain gene is mutated in a severe speech and language disorder. Nature 413, 519–523 (2001).
Teramitsu, I., Kudo, L. C., London, S. E., Geschwind, D. H. & White, S. A. Parallel FoxP1 and FoxP2 expression in songbird and human brain predicts functional interaction. J. Neurosci. 24, 3152–3163 (2004).
Clark, A. G. et al. Inferring nonneutral evolution from human-chimp-mouse orthologous gene trios. Science 302, 1960–1963 (2003). The first large-scale comparison of human, chimpanzee and mouse coding sequences, identifying hundreds of genes that probably underwent selective changes during human evolution.
King, M. C. & Wilson, A. C. Evolution at two levels in humans and chimpanzees. Science 188, 107–116 (1975). A classic paper in human evolutionary biology, in which the authors argue that phenotypic differences between humans and chimpanzees mainly reflect changes in the regulation of gene expression.
De Beer, G. Embryos and Ancestors (The Clarendon Press, Oxford, 1940).
Gould, S. J. Ontogeny and Phylogeny (Belknap, Harvard Univ. Press, US, 1977).
Bonner, J. T. & Dawid, I. (eds). Evolution and Development: Report of the Dahlem Workshop on Evolution and Development (Springer, Berlin; New York, 1982).
Raff, R. A. The Shape of Life: Genes, Development, and the Evolution of Animal Form (Univ. Chicago Press, US, 1996).
Carroll, S. B., Grenier, J. K. & Weatherbee, S. D. From DNA to Diversity: Molecular Genetics and the Evolution of Animal Design (Blackwell Science, US, 2001).
Cáceres, M. et al. Elevated gene expression levels distinguish human from non-human primate brains. Proc. Natl Acad. Sci. USA 100, 13030–13035 (2003). The second independent microarray comparison of the human and chimpanzee cerebral cortices, which provides strong support for upregulation of gene-expression changes during human brain evolution.
Enard, W. et al. Intra- and interspecific variation in primate gene expression patterns. Science 296, 340–343 (2002). The first microarray study of gene-expression patterns in primates and the first to propose that gene-expression changes accelerated during human brain evolution.
Karaman, M. W. et al. Comparative analysis of gene-expression patterns in human and African great ape cultured fibroblasts. Genome Res. 13, 1619–1630 (2003). This microarray study compares fibroblasts among a large number of humans and apes, and provides strong evidence for the confounding effect of interspecies sequence differences in microarray results.
Uddin, M. et al. Sister grouping of chimpanzees and humans as revealed by genome-wide phylogenetic analysis of brain-gene expression profiles. Proc. Natl Acad. Sci. USA 101, 2957–2962 (2004).
Mirnics, K. & Pevsner, J. Progress in the use of microarray technology to study the neurobiology of disease. Nature Neurosci. 7, 434–439 (2004).
Karsten, S. & Geschwind, D. in Current Protocols in Neuroscience (eds Crawley, J. et al.) Suppl. 20, Section 4: Unit 4. 28. (John Wiley & Sons, Inc., New York, 2002).
Geschwind, D. H. Mice, microarrays, and the genetic diversity of the brain. Proc. Natl Acad. Sci. USA 97, 10676–10678 (2000). An early discussion of the key issues, such as tissue heterogeneity, that complicate studies of gene expression in the mammalian brain.
Luo, L. et al. Gene expression profiles of laser-captured adjacent neuronal subtypes. Nature Med. 5, 117–122 (1999).
Bahn, S. et al. Gene expression profiling in the post-mortem human brain — no cause for dismay. J. Chem. Neuroanat. 22, 79–94 (2001).
Li, J. Z. et al. Systematic changes in gene expression in postmortem human brains associated with tissue pH and terminal medical conditions. Hum. Mol. Genet. 13, 609–616 (2004).
Hillis, D. M., Moritz, C. & Mable, B. K. Molecular Systematics (Sinauer Associates, Sunderland, Massachusetts, 1996).
Felsenstein, J. Inferring Phylogenies (Sinauer Associates, Sunderland, Massachusetts, 2004).
Fujiyama, A. et al. Construction and analysis of a human-chimpanzee comparative clone map. Science 295, 131–134 (2002).
Thomas, J. W. et al. Comparative analyses of multi-species sequences from targeted genomic regions. Nature 424, 788–793 (2003).
Chee, M. et al. Accessing genetic information with high-density DNA arrays. Science 274, 610–614 (1996).
Quackenbush, J. Computational analysis of microarray data. Nature Rev. Genet. 2, 418–427 (2001). A useful review of methods for the normalization and analysis of microarray data.
