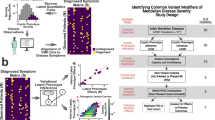
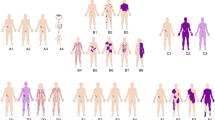
Abstract
The study of inherited monogenic diseases has contributed greatly to our mechanistic understanding of pathogenic mutations and gene regulation, and to the development of effective diagnostic tools. But interest has gradually shifted away from monogenic diseases, which collectively affect only a small fraction of the world's population, towards multifactorial, common diseases. The quest for the genetic variability associated with common traits should not be done at the expense of Mendelian disorders, because the latter could still contribute greatly to understanding the aetiology of complex traits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
The ENCODE (ENCyclopedia Of DNA Elements) project. Science 306, 636–640 (2004).
Hamosh, A., Scott, A. F., Amberger, J. S., Bocchini, C. A. & McKusick, V. A. Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res. 33, D514–D517 (2005).
Antonarakis, S. E. & McKusick, V. A. OMIM passes the 1,000-disease-gene mark. Nature Genet. 25, 11 (2000).
Stenson, P. D. et al. Human Gene Mutation Database (HGMD): 2003 update. Hum. Mutat. 21, 577–581 (2003).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Sonnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434, 462–469 (2005).
Bittles, A. Consanguinity and its relevance to clinical genetics. Clin. Genet. 60, 89–98 (2001).
Hackstein, J. H., Hochstenbach, R. & Pearson, P. L. Towards an understanding of the genetics of human male infertility: lessons from flies. Trends Genet. 16, 565–572 (2000).
Strachan, T., Abitbol, M., Davidson, D. & Beckmann, J. S. A new dimension for the human genome project: towards comprehensive expression maps. Nature Genet. 16, 126–132 (1997).
de Vries, B. B. et al. Diagnostic genome profiling in mental retardation. Am. J. Hum. Genet. 77, 606–616 (2005).
Vissers, L. E. et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nature Genet. 36, 955–957 (2004).
Auwerx, J. et al. The European dimension for the mouse genome mutagenesis program. Nature Genet. 36, 925–927 (2004).
Austin, C. P. et al. The knockout mouse project. Nature Genet. 36, 921–924 (2004).
Ben Yaou, R. et al. Genetics of laminopathies. Novartis Found. Symp. 264, 81–90; discussion 90–97, 227–230 (2005).
Liu, J. et al. Dysferlin, a novel skeletal muscle gene, is mutated in Miyoshi myopathy and limb girdle muscular dystrophy. Nature Genet. 20, 31–36 (1998).
Bashir, R. et al. A gene related to Caenorhabditis elegans spermatogenesis factor fer-1 is mutated in limb-girdle muscular dystrophy type 2B. Nature Genet. 20, 37–42 (1998).
Gambetti, P., Parchi, P. & Chen, S. G. Hereditary Creutzfeldt–Jakob disease and fatal familial insomnia. Clin. Lab. Med. 23, 43–64 (2003).
Phinney, A. L. et al. Mouse models of Alzheimer's disease: the long and filamentous road. Neurol. Res. 25, 590–600 (2003).
Kuehn, M. R., Bradley, A., Robertson, E. J. & Evans, M. J. A potential animal model for Lesch–Nyhan syndrome through introduction of HPRT mutations into mice. Nature 326, 295–298 (1987).
Pearson, C. E., Edamura, K. N. & Cleary, J. D. Repeat instability: mechanisms of dynamic mutations. Nature Rev. Genet. 6, 729–742 (2005).
Rabionet, R., Gasparini, P. & Estivill, X. Molecular genetics of hearing impairment due to mutations in gap junction genes encoding β connexins. Hum. Mutat. 16, 190–202 (2000).
Buratti, E., Brindisi, A., Pagani, F. & Baralle, F. E. Nuclear factor TDP-43 binds to the polymorphic TG repeats in CFTR intron 8 and causes skipping of exon 9: a functional link with disease penetrance. Am. J. Hum. Genet. 74, 1322–1325 (2004).
Cutting, G. R. Modifier genetics: cystic fibrosis. Annu. Rev. Genomics Hum. Genet. 6 237–260 (2005).
Weatherall, D. J. Phenotype–genotype relationships in monogenic disease: lessons from the thalassaemias. Nature Rev. Genet. 2, 245–255 (2001).
Karmous-Benailly, H. et al. Antenatal presentation of Bardet–Biedl syndrome may mimic Meckel syndrome. Am. J. Hum. Genet. 76, 493–504 (2005).