Geschwind, D. H. et al. A genetic analysis of neural progenitor differentiation. Neuron 29, 325–339 (2001).
Chuaqui, R. F. et al. Post-analysis follow-up and validation of microarray experiments. Nature Genet. 32 (Suppl.), 509–514 (2002).
Griffin, T. J. et al. Complementary profiling of gene expression at the transcriptome and proteome levels in Saccharomyces cerevisiae. Mol. Cell Proteomics 1, 323–333 (2002).
Washburn, M. P. et al. Protein pathway and complex clustering of correlated mRNA and protein expression analyses in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 100, 3107–3112 (2003).
Marvanova, M. et al. Microarray analysis of nonhuman primates: validation of experimental models in neurological disorders. FASEB J. 17, 929–931 (2003).
Gu, J. & Gu, X. Induced gene expression in human brain after the split from chimpanzee. Trends Genet. 19, 63–65 (2003). A statistical re-analysis of the results from reference 33 and the first to argue for the predominance of upregulation of gene expression during human brain evolution.
Hsieh, W. P., Chu, T. M., Wolfinger, R. D. & Gibson, G. Mixed-model reanalysis of primate data suggests tissue and species biases in oligonucleotide-based gene expression profiles. Genetics 165, 747–757 (2003).
Gu, J. & Gu, X. Further statistical analysis for genome-wide expression evolution in primate brain/liver/fibroblast tissues. Hum. Genomics 1, 247–254 (2004).
Khaitovich, P. et al. Regional patterns of gene expression in human and chimpanzee brains. Genome Res. 14, 1462–1473 (2004). The first extensive regional analysis of gene expression in human and chimpanzee brains at one developmental stage (adult).
Pennisi, E. Primate evolution. Gene activity clocks brain's fast evolution. Science 296, 233–235 (2002).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genet. 25, 25–29 (2000).
Preuss, T. M. in The Cognitive Neurosciences 3rd edn (ed. Gazzaniga, M. S.) (MIT Press, Cambridge, Massachusetts, in the press). Reviews current evidence regarding evolutionary specializations of the human brain.
Cavalieri, P. & Singer, P. The Great Ape Project: Equality Beyond Humanity (St. Martin's, New York, 1994).
de Waal, F. B. M. Chimpanzee Politics: Power and Sex Among Apes (Harper & Row, New York, 1982).
Povinelli, D. J. Folk Physics for Apes: the Chimpanzee's Theory of How the World Works (Oxford Univ. Press, Oxford; New York, 2000).
Povinelli, D. J. & Preuss, T. M. Theory of mind: evolutionary history of a cognitive specialization. Trends Neurosci. 18, 418–424 (1995).
Savage-Rumbaugh, E. S. & Lewin, R. Kanzi: the Ape at the Brink of the Human Mind (Wiley, New York, 1994).
Tomasello, M., Call, J. & Hare, B. Chimpanzees understand psychological states — the question is which ones and to what extent. Trends Cogn. Sci. 7, 153–156 (2003).
Wallman, J. Aping Language (Cambridge Univ. Press, Cambridge; New York, 1992).
Nimchinsky, E. A. et al. neuronal morphologic type unique to humans and great apes. Proc. Natl Acad. Sci. USA 96, 5268–5273 (1999).
Morris, C. A. et al. GTF2I hemizygosity implicated in mental retardation in Williams syndrome: genotype-phenotype analysis of five families with deletions in the Williams syndrome region. Am. J. Med. Genet. 123A, 45–59 (2003).
Doniger, S. W. et al. MAPPFinder: using Gene Ontology and GenMAPP to create a global gene-expression profile from microarray data. Genome Biol. 4, R7 (2003).
Hosack, D. A., Dennis, G. Jr, Sherman, B. T., Lane, H. C. & Lempicki, R. A. Identifying biological themes within lists of genes with EASE. Genome Biol. 4, R70 (2003).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nature Genet. 34, 166–176 (2003).
Marques-Bonet, T. et al. Chromosomal rearrangements and the genomic distribution of gene-expression divergence in humans and chimpanzees. Trends Genet. 20, 524–528 (2004).
Enard, W. et al. Differences in DNA methylation patterns between humans and chimpanzees. Curr. Biol. 14, R148–149 (2004).
Khaitovich, P. et al. A neutral model of transcriptome evolution. PLoS Biol 2, E132 (2004).
Haesler, S. et al. FoxP2 expression in avian vocal learners and non-learners. J. Neurosci. 24, 3164–3175 (2004).
Stewart, C. B. & Disotell, T. R. Primate evolution — in and out of Africa. Curr. Biol. 8, R582–588 (1998).
Fleagle, J. G. Primate Adaptation and Evolution (Academic Press, San Diego, 1998).
Acknowledgements
T.M.P. and M.C. contributed equally to this work. We thank the different authors of the primate microarray studies for making their data sets publicly available, James Thomas for the comparison of the array probes to the chimpanzee genome sequence, and David Kornack for his insightful comments. We would also like to thank the James S. McDonnell Foundation for their support of our research through a Collaborative Activities Grant (T.P., D.H.G.) and acknowledge support from the National Institute of Mental Health (D.H.G.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
OMIM
FURTHER INFORMATION
Affymetrix GeneChip Microarray Data Files
Affymetrix GeneChips in TeraGenomics
Glossary
- HUMAN SPECIALIZATION
-
A phenotypic characteristic of modern humans that has emerged since the divergence of the human lineage from the common ancestor of humans and chimpanzees.
- POSITIVE SELECTION
-
A form of evolutionary change in which a mutation has a favourable effect and increases its frequency in the population at a rate greater than that predicted by neutral drift.
- AGONAL STATE
-
The state of an individual during the time immediately preceeding death. For example, prolonged hypoxia or acidosis, can significantly affect gene expression.
- OUTGROUP METHOD
-
Comparison of closely related species to infer the state of a common ancestor.
- OLIGONUCLEOTIDE ARRAY
-
A microarray made with synthetic probes, usually 25–60 bases long, each designed to hybridize to a specific mRNA. These are fabricated either in situ or by deposition and attachment onto a solid surface. The oligonucleotide arrays that are currently available can measure expression levels for 10,000–40,000 genes simultaneously.
- INDELS
-
Insertions or deletions of DNA sequences in chromosomes.
- NORTHERN BLOT
-
An experimental technique for determining the abundance and size of the transcript(s) for a particular gene in a given tissue. mRNAs are separated electrophoretically on a gel and then transferred to a membrane (blot) by capillary action. The membrane is then immersed in a labelled probe designed to hybridize to a specific mRNA.
- RT-PCR
-
Reverse transcription PCR. Using the enzyme reverse transcriptase, RNA is converted into DNA, which is then amplified with specific primers.
- NORMALIZATION
-
Mathematical processing of raw data to reduce the effects of variables introduced by the experimental design or method used. For microarrays, such variables might include differences in fluorescent dye incorporation, the amount of cRNA or cDNA hybridized to the array, hybridization conditions or the arrays themselves.
- cDNA MICROARRAY
-
A microarray made by deposition of gene-specific, PCR-amplified inserts from cDNA clones, which can be from several hundred to several thousand bases long. cDNA arrays typically measure expression levels for 5,000–30,000 genes.
- QUANTITATIVE REAL-TIME RT-PCR
-
A procedure in which DNA amplification in a PCR reaction is measured during its log-linear phase by monitoring the accumulating signal that is provided by a fluorescent dye or gene-specific fluorescent probe incorporated into the PCR product.
- DISTANCE METRIC
-
A measure of similarity or dissimilarity that can be used to organize groups according to their degree of relation to one another. For example, the Euclidian distance metric that distinguishes two genes, or groups of genes, is often defined as the square root of the sum of their squared expression differences.
- GENE ONTOLOGY
-
A framework for classifying gene products hierarchically in three dimensions according to the biological process in which they are involved, the molecular function that they perform and the cellular component in which they are located.
- NETWORK ANALYSIS
-
Analysis of the individual interactions between constituents, which, when grouped together, describe a network. In the case of gene-expression data, network analysis entails the identification of relationships among genes or groups of genes across different experimental conditions or tissue samples.
- DIVERSIFYING SELECTION
-
Natural selection against the mean value of a quantitative trait, therefore favouring individuals at the two tails of the phenotypic distribution.
Rights and permissions
About this article
Cite this article
Preuss, T., Cáceres, M., Oldham, M. et al. Human brain evolution: insights from microarrays. Nat Rev Genet 5, 850–860 (2004). https://doi.org/10.1038/nrg1469
Issue Date:
DOI: https://doi.org/10.1038/nrg1469
This article is cited by
-
Evolution of DNA methylation in the human brain
Nature Communications (2021)
-
Cognitive-behavioral phenotypes of Williams syndrome are associated with genetic variation in the GTF2I gene, in a healthy population
BMC Neuroscience (2014)
-
Aerobic glycolysis in the primate brain: reconsidering the implications for growth and maintenance
Brain Structure and Function (2014)
-
Emerging roles of astrocytes in neural circuit development
Nature Reviews Neuroscience (2013)
-
Genes and human brain evolution
Nature (2012)