Hinds, D. A. et al. Whole-genome patterns of common DNA variation in three human populations. Science 307, 1072–1079 (2005).
Consortium., T. I. H. The international HapMap project. Nature 426, 789–796 (2003).
Altshuler, D. et al. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Abbadi, N. et al. Additional case of female monozygotic twins discordant for the clinical manifestations of Duchenne muscular dystrophy due to opposite X-chromosome inactivation. Am. J. Med. Genet. 52, 198–206 (1994).
Bras, J. M. et al. G2019S dardarin substitution is a common cause of Parkinson's disease in a Portuguese cohort. Mov. Disord. 20, 1653–1655 (2005).
Fraga, M. F. et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proc. Natl Acad. Sci. USA 102, 10604–10609 (2005).
Kajiwara, K., Berson, E. L. & Dryja, T. P. Digenic retinitis pigmentosa due to mutations at the unlinked peripherin/RDS and ROM1 loci. Science 264, 1604–1608 (1994).
Katsanis, N. et al. Triallelic inheritance in Bardet–Biedl syndrome, a Mendelian recessive disorder. Science 293, 2256–2259 (2001).
Beckmann, J. S. The Reunion paradox and the digenic model. Am. J. Hum. Genet. 59, 1400–1402 (1996).
Wang, D., Johnson, A. D., Papp, A. C., Kroetz, D. L. & Sadee, W. Multidrug resistance polypeptide 1 (MDR1, ABCB1) variant 3435C>T affects mRNA stability. Pharmacogenet. Genomics 15, 693–704 (2005).
Williams, T. N. et al. An immune basis for malaria protection by the sickle cell trait. PLoS Med. 2, e128 (2005).
Nagel, R. L. Epistasis and the genetics of human diseases. C. R. Biol. 328, 606–615 (2005).
Nadeau, J. H. Modifier genes in mice and humans. Nature Rev. Genet. 2, 165–174 (2001).
Antonarakis, S. E., Krawczak, M., Cooper, D. M. in The Metabolic and Molecular Basis of Inherited Disease (eds Scriver, C. R. et al.) 343–378 (McGraw–Hill, New York, 2001).
Cohen, J. C. et al. Multiple rare alleles contribute to low plasma levels of HDL cholesterol. Science 305, 869–872 (2004).
Frikke-Schmidt, R., Nordestgaard, B. G., Jensen, G. B. & Tybjaerg-Hansen, A. Genetic variation in ABC transporter A1 contributes to HDL cholesterol in the general population. J. Clin. Invest. 114, 1343–1353 (2004).
Lindsay, E. A. et al. Schizophrenia and chromosomal deletions within 22q11.2. Am. J. Hum. Genet. 56, 1502–1503 (1995).
Millar J. K., et al. DISC1 and PDE4B are interacting genetic factors in schizophrenia that regulate cAMP signaling. Science 310, 1187–1191 (2005).
Harper, P. S., Harley, H. G., Reardon, W. & Shaw, D. J. Anticipation in myotonic dystrophy: new light on an old problem. Am. J. Hum. Genet. 51, 10–16 (1992).
Ibanez, P. et al. Causal relation between α-synuclein gene duplication and familial Parkinson's disease. Lancet 364, 1169–1171 (2004).
Ledbetter, D. H. & Engel, E. Uniparental disomy in humans — development of an imprinting map and its implications for prenatal diagnosis. Hum. Mol. Genet. 4, 1757–1764 (1995).
da Rocha, S. T. & Ferguson-Smith, A. C. Genomic imprinting. Curr. Biol. 14, R646–R649 (2004).
Tupler, R. & Gabellini, D. Molecular basis of facioscapulohumeral muscular dystrophy. Cell. Mol. Life Sci. 61, 557–566 (2004).
Rovelet-Lecrux, A. et al. APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nature Genet. 38, 24–26 (2006).
Singleton, A. B. et al. α-Synuclein locus triplication causes Parkinson's disease. Science 302, 841 (2003).
Eriksen, J. L., Przedborski, S. & Petrucelli, L. Gene dosage and pathogenesis of Parkinson's disease. Trends Mol. Med. 11, 91–96 (2005).
Sharp, A. J. et al. Segmental duplications and copy-number variation in the human genome. Am. J. Hum. Genet. 77, 78–88 (2005).
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Feuk, L., Carson, A. R. & Scherer, S. W. Structural variation in the human genome. Nature Rev. Genet. 7, 85–97 (2006).
Aitman, T. J. et al. Copy number polymorphism in Fcgr3 predisposes to glomerulonephritis in rats and humans. Nature 439, 851–855 (2006).
Dermitzakis, E. T., Reymond, A. & Antonarakis, S. E. Conserved non-genic sequences — an unexpected feature of mammalian genomes. Nature Rev. Genet. 6, 151–157 (2005).
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nature Rev. Genet. 5, 522–531 (2004).
Eddy, S. R. Non-coding RNA genes and the modern RNA world. Nature Rev. Genet. 2, 919–929 (2001).
Brunner, H. G. & van Driel, M. A. From syndrome families to functional genomics. Nature Rev. Genet. 5, 545–551 (2004).
Gavin, A. C. & Superti-Furga, G. Protein complexes and proteome organization from yeast to man. Curr. Opin. Chem. Biol. 7, 21–27 (2003).
von Mering, C. et al. Comparative assessment of large-scale data sets of protein–protein interactions. Nature 417, 399–403 (2002).
D'Andrea, A. D. & Grompe, M. The Fanconi anaemia/BRCA pathway. Nature Rev. Cancer 3, 23–34 (2003).
Acknowledgements
We thank all the past and present members of our laboratories for ideas, debates and discussions. Mrs J. Amberger at OMIM, John Hopkins University, is gratefully acknowledged for her assistance in providing the data of figure 1a. S.E.A.'s laboratory is supported by the Swiss National Science Foundation (SNF), European Union, National Institutes of Health and the Lejeune and ChildCare Foundations. Work in J.S.B.'s laboratory is funded by grants from the SNF and the University of Lausanne.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
OMIM
facioscapulohumeral muscular dystrophy
limb girdle muscular dystrophy type 2B
FURTHER INFORMATION
Online Mendelian Inheritance in Man (OMIM)
Human Gene Mutation Database (HGMD)
Glossary
- Anticipation
-
A phenomenon that describes the property of some disorders to increase in severity in successive generations.
- Copy-number polymorphism (CNP) or variant (CNV)
-
A structural genomic variant, resulting in confined copy-number changes in a defined chromosomal region. If its population allele frequency is less than 1%, then one refers to it as a variant; the term polymorphism refers instead to variants that occur at an allelic frequency of 1% or higher.
- Discordant monozygotic twins
-
Twins that share identical genomes but are not affected with the same disorder; one twin being affected but the other not.
- Epistatic interaction
-
The influence of the interaction of multiple loci on phenotypic variation.
- Expressivity
-
The extent to which a genetic defect is expressed.
- High-resolution tiling path CGH arrays
-
Arrays for comparative genomic hybridization (CGH) offering a resolution in the order of kilobases. The arrays are currently based on BACs or long oligonucleotides.
- Mendelian disease
-
A disease for which alternative genotypes fall into distinct, discrete phenotypic classes, following Gregor Mendel's laws of inheritance.
- Monogenic disorder
-
A disease that is mainly caused by variants in a single gene.
- Multifactorial
-
Multifactorial traits are determined by a combination of genetic as well as non-genetic factors, each contributing to the overall phenotype.
- Parental imprinting
-
Differential allele expression that depends on parental origin.
- Penetrance
-
The frequency with which a given genotype manifests itself in a given phenotype.
- Triplet-repeat copy number
-
Triplet-repeat copy-number mutations are dynamic mutations that are due to the expansion or contraction in the number of triplet nucleotide repeats.
- Uniparental disomy
-
(UPD). A state wherein both homologues (alleles) at a locus derive from the same parent; for some chromosomal segments, UPD generates characteristic syndromes.
Rights and permissions
About this article
Cite this article
Antonarakis, S., Beckmann, J. Mendelian disorders deserve more attention. Nat Rev Genet 7, 277–282 (2006). https://doi.org/10.1038/nrg1826
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg1826
This article is cited by
-
Rare disorders have many faces: in silico characterization of rare disorder spectrum
Orphanet Journal of Rare Diseases (2022)
-
Severe testing with high-dimensional omics data for enhancing biomedical scientific discovery
npj Systems Biology and Applications (2022)
-
Cruxome: a powerful tool for annotating, interpreting and reporting genetic variants
BMC Genomics (2021)
-
The impact of non-additive genetic associations on age-related complex diseases
Nature Communications (2021)
-
Altered evolution: are reproductive endocrinology and infertility specialists ready for the genetically engineered future?
Journal of Assisted Reproduction and Genetics (2020